

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0219.2
(365 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
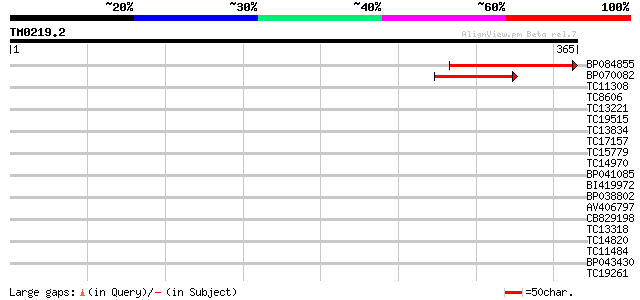
Score E
Sequences producing significant alignments: (bits) Value
BP084855 113 4e-26
BP070082 86 8e-18
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 34 0.034
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 32 0.13
TC13221 similar to UP|PPAN_ARATH (Q9ASU7) Peter Pan-like protein... 32 0.22
TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, pa... 30 0.49
TC13834 homologue to UP|Q948R9 (Q948R9) RPS6-like protein (AT3g1... 30 0.64
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 30 0.84
TC15779 similar to UP|O24093 (O24093) L-ascorbate oxidase precur... 30 0.84
TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) prec... 29 1.1
BP041085 29 1.4
BI419972 29 1.4
BP038802 29 1.4
AV406797 28 1.9
CB829198 28 1.9
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 28 2.4
TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate p... 27 4.1
TC11484 similar to PIR|JC5965|JC5965 cytochrome P450 CYP86A1 - A... 27 4.1
BP043430 27 4.1
TC19261 similar to UP|BAD06577 (BAD06577) Cell wall protein Awa1... 27 4.1
>BP084855
Length = 373
Score = 113 bits (283), Expect = 4e-26
Identities = 58/83 (69%), Positives = 67/83 (79%), Gaps = 1/83 (1%)
Frame = -2
Query: 284 QISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLDLLPQFDDESEPEEDG-EDGNEQHKNE 342
QISVVEP FDLS IG+LKEIKDGQVVGDDD+ +DLLPQF+ ESE EE+ EDG+ + KN
Sbjct: 372 QISVVEPDFDLSCIGFLKEIKDGQVVGDDDVDIDLLPQFNSESEYEEEKCEDGHGEDKNG 193
Query: 343 DHEKEDPQAGTSQGNNANNENLA 365
D + DP+A SQ NNANNENLA
Sbjct: 192 DEGENDPKAAASQDNNANNENLA 124
>BP070082
Length = 489
Score = 86.3 bits (212), Expect = 8e-18
Identities = 39/54 (72%), Positives = 49/54 (90%)
Frame = -1
Query: 274 FAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLDLLPQFDDESE 327
+ +GF+AAKEQ+SVV+P FDLSRIG+LK+I+DGQV+G DDI LDLLP+FD ESE
Sbjct: 345 YVEGFVAAKEQVSVVDPDFDLSRIGFLKKIQDGQVIGHDDIDLDLLPKFDSESE 184
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 34.3 bits (77), Expect = 0.034
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = -2
Query: 310 GDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDP 349
GD+D S + D+E E E + E+G E + ED E EDP
Sbjct: 350 GDEDGSRFHEIENDNEGEEEVEAEEGEEDEEEEDDEDEDP 231
Score = 27.7 bits (60), Expect = 3.2
Identities = 12/55 (21%), Positives = 24/55 (42%)
Frame = -2
Query: 308 VVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNE 362
V+GDD + ++ + E EDG+ H+ E+ + + + +G E
Sbjct: 419 VIGDDGAETTSVDPVEENARAEPGDEDGSRFHEIENDNEGEEEVEAEEGEEDEEE 255
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 32.3 bits (72), Expect = 0.13
Identities = 14/50 (28%), Positives = 27/50 (54%), Gaps = 10/50 (20%)
Frame = -2
Query: 324 DESEPEEDGEDGNEQHKNEDHEKEDP----------QAGTSQGNNANNEN 363
++ + + +GEDGN+ +ED + + P + G GNN+NN++
Sbjct: 641 EDKQHDAEGEDGNDDEGDEDDDGDGPFGEGEEDLSSEDGAGYGNNSNNKS 492
Score = 30.4 bits (67), Expect = 0.49
Identities = 21/53 (39%), Positives = 30/53 (55%), Gaps = 2/53 (3%)
Frame = -2
Query: 301 KEIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEK--EDPQA 351
+E +DG+ DDD + + DD E EE E+G E+ NED E+ ED +A
Sbjct: 425 EEDEDGEDQEDDDDDDEDDDEEDDGGEDEE--EEGVEEEDNEDEEEDEEDEEA 273
Score = 29.6 bits (65), Expect = 0.84
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 2/41 (4%)
Frame = -2
Query: 310 GDDDISLDLLPQFDDESEPEEDGE--DGNEQHKNEDHEKED 348
G+D+ D Q DD+ + E+D E DG E + E E+ED
Sbjct: 434 GEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEED 312
Score = 28.1 bits (61), Expect = 2.4
Identities = 10/36 (27%), Positives = 20/36 (54%)
Frame = -2
Query: 320 PQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
P+ + E E +EDGED + +++ + E+ G +
Sbjct: 446 PEENGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDE 339
>TC13221 similar to UP|PPAN_ARATH (Q9ASU7) Peter Pan-like protein, partial
(15%)
Length = 521
Score = 31.6 bits (70), Expect = 0.22
Identities = 14/36 (38%), Positives = 24/36 (65%)
Frame = +3
Query: 323 DDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNN 358
DDE + EDGED +E ++ED + EDP+ + + ++
Sbjct: 183 DDEMQDSEDGED-SEGSEDEDQDGEDPEEASLEDDD 287
>TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, partial (3%)
Length = 504
Score = 30.4 bits (67), Expect = 0.49
Identities = 18/55 (32%), Positives = 29/55 (52%)
Frame = +3
Query: 311 DDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNENLA 365
DDD D D ESE EED +D ++D + EDP G G +++++++
Sbjct: 57 DDDSDPD-----DAESEDEEDDDD------DDDGDDEDPLLGPGLGRESDSDDMS 188
>TC13834 homologue to UP|Q948R9 (Q948R9) RPS6-like protein
(AT3g17170/K14A17_29), partial (13%)
Length = 517
Score = 30.0 bits (66), Expect = 0.64
Identities = 16/44 (36%), Positives = 26/44 (58%), Gaps = 2/44 (4%)
Frame = +1
Query: 311 DDDISLDLLPQFDDESEPEE--DGEDGNEQHKNEDHEKEDPQAG 352
DDD +L +DD+ + EE DGEDG+ + D +++D +G
Sbjct: 151 DDDDDEELDEDYDDDWDGEEDADGEDGDAEFIVIDDDQDDVDSG 282
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 29.6 bits (65), Expect = 0.84
Identities = 10/27 (37%), Positives = 19/27 (70%)
Frame = +2
Query: 322 FDDESEPEEDGEDGNEQHKNEDHEKED 348
F+D E +EDG++ ++ ++D E+ED
Sbjct: 761 FEDIEEDDEDGDEDEDEDDDDDEEEED 841
Score = 26.6 bits (57), Expect = 7.1
Identities = 13/52 (25%), Positives = 26/52 (50%)
Frame = +2
Query: 302 EIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGT 353
E D + + +DD D DD+ + EE+ +D +++ E K ++G+
Sbjct: 749 EQSDFEDIEEDDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSGS 904
Score = 26.2 bits (56), Expect = 9.2
Identities = 15/62 (24%), Positives = 27/62 (43%)
Frame = +2
Query: 301 KEIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNAN 360
++I++ GD+D D DD+ E E+D +D + + E K + G +
Sbjct: 764 EDIEEDDEDGDEDEDED----DDDDEEEEDDDDDDEDDEEGEGKSKSKSGSKARPGKDQP 931
Query: 361 NE 362
E
Sbjct: 932 TE 937
>TC15779 similar to UP|O24093 (O24093) L-ascorbate oxidase precursor, partial
(50%)
Length = 1304
Score = 29.6 bits (65), Expect = 0.84
Identities = 18/71 (25%), Positives = 31/71 (43%), Gaps = 3/71 (4%)
Frame = +1
Query: 62 PLHQSTLDPKGRPTEKKKGHDNVPPHQPDSGALINR---PSTPFIQAGPSSAIGGEALPP 118
P H+ P+G + + +N PPH+P + + PS+P P+ +G
Sbjct: 988 PHHRPEPKPQGTARQSESEAENPPPHKPSPSSTTKQSPAPSSPRTPPPPTPRMGRFRTQL 1167
Query: 119 LLNLSDPRFNG 129
L+ + R NG
Sbjct: 1168 SLHQENHRGNG 1200
>TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) precursor,
partial (51%)
Length = 1566
Score = 29.3 bits (64), Expect = 1.1
Identities = 19/63 (30%), Positives = 29/63 (45%), Gaps = 19/63 (30%)
Frame = +1
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQH-------------------KNEDHEKEDP 349
V DD+S + P+ DE E EED ++ EQH ++ED ++E+
Sbjct: 463 VEGDDLSF-ISPESADEDEDEEDEDEDQEQHGRHSRPSGRRTRRPSREEDEDEDEDEEER 639
Query: 350 QAG 352
Q G
Sbjct: 640 QGG 648
>BP041085
Length = 449
Score = 28.9 bits (63), Expect = 1.4
Identities = 21/60 (35%), Positives = 29/60 (48%), Gaps = 5/60 (8%)
Frame = +1
Query: 3 KSKVDANPIKMK-----EYLAQSAAAAKKRAAETEQKKKNEGTSGSDNVRDPKRQKTSSA 57
+S + A+ IKM +YL Q A++ AA + K N GTS S N + Q S A
Sbjct: 7 QSNLGAS*IKMAGLSRIQYLRQQLLEAEEAAAREMEPKSNHGTSNSFNNQGGGGQNFSGA 186
>BI419972
Length = 498
Score = 28.9 bits (63), Expect = 1.4
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 13/106 (12%)
Frame = +1
Query: 271 SLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEI-----------KDGQVVGDDDISLD-- 317
SLL A +AA IS V+ L+ IG+L + KDG+ +D +
Sbjct: 166 SLLHA---VAADTLISAVQ---SLTLIGYLLTMITDLVPPKLNSKDGRFSLEDLLGRKPA 327
Query: 318 LLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNEN 363
L D E+E +EDG+D ++ ++D ++ + +G +G A+ E+
Sbjct: 328 YLANKDSETEDDEDGDDEDDDADDDDDDEGEDYSG-DEGEEADPED 462
>BP038802
Length = 533
Score = 28.9 bits (63), Expect = 1.4
Identities = 16/46 (34%), Positives = 24/46 (51%)
Frame = +1
Query: 311 DDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQG 356
DDD D P FD E ++ EDG +Q+ N H +E+ + +G
Sbjct: 298 DDDDGGDDGPFFDLEFTVPDESEDG-QQNPNHTHPEEEEECDEGEG 432
>AV406797
Length = 418
Score = 28.5 bits (62), Expect = 1.9
Identities = 19/63 (30%), Positives = 28/63 (44%), Gaps = 3/63 (4%)
Frame = +1
Query: 44 DNVRDPKRQKTSSAAGV---KPLHQSTLDPKGRPTEKKKGHDNVPPHQPDSGALINRPST 100
D +R+ + +AGV K +S G P+ +K PP QP S N+P
Sbjct: 163 DMIREEGKNMEKPSAGVVGRKTYSRSVSWTDGSPSRRKPP----PPSQPPSSNSSNKPPR 330
Query: 101 PFI 103
PF+
Sbjct: 331 PFL 339
>CB829198
Length = 562
Score = 28.5 bits (62), Expect = 1.9
Identities = 13/42 (30%), Positives = 23/42 (53%)
Frame = +3
Query: 321 QFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNE 362
Q DD+ EP+ D E+ ++ D ++E + G+N NN+
Sbjct: 240 QHDDQEEPQNDDV---EESQHSDSKEETEEEEDEDGDNENNK 356
Score = 26.9 bits (58), Expect = 5.4
Identities = 13/52 (25%), Positives = 25/52 (48%)
Frame = +3
Query: 311 DDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNE 362
+DD+ +E+E EED + NE +K + + + + N ++NE
Sbjct: 267 NDDVEESQHSDSKEETEEEEDEDGDNENNKPKRAKAAKKKKNKERSNKSDNE 422
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 28.1 bits (61), Expect = 2.4
Identities = 11/40 (27%), Positives = 21/40 (52%)
Frame = +3
Query: 323 DDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNE 362
DDE E +ED +D ++ ++D E+E+ + + E
Sbjct: 267 DDEEEDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEGSEE 386
Score = 27.3 bits (59), Expect = 4.1
Identities = 13/44 (29%), Positives = 25/44 (56%)
Frame = +3
Query: 305 DGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKED 348
D + DDD D + DD+ ++D +DG E+ ++D ++E+
Sbjct: 246 DNEEEEDDDEEEDDEDEDDDDD--DDDDDDGEEEEDDDDDDEEE 371
Score = 26.9 bits (58), Expect = 5.4
Identities = 15/51 (29%), Positives = 26/51 (50%)
Frame = +3
Query: 302 EIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAG 352
E +D + DDD D DD+ E EED +D +E+ + + + + + G
Sbjct: 273 EEEDDEDEDDDDDDDD-----DDDGEEEEDDDDDDEEEEGSEEVEVEGKDG 410
>TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate protein 80,
partial (53%)
Length = 1180
Score = 27.3 bits (59), Expect = 4.1
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +1
Query: 323 DDESEPEEDGEDGNEQHKNEDHEKED 348
DDE E E+ E+G + +E+ EKE+
Sbjct: 733 DDEDEEEKKDEEGKVEDVDEEKEKEE 810
>TC11484 similar to PIR|JC5965|JC5965 cytochrome P450 CYP86A1 - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (43%)
Length = 752
Score = 27.3 bits (59), Expect = 4.1
Identities = 19/60 (31%), Positives = 28/60 (46%), Gaps = 3/60 (5%)
Frame = +2
Query: 45 NVRDPKRQKTSSAAGVKPLHQSTLDPKGRP--TEKKKGHDNVPPHQPD-SGALINRPSTP 101
+ R P +KTSS + + L + DP GRP T + +P S A + R S+P
Sbjct: 281 STRSPATRKTSSTSSERGLTTTRKDPGGRPPSTTSSATESSTATAKPGLSSAKLPRSSSP 460
>BP043430
Length = 493
Score = 27.3 bits (59), Expect = 4.1
Identities = 13/39 (33%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Frame = -1
Query: 311 DDDISLDLLPQFDDESEPEEDGEDGNEQHK-NEDHEKED 348
DD+ + + + + D+E E D +DG+E + N+D E D
Sbjct: 349 DDEGNDNEVDEEDEEEEDSNDDDDGSESEEFNDDEEDSD 233
>TC19261 similar to UP|BAD06577 (BAD06577) Cell wall protein Awa1p, partial
(3%)
Length = 495
Score = 27.3 bits (59), Expect = 4.1
Identities = 18/65 (27%), Positives = 27/65 (40%), Gaps = 6/65 (9%)
Frame = +1
Query: 21 AAAAKKRAAETEQKKKNEGTSGSDNVRDPKRQKTSSAAGVKPLHQ------STLDPKGRP 74
AAAA + A + + NEGT + + SS P+H + +PKG
Sbjct: 229 AAAAPRIAEKASSRGTNEGTGNEGGSKRVHQPDPSSILIASPIHSRRGHGLQSKEPKGLA 408
Query: 75 TEKKK 79
T + K
Sbjct: 409 TRRLK 423
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.308 0.127 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,907,499
Number of Sequences: 28460
Number of extensions: 61311
Number of successful extensions: 424
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 405
length of query: 365
length of database: 4,897,600
effective HSP length: 91
effective length of query: 274
effective length of database: 2,307,740
effective search space: 632320760
effective search space used: 632320760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0219.2


