

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
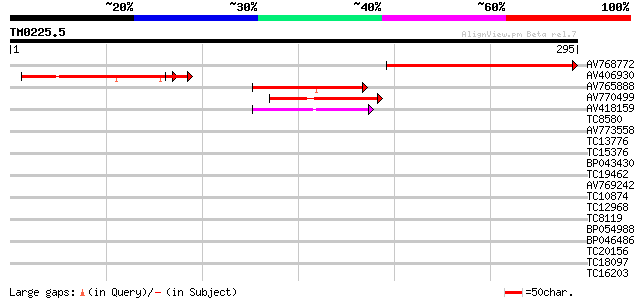
Sequences producing significant alignments: (bits) Value
AV768772 196 4e-51
AV406930 100 1e-24
AV765888 69 1e-12
AV770499 49 1e-06
AV418159 49 1e-06
TC8580 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (NAD... 32 0.17
AV773558 31 0.28
TC13776 similar to GB|AAO44003.1|28466789|BT004737 At5g24165 {Ar... 30 0.63
TC15376 weakly similar to UP|Q7SYH2 (Q7SYH2) Cystatin domain fet... 29 1.1
BP043430 29 1.1
TC19462 similar to UP|Q9SI74 (Q9SI74) F23N19.12, partial (5%) 28 1.8
AV769242 28 1.8
TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxyge... 28 2.4
TC12968 weakly similar to UP|AAS45959 (AAS45959) Envelope glycop... 28 2.4
TC8119 similar to UP|Q9FFF5 (Q9FFF5) Nucleic acid binding protei... 27 3.1
BP054988 27 3.1
BP046486 27 3.1
TC20156 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {Ar... 27 4.1
TC18097 similar to UP|Q9LNH5 (Q9LNH5) F21D18.5, partial (10%) 27 4.1
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 27 4.1
>AV768772
Length = 529
Score = 196 bits (498), Expect = 4e-51
Identities = 98/99 (98%), Positives = 99/99 (99%)
Frame = -2
Query: 197 FAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGS 256
FAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVT+GS
Sbjct: 528 FAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTNGS 349
Query: 257 DLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
DLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES
Sbjct: 348 DLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 232
>AV406930
Length = 433
Score = 100 bits (249), Expect(2) = 1e-24
Identities = 60/95 (63%), Positives = 67/95 (70%), Gaps = 14/95 (14%)
Frame = +3
Query: 7 HHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTSIALSFT----NADSAYAF 62
H + LH H+ + P P +LSLL+S TPKTCF+S LILCTS+ALSFT N DSA AF
Sbjct: 123 HLSKSLHLHTPHHPISHP-TLSLLQSPTPKTCFDSALILCTSLALSFTLFFTNTDSASAF 299
Query: 63 VVTPPRKLQSDELAT----------VVYITNLAVK 87
VVTPPRKLQSDELAT VVYITNLAVK
Sbjct: 300 VVTPPRKLQSDELATVRLFQENTPSVVYITNLAVK 404
Score = 28.5 bits (62), Expect(2) = 1e-24
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +1
Query: 82 TNLAVKMRLRWMCW 95
T L+ +MRLRWMCW
Sbjct: 391 TLLSSRMRLRWMCW 432
>AV765888
Length = 581
Score = 68.6 bits (166), Expect = 1e-12
Identities = 33/62 (53%), Positives = 47/62 (75%), Gaps = 2/62 (3%)
Frame = -3
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYS--PSGASSGVGFSIPLDTVNGIVDQLVK 184
+ +DAAIN GNSGGPL+DS G++IG+NT+ ++ +G SSGV F+IP+DTV V L+
Sbjct: 357 IQTDAAINAGNSGGPLIDSRGHVIGVNTSTFTRKGTGVSSGVNFAIPIDTVIRNVPYLIV 178
Query: 185 FG 186
+G
Sbjct: 177 YG 172
>AV770499
Length = 491
Score = 48.9 bits (115), Expect = 1e-06
Identities = 24/59 (40%), Positives = 36/59 (60%)
Frame = -2
Query: 136 GNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILG 194
GNSGGPLL+ G ++G+N + G FSIP+D+V+ +++ L K G + RP G
Sbjct: 490 GNSGGPLLNMDGEVVGVNIM---KAKDVDGFSFSIPIDSVDKVIEHLQKRGSLERPRKG 323
>AV418159
Length = 282
Score = 48.9 bits (115), Expect = 1e-06
Identities = 25/63 (39%), Positives = 34/63 (53%)
Frame = +3
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAAINPGNSGGP + G IG+ +Y S + +G+ IP V+ + K G
Sbjct: 96 IQIDAAINPGNSGGPAFNDQGECIGVAFQVYR-SEEAENIGYVIPTTVVSHFLTDYEKNG 272
Query: 187 KVT 189
K T
Sbjct: 273 KYT 281
>TC8580 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (NAD+) ,
partial (37%)
Length = 852
Score = 31.6 bits (70), Expect = 0.17
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 11/133 (8%)
Frame = -2
Query: 38 CFNSILILCTSIALSFTNADSAYAFVVTPPRKLQSDELATVVYITN-LAVKMRLRWMCWR 96
C +I C+++ S S Y F P K + + ++ Y+T L + L W
Sbjct: 482 CIKAIYPHCSNLQCSSQCICSVYVFCEDPSCKAITCVICSLNYLTKVLEL*YGLHWTKDL 303
Query: 97 FLKEGNIVTNYHVIPGA-SDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNL------ 149
FL N+HVI D L++ + +I+P G P+L++ N+
Sbjct: 302 FL------CNHHVILDI*EDSRLNIEAFV-----TKSISPSFKGSPILNTGSNILQYFLK 156
Query: 150 ---IGINTAIYSP 159
I + T +YSP
Sbjct: 155 LV*INLRTLLYSP 117
>AV773558
Length = 469
Score = 30.8 bits (68), Expect = 0.28
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +3
Query: 7 HHKQRLHYHSHYSPHFTP 24
HH H+H H+ PH+TP
Sbjct: 324 HHHHHHHHHHHHHPHYTP 377
Score = 28.1 bits (61), Expect = 1.8
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = +3
Query: 4 HYKHHKQRLHYHSHYSP 20
H+ HH H+H HY+P
Sbjct: 327 HHHHHHHHHHHHPHYTP 377
>TC13776 similar to GB|AAO44003.1|28466789|BT004737 At5g24165 {Arabidopsis
thaliana;}, partial (56%)
Length = 505
Score = 29.6 bits (65), Expect = 0.63
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +1
Query: 1 MIQHYKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCF 39
M+ H+ QRL HS P F+PN L L + C+
Sbjct: 187 MVVHHSFQVQRLQQHSEGRPLFSPNELPCLIRRLRQFCW 303
>TC15376 weakly similar to UP|Q7SYH2 (Q7SYH2) Cystatin domain fetuin-like
protein, partial (6%)
Length = 582
Score = 28.9 bits (63), Expect = 1.1
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +1
Query: 3 QHYKHHKQRLHYHSHYSPHFTP 24
+HY HH + H+H H+ H P
Sbjct: 394 RHYHHHHRHRHHHRHHHHHPLP 459
>BP043430
Length = 493
Score = 28.9 bits (63), Expect = 1.1
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +1
Query: 7 HHKQRLHYHSHYSPHFTPNSLSLL 30
HH+ LH HSH+ H+ P L LL
Sbjct: 247 HHR*ILHSHSHHHHHYYPPLLHLL 318
>TC19462 similar to UP|Q9SI74 (Q9SI74) F23N19.12, partial (5%)
Length = 535
Score = 28.1 bits (61), Expect = 1.8
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = -2
Query: 1 MIQHYKHHKQRLHYHSH 17
+ Q++ HH R HYHSH
Sbjct: 309 LYQYHYHHHHRRHYHSH 259
>AV769242
Length = 219
Score = 28.1 bits (61), Expect = 1.8
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 4/33 (12%)
Frame = +3
Query: 12 LHYHSHYSPHF-TPNSLSLLRSSTPK---TCFN 40
+HYH H+ P T +L+L S PK TCFN
Sbjct: 99 VHYHDHHDPRIPTSTTLTLPFSFHPKSITTCFN 197
>TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxygenase-like
protein, partial (38%)
Length = 723
Score = 27.7 bits (60), Expect = 2.4
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 4/32 (12%)
Frame = -2
Query: 29 LLRSSTPKTCFNSILILCT----SIALSFTNA 56
LL++ K F SIL +CT SI +SFTN+
Sbjct: 584 LLQTKEEKQLFQSILYICTYIVYSICVSFTNS 489
>TC12968 weakly similar to UP|AAS45959 (AAS45959) Envelope glycoprotein,
partial (4%)
Length = 400
Score = 27.7 bits (60), Expect = 2.4
Identities = 13/33 (39%), Positives = 18/33 (54%), Gaps = 1/33 (3%)
Frame = +1
Query: 2 IQHYKHHKQRLHYHSHYSPHFTP-NSLSLLRSS 33
+ H + R H+H H+ HF P + L LRSS
Sbjct: 220 LHHERPRLPRRHHHRHHRTHFPPCHRLHTLRSS 318
>TC8119 similar to UP|Q9FFF5 (Q9FFF5) Nucleic acid binding protein-like,
partial (85%)
Length = 1249
Score = 27.3 bits (59), Expect = 3.1
Identities = 19/49 (38%), Positives = 22/49 (44%), Gaps = 15/49 (30%)
Frame = +1
Query: 4 HYKHHKQRLHYH---------SHYSPHF----TPNSLSLLRS--STPKT 37
HY HH H+H H P F PNSLS L S S+P+T
Sbjct: 124 HYHHHHHHHHHHHMHLRSLLLMHPHPSF*NQPNPNSLSDLNSMASSPRT 270
>BP054988
Length = 442
Score = 27.3 bits (59), Expect = 3.1
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +2
Query: 4 HYKHHKQRLHYHSHY 18
HY HH LH+H H+
Sbjct: 122 HYHHHHHLLHHHGHH 166
>BP046486
Length = 452
Score = 27.3 bits (59), Expect = 3.1
Identities = 12/44 (27%), Positives = 20/44 (45%)
Frame = +1
Query: 3 QHYKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILC 46
+H H+ R+H H+HY H + + P C +L+ C
Sbjct: 295 RHQCCHQHRVHIHTHYYNHNHQTQMLQQEPNIPSCC---VLLTC 417
>TC20156 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {Arabidopsis
thaliana;}, partial (10%)
Length = 504
Score = 26.9 bits (58), Expect = 4.1
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = +2
Query: 158 SPSGASSGVGFSIPLDTVNGIVDQLVKF 185
SPS +++G+ FS L VDQL KF
Sbjct: 176 SPSLSTTGIRFSHDLKIATAFVDQLFKF 259
>TC18097 similar to UP|Q9LNH5 (Q9LNH5) F21D18.5, partial (10%)
Length = 528
Score = 26.9 bits (58), Expect = 4.1
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = -2
Query: 10 QRLHYHSHYSPHF 22
QRL++H HY PHF
Sbjct: 161 QRLYHHLHYFPHF 123
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation
aberrant root formation protein), complete
Length = 3308
Score = 26.9 bits (58), Expect = 4.1
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = -2
Query: 241 GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGD 279
GD+ + G +V + + CKVG K+ EV+ GD
Sbjct: 538 GDVTRELAGEEVVGDVEDLEGGEACKVGGKLVSEVVHGD 422
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,183,530
Number of Sequences: 28460
Number of extensions: 73004
Number of successful extensions: 831
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 748
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 804
length of query: 295
length of database: 4,897,600
effective HSP length: 90
effective length of query: 205
effective length of database: 2,336,200
effective search space: 478921000
effective search space used: 478921000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0225.5


