

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
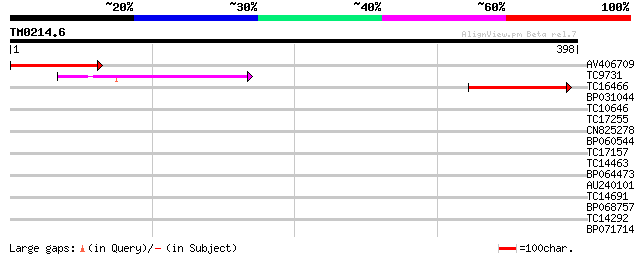
Score E
Sequences producing significant alignments: (bits) Value
AV406709 139 1e-33
TC9731 80 6e-16
TC16466 weakly similar to UP|TDE2_MOUSE (Q9QZI8) Tumor different... 79 1e-15
BP031044 31 0.41
TC10646 weakly similar to UP|AAS55083 (AAS55083) UDP-glucose glu... 28 2.7
TC17255 28 2.7
CN825278 28 3.5
BP060544 27 4.6
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 27 4.6
TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein (At2g... 27 4.6
BP064473 27 4.6
AU240101 27 6.0
TC14691 27 6.0
BP068757 27 7.8
TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precur... 27 7.8
BP071714 27 7.8
>AV406709
Length = 430
Score = 139 bits (349), Expect = 1e-33
Identities = 65/65 (100%), Positives = 65/65 (100%)
Frame = +1
Query: 1 METGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRGAL 60
METGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRGAL
Sbjct: 235 METGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRGAL 414
Query: 61 TEMER 65
TEMER
Sbjct: 415 TEMER 429
>TC9731
Length = 662
Score = 80.1 bits (196), Expect = 6e-16
Identities = 46/142 (32%), Positives = 73/142 (51%), Gaps = 5/142 (3%)
Frame = +1
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGK-----DCLGAEGVLRVSLGC 88
AR Y +F + L+A R+ A ME + N K + + VLRVSLG
Sbjct: 220 ARIAYCGLFAFSLLVAGILREV---AAPLMESIPWINHFKHTPSREWFETDAVLRVSLGN 390
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+ I+ + + RD H G W +KI+ W + + F LP+E I Y ++ F
Sbjct: 391 FLFFTILAVLMVGVKNQKDPRDGLHHGGWMMKIICWFLLVIFMFFLPNEIISFYETISKF 570
Query: 149 GAGVFLLIQLISIISFITWLND 170
G+G+FLL+Q++ ++ F+ ND
Sbjct: 571 GSGMFLLVQVVLLLDFVHGWND 636
>TC16466 weakly similar to UP|TDE2_MOUSE (Q9QZI8) Tumor differentially
expressed protein 2 (Tumor differentially expressed 1
protein like) (Membrane protein TMS-2) (Axotomy induced
glyco/Golgi protein 2), partial (6%)
Length = 584
Score = 79.3 bits (194), Expect = 1e-15
Identities = 34/72 (47%), Positives = 45/72 (62%)
Frame = +1
Query: 323 VPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLW 382
V Y Y FFH +F+ +MY AMLL GW++ +DVGW S WVRIV W ++LW
Sbjct: 139 VTYSYSFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIVTCWATAILFLW 318
Query: 383 MLVAPIIWKSRQ 394
LVAPI++ R+
Sbjct: 319 SLVAPIMFPERE 354
>BP031044
Length = 381
Score = 30.8 bits (68), Expect = 0.41
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = -3
Query: 211 PSCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILT 245
PSC + FF+ L+L QL S S P++ AG+LT
Sbjct: 256 PSCHIPSFFVLMNLLLQQL*MSFSNLPELVAGVLT 152
>TC10646 weakly similar to UP|AAS55083 (AAS55083) UDP-glucose
glucosyltransferase, partial (8%)
Length = 501
Score = 28.1 bits (61), Expect = 2.7
Identities = 10/34 (29%), Positives = 19/34 (55%)
Frame = +3
Query: 56 GRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCF 89
GR + + + + C GG+D + ++ +SL CF
Sbjct: 162 GRSRVIQKQSTQTCTGGQDSRARQRIISLSLDCF 263
>TC17255
Length = 485
Score = 28.1 bits (61), Expect = 2.7
Identities = 14/54 (25%), Positives = 26/54 (47%), Gaps = 1/54 (1%)
Frame = -3
Query: 115 GWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGV-FLLIQLISIISFITW 167
GWW K++ W+ + + SEF I + + G+ F ++ + +S I W
Sbjct: 216 GWWREKVIQWISRFWLVLICFSEFSIICSDHSINVHGIKFHWLRKLRCVSMIAW 55
>CN825278
Length = 659
Score = 27.7 bits (60), Expect = 3.5
Identities = 13/36 (36%), Positives = 20/36 (55%)
Frame = +2
Query: 98 TTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFL 133
T+A+ V D W G + ++VL+ TV+PFL
Sbjct: 305 TSAQKDSAWMVADPWRCGKFEQRLVLYPKRTVLPFL 412
>BP060544
Length = 446
Score = 27.3 bits (59), Expect = 4.6
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = -2
Query: 197 LVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTS 232
LVG + I P CL F + W L+L +L++S
Sbjct: 442 LVGFWIHTIG*RPTVFCLFQCFLLVWLLLLFKLLSS 335
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein 1-like
protein 3, partial (81%)
Length = 1298
Score = 27.3 bits (59), Expect = 4.6
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +3
Query: 317 IPAEDDVPYGYGFFHFVFATGAMYFAMLLI 346
+P + D YGFFHF F YF+ML++
Sbjct: 1131 LPCDVDFIPLYGFFHFFFIES--YFSMLIL 1214
>TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein
(At2g16660/T24I21.7), partial (40%)
Length = 1122
Score = 27.3 bits (59), Expect = 4.6
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = +1
Query: 188 FATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLL 227
F V +V + L+ + P PS L++ F+A LVLL
Sbjct: 748 FGVCNAVAVVVAVYLLAYGFVPSPSSLVSRVFVAVLLVLL 867
>BP064473
Length = 471
Score = 27.3 bits (59), Expect = 4.6
Identities = 13/26 (50%), Positives = 14/26 (53%)
Frame = -3
Query: 201 ILMFIWYAPKPSCLLNIFFIAWTLVL 226
IL+FIW LL I F WTL L
Sbjct: 193 ILLFIWNQDFLKTLLTILFSVWTLFL 116
>AU240101
Length = 300
Score = 26.9 bits (58), Expect = 6.0
Identities = 16/50 (32%), Positives = 24/50 (48%)
Frame = -3
Query: 158 LISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWY 207
L++II + LN S Y A Q H F ++VCL I + +W+
Sbjct: 145 LVTIILLVIRLNS--SSSHYEAHHQQHCNQFIARHFLVCLTLSINICVWF 2
>TC14691
Length = 1255
Score = 26.9 bits (58), Expect = 6.0
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -2
Query: 123 LWVGMTVIPFLLPSEFIQI 141
LWV V+P LP+EF++I
Sbjct: 321 LWVSEEVLPVSLPAEFVEI 265
>BP068757
Length = 519
Score = 26.6 bits (57), Expect = 7.8
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = -2
Query: 110 DTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGA 150
D W WW V+ L ++ P+LLP + I +Y + H G+
Sbjct: 170 DDW---WWGVEFWLSD**SIQPYLLPLDMIVLY--IKHIGS 63
>TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (95%)
Length = 1792
Score = 26.6 bits (57), Expect = 7.8
Identities = 18/97 (18%), Positives = 37/97 (37%), Gaps = 32/97 (32%)
Frame = +3
Query: 303 GIDSKCFQFRKD---------DDIPAEDDVP-----------------------YGYGFF 330
G+D +C Q+RK+ +D+P D++ Y G F
Sbjct: 726 GVDGRCDQYRKNAKVVSIDDYEDVPTYDEIALKKAVANQPISVAIEGGGREFQLYDSGIF 905
Query: 331 HFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTW 367
T A+ ++ +G+ + + + W + W ++W
Sbjct: 906 TGRCGT-ALDHGVVAVGYGTENGLDYWIVRNSWGASW 1013
>BP071714
Length = 411
Score = 26.6 bits (57), Expect = 7.8
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = +1
Query: 197 LVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLH 236
LV LMFIW+ + CLL + + + L+ MTS+ +H
Sbjct: 109 LVRSTLMFIWFLQEFPCLL*VL----SQMHLECMTSIFVH 216
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,136,848
Number of Sequences: 28460
Number of extensions: 150800
Number of successful extensions: 1161
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1152
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1160
length of query: 398
length of database: 4,897,600
effective HSP length: 92
effective length of query: 306
effective length of database: 2,279,280
effective search space: 697459680
effective search space used: 697459680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0214.6


