

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0218.20
(631 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
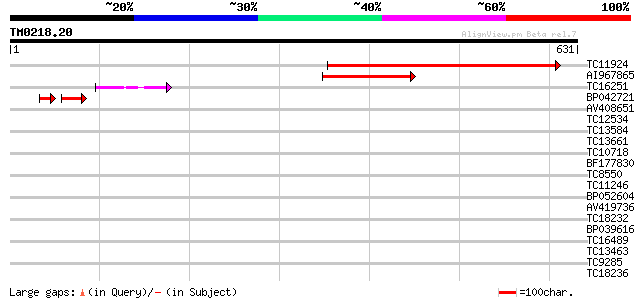
Score E
Sequences producing significant alignments: (bits) Value
TC11924 similar to UP|P93484 (P93484) BP-80 vacuolar sorting rec... 558 e-160
AI967865 170 6e-43
TC16251 similar to UP|Q945B9 (Q945B9) Growth-on protein GRO11, p... 53 2e-07
BP042721 44 2e-07
AV408651 33 0.18
TC12534 similar to UP|Q9MAJ1 (Q9MAJ1) F27F5.26, partial (13%) 30 1.5
TC13584 30 1.5
TC13661 similar to UP|Q7VRW0 (Q7VRW0) NADH dehydrogenase I chain... 30 1.5
TC10718 similar to UP|Q9FXS6 (Q9FXS6) Nt-SubE80 protein, partial... 29 2.6
BF177830 29 2.6
TC8550 similar to UP|Q94IP5 (Q94IP5) Mono-lipoyl E2, partial (21%) 28 3.4
TC11246 28 3.4
BP052604 28 3.4
AV419736 28 3.4
TC18232 similar to UP|Q7PS48 (Q7PS48) ENSANGP00000015589 (Fragme... 28 4.5
BP039616 28 5.8
TC16489 similar to UP|O64885 (O64885) Expressed protein (At2g445... 28 5.8
TC13463 28 5.8
TC9285 weakly similar to PIR|T50669|T50669 villin 2 [imported] -... 27 9.9
TC18236 weakly similar to UP|Q9LH79 (Q9LH79) MtN3-like protein (... 27 9.9
>TC11924 similar to UP|P93484 (P93484) BP-80 vacuolar sorting receptor,
partial (41%)
Length = 783
Score = 558 bits (1439), Expect = e-160
Identities = 259/260 (99%), Positives = 260/260 (99%)
Frame = +3
Query: 354 NPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCL 413
+PVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCL
Sbjct: 3 DPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCL 182
Query: 414 SSDVETNECLENNGGCWKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPG 473
SSDVETNECLENNGGCWKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPG
Sbjct: 183 SSDVETNECLENNGGCWKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPG 362
Query: 474 RCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPE 533
RCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPE
Sbjct: 363 RCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPE 542
Query: 534 CSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLI 593
CSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLI
Sbjct: 543 CSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLI 722
Query: 594 YKYRIRSYMDSEIRAIMAQY 613
YKYRIRSYMDSEIRAIMAQY
Sbjct: 723 YKYRIRSYMDSEIRAIMAQY 782
>AI967865
Length = 326
Score = 170 bits (431), Expect = 6e-43
Identities = 76/103 (73%), Positives = 90/103 (86%)
Frame = +2
Query: 349 DADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTE 408
+AD +N VLK EQ QVG+GSRGDVTILPTLV+NN QY GKLE+ V+KA+C+GF+E+TE
Sbjct: 17 EADVENEVLKNEQQVQVGRGSRGDVTILPTLVINNVQYTGKLERTPVLKAVCAGFKESTE 196
Query: 409 PAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRGRVCEC 451
P VCLS D+ETNECLE NGGCW+DK A+I+ACKDTFRGRVCEC
Sbjct: 197 PPVCLSGDIETNECLERNGGCWQDKNASISACKDTFRGRVCEC 325
>TC16251 similar to UP|Q945B9 (Q945B9) Growth-on protein GRO11, partial (9%)
Length = 905
Score = 52.8 bits (125), Expect = 2e-07
Identities = 35/85 (41%), Positives = 43/85 (50%)
Frame = +3
Query: 96 TIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPS 155
++ L RG C F K AQ GASA+L+ +D EE L M G NI+IP
Sbjct: 303 SVALCTRGGCDFTTKAAFAQSGGASAMLLIND-EEDLFEMACSNSTGG------NISIPV 461
Query: 156 ALIEKSFGEKLKKSISGGEMVNVNL 180
LI KS GE KS++ G V V L
Sbjct: 462 VLITKSAGEAFNKSLASGRKVEVLL 536
>BP042721
Length = 504
Score = 43.5 bits (101), Expect(2) = 2e-07
Identities = 19/28 (67%), Positives = 22/28 (77%)
Frame = +2
Query: 58 QYGGSMAGNVVYPKDNHKGCKEFDESGI 85
Q GG +A N VYPK+ H+GCKEFDE GI
Sbjct: 278 QCGGCVARNGVYPKEIHQGCKEFDEYGI 361
Score = 28.5 bits (62), Expect(2) = 2e-07
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +1
Query: 34 SLTVTSPEDIKGTHDSAI 51
+L + SP+ IKGTHDSAI
Sbjct: 217 ALPMISPQTIKGTHDSAI 270
>AV408651
Length = 405
Score = 32.7 bits (73), Expect = 0.18
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = +1
Query: 520 VDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRD 556
+++C E C+ C C+N W DCS + IRD
Sbjct: 67 INQCSEHGHCRGGFCQCENGWYGADCSTPSVISSIRD 177
>TC12534 similar to UP|Q9MAJ1 (Q9MAJ1) F27F5.26, partial (13%)
Length = 941
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/53 (30%), Positives = 22/53 (41%), Gaps = 7/53 (13%)
Frame = +1
Query: 503 CQCPAGFKGDGVKNCQDVDECKEKKACQC-------PECSCKNTWGSYDCSCS 548
C+C +K K+ C K + +C CSCKN+W C CS
Sbjct: 1 CKCKHSWKCCSFKHSWKC--CSFKHSWKCCFFKYSWKHCSCKNSWKHCSCKCS 153
>TC13584
Length = 596
Score = 29.6 bits (65), Expect = 1.5
Identities = 14/21 (66%), Positives = 16/21 (75%)
Frame = -1
Query: 29 VVEKNSLTVTSPEDIKGTHDS 49
VVE LTVTSP++I THDS
Sbjct: 119 VVEFTRLTVTSPKNINRTHDS 57
>TC13661 similar to UP|Q7VRW0 (Q7VRW0) NADH dehydrogenase I chain J ,
partial (7%)
Length = 559
Score = 29.6 bits (65), Expect = 1.5
Identities = 13/41 (31%), Positives = 19/41 (45%), Gaps = 9/41 (21%)
Frame = -3
Query: 531 CPECSCKNTWGSYDCSCSGDLLYIR---------DHDTCIS 562
C +CKN W S C G+ +I+ D DTC++
Sbjct: 197 CIRIACKNCWSSRTAKCVGNTSFIKDRFQLGNIPDRDTCMT 75
>TC10718 similar to UP|Q9FXS6 (Q9FXS6) Nt-SubE80 protein, partial (74%)
Length = 557
Score = 28.9 bits (63), Expect = 2.6
Identities = 13/21 (61%), Positives = 18/21 (84%)
Frame = +1
Query: 8 LGFLLGFLLLLSVTPSSMARF 28
+G LLGFLL+LS++ SS+A F
Sbjct: 355 IGALLGFLLMLSLSHSSLASF 417
>BF177830
Length = 476
Score = 28.9 bits (63), Expect = 2.6
Identities = 18/58 (31%), Positives = 30/58 (51%), Gaps = 2/58 (3%)
Frame = +2
Query: 514 VKNCQDVDECKEKKACQC--PECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGG 569
V++ D + +E+ C+ C +NT+ +CSC GDL + H+ C+ K S G
Sbjct: 188 VESAADEEIPEEEAVCRICFDVCDERNTF-KMECSCKGDLTLV--HEECLIKWFSLKG 352
>TC8550 similar to UP|Q94IP5 (Q94IP5) Mono-lipoyl E2, partial (21%)
Length = 852
Score = 28.5 bits (62), Expect = 3.4
Identities = 17/64 (26%), Positives = 24/64 (36%), Gaps = 8/64 (12%)
Frame = -3
Query: 198 WTNSNDECGVKCDMLMEFVKDFKGAAQILEK--------GGYTQFTPHYITWYCPKAFTL 249
W N+ +KC + +K+FK A I + G P + TW P T
Sbjct: 586 WKNTISGFKIKCRIYPRLLKNFKSAKFIFQGLKTSIKNWSGIQSINPKWETWKYPDEQTS 407
Query: 250 SKQC 253
S C
Sbjct: 406 SSAC 395
>TC11246
Length = 563
Score = 28.5 bits (62), Expect = 3.4
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 3/37 (8%)
Frame = -3
Query: 523 CKEKKACQCPECSCKN---TWGSYDCSCSGDLLYIRD 556
C+ +AC CSC T Y C+CS +L + RD
Sbjct: 327 CR*MRACWGRSCSCDTSAKTSTPYGCNCSCELQFSRD 217
>BP052604
Length = 506
Score = 28.5 bits (62), Expect = 3.4
Identities = 20/53 (37%), Positives = 24/53 (44%), Gaps = 8/53 (15%)
Frame = -1
Query: 506 PAGFKGDGVKNCQDVDE----CKEKKACQCPECSCKNT----WGSYDCSCSGD 550
P + D + +C VDE C E Q P C C N WG Y CSC+ D
Sbjct: 302 PEFYSIDCLHDCLAVDEEPFICLECLFSQVPLCLCCNCMCLYWGLY-CSCNCD 147
>AV419736
Length = 427
Score = 28.5 bits (62), Expect = 3.4
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +1
Query: 464 YTTCEASGPGRCKIKNGGCWH 484
+ +C+A G RC N GCWH
Sbjct: 358 FWSCKAIG**RCPCYNSGCWH 420
>TC18232 similar to UP|Q7PS48 (Q7PS48) ENSANGP00000015589 (Fragment),
partial (8%)
Length = 907
Score = 28.1 bits (61), Expect = 4.5
Identities = 14/35 (40%), Positives = 16/35 (45%)
Frame = +1
Query: 511 GDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDC 545
G+GV C D C + C CK WGSY C
Sbjct: 187 GEGV-TC*DCYSCMP*PSGPC*SSCCKGHWGSYCC 288
>BP039616
Length = 564
Score = 27.7 bits (60), Expect = 5.8
Identities = 15/40 (37%), Positives = 17/40 (42%)
Frame = +3
Query: 471 GPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFK 510
G G C + NGGCW A FS D C GF+
Sbjct: 351 GTGHCSL-NGGCWLGAHTRFGFSCKFSDLVHLCSGIMGFQ 467
>TC16489 similar to UP|O64885 (O64885) Expressed protein
(At2g44510/F4I1.32), partial (29%)
Length = 709
Score = 27.7 bits (60), Expect = 5.8
Identities = 16/44 (36%), Positives = 19/44 (42%), Gaps = 2/44 (4%)
Frame = -1
Query: 537 KNTWGSYDCSCSGDLLYIRDHD-TCISKTASQGGRSA-WAAFWV 578
K T G SC L Y +DH+ CI K G + W WV
Sbjct: 472 KTTGGKSSISCQ*SLQYKKDHNKICIKKFPKFGSSNTFWLPLWV 341
>TC13463
Length = 601
Score = 27.7 bits (60), Expect = 5.8
Identities = 14/47 (29%), Positives = 20/47 (41%), Gaps = 2/47 (4%)
Frame = +2
Query: 509 FKGDGVKNCQDVDE--CKEKKACQCPECSCKNTWGSYDCSCSGDLLY 553
+K DG C + C + C SCK+ G+ DCS L +
Sbjct: 155 WKRDGCSKCSSNSKAVCLNNQDCALQTSSCKSHGGTVDCSLGIQLAF 295
>TC9285 weakly similar to PIR|T50669|T50669 villin 2 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(7%)
Length = 657
Score = 26.9 bits (58), Expect = 9.9
Identities = 10/27 (37%), Positives = 15/27 (55%)
Frame = -1
Query: 237 HYITWYCPKAFTLSKQCKSQCINHGRY 263
H TW A ++ + K+ CI+HG Y
Sbjct: 390 HIQTWILTTAMSIMAESKTLCIDHGIY 310
>TC18236 weakly similar to UP|Q9LH79 (Q9LH79) MtN3-like protein
(AT3g14770/T21E2_2), partial (42%)
Length = 1039
Score = 26.9 bits (58), Expect = 9.9
Identities = 13/32 (40%), Positives = 15/32 (46%)
Frame = +3
Query: 526 KKACQCPECSCKNTWGSYDCSCSGDLLYIRDH 557
KK Q EC W ++C CS LL DH
Sbjct: 81 KKKVQGHECPSFCVWHIWECFCSVPLLGTSDH 176
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,356,123
Number of Sequences: 28460
Number of extensions: 199220
Number of successful extensions: 1018
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 1003
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1015
length of query: 631
length of database: 4,897,600
effective HSP length: 96
effective length of query: 535
effective length of database: 2,165,440
effective search space: 1158510400
effective search space used: 1158510400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0218.20


