

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.7
(451 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
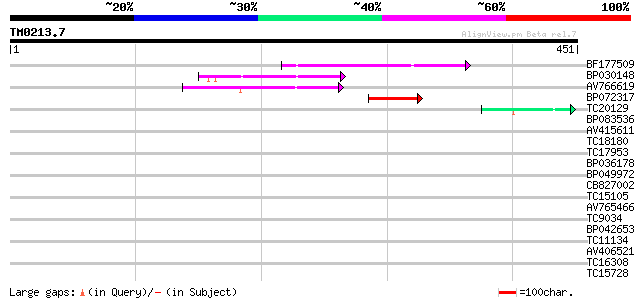
Score E
Sequences producing significant alignments: (bits) Value
BF177509 70 7e-13
BP030148 52 2e-07
AV766619 43 1e-04
BP072317 42 2e-04
TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-li... 41 5e-04
BP083536 40 0.001
AV415611 39 0.002
TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutat... 37 0.005
TC17953 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17... 37 0.009
BP036178 36 0.011
BP049972 35 0.019
CB827002 30 1.1
TC15105 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein... 30 1.1
AV765466 29 1.4
TC9034 29 1.8
BP042653 28 2.4
TC11134 28 2.4
AV406521 27 5.3
TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I... 27 6.9
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 27 6.9
>BF177509
Length = 472
Score = 70.1 bits (170), Expect = 7e-13
Identities = 42/150 (28%), Positives = 70/150 (46%)
Frame = +1
Query: 217 WLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFA 276
W+ +IP R W A+ + L N+ ES N IL P++ M++ IR +M F
Sbjct: 19 WIRRIPPRLWAT-AYFEGQRFGHLTANIVESLNTWILEASGLPIIQMMECIRRQLMTWFN 195
Query: 277 TMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHT 336
E ++ ++P + + +E R + R E ++EV + + +V++
Sbjct: 196 ERRETSMQWASILVPSAERCVAEALERARTYQVLRANEAEFEV--ISHEGSNIVDIRNRC 369
Query: 337 CSCNFWELVGIPCRHAVAAISHSSQRPEDF 366
C C W+L G+PC HAVAA+ Q F
Sbjct: 370 CLCRGWQLYGLPCAHAVAALLSCRQNVHRF 459
>BP030148
Length = 398
Score = 52.4 bits (124), Expect = 2e-07
Identities = 37/128 (28%), Positives = 64/128 (49%), Gaps = 11/128 (8%)
Frame = +3
Query: 151 LTVFDD----LMGGVE-------HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQL 199
LT+ D ++ GVE H FC+RHL +F+K+F +++ NL+ AA
Sbjct: 21 LTILSDRQKGIVDGVEANFPTAFHGFCMRHLSDSFRKEFNNTMLV-NLLWEAAHVLTVIE 197
Query: 200 WTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKP 259
+ K++E++ + +A W+ +IP R W A+ + L N+ ES + IL P
Sbjct: 198 FESKILEIEEISQDAAYWIRRIPPRLWAT-AYFEGQRFGHLTANIVESLDTWILDASGLP 374
Query: 260 VLTMVDWI 267
++ M + I
Sbjct: 375 IVQMTECI 398
>AV766619
Length = 412
Score = 42.7 bits (99), Expect = 1e-04
Identities = 32/134 (23%), Positives = 55/134 (40%), Gaps = 6/134 (4%)
Frame = +2
Query: 138 EPIYEVS*ILQGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGV------VIRNLMMAA 191
EPI V+ GL ++ H +CLR+L K G + N AA
Sbjct: 14 EPITFVANCQNGLKKSLSEIFEKCHHSYCLRYLAEKLNKDLKGQFSHEARRFMINDFYAA 193
Query: 192 AKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNAT 251
A A + + + ++ + EA+ W+MQ W +AF + + L +N + F
Sbjct: 194 AYAPKQETFERSVENMKGISSEAFNWVMQNEPEHWA-NAFFNGARYNHLTSNFGQQFYNW 370
Query: 252 ILLVRDKPVLTMVD 265
+ + P+ M+D
Sbjct: 371 MSEAHELPITQMID 412
>BP072317
Length = 436
Score = 42.0 bits (97), Expect = 2e-04
Identities = 20/43 (46%), Positives = 28/43 (64%)
Frame = +3
Query: 286 KGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQF 328
+G+VMPKPLKRL E++ + GE YEVKH+V+ + F
Sbjct: 306 QGQVMPKPLKRLAWEVKGTSNGLPQASGEGLYEVKHVVTLEGF 434
>TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (12%)
Length = 588
Score = 40.8 bits (94), Expect = 5e-04
Identities = 27/85 (31%), Positives = 33/85 (38%), Gaps = 10/85 (11%)
Frame = +3
Query: 376 YMATYGHVITPINGQKLWPTTNDP----------YILPPLYKRAPGRPKKLRRRDPHEDT 425
Y TY I PI + LW ++ I PP R PGRP+K R R
Sbjct: 27 YRKTYSQTIHPIPDKSLWKELSEGDENAAKADQVLINPPKSLRPPGRPRKKRVRAEDRGR 206
Query: 426 TDNQTRLRSRCGQFGHNSRSCRNPV 450
SRC Q GH +C P+
Sbjct: 207 VKRVVHC-SRCNQTGHFRTTCAAPI 278
>BP083536
Length = 377
Score = 39.7 bits (91), Expect = 0.001
Identities = 14/22 (63%), Positives = 17/22 (76%)
Frame = +1
Query: 337 CSCNFWELVGIPCRHAVAAISH 358
C CN WELV IPCRHA+A + +
Sbjct: 307 CCCNLWELVEIPCRHALAGVHY 372
>AV415611
Length = 219
Score = 38.5 bits (88), Expect = 0.002
Identities = 14/28 (50%), Positives = 19/28 (67%)
Frame = +3
Query: 329 VVNLAEHTCSCNFWELVGIPCRHAVAAI 356
VV++ CSC W+L G+PC HA+A I
Sbjct: 108 VVDIDRWECSCKTWQLTGVPCCHAIAVI 191
>TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (4%)
Length = 513
Score = 37.4 bits (85), Expect = 0.005
Identities = 25/87 (28%), Positives = 38/87 (42%), Gaps = 4/87 (4%)
Frame = +3
Query: 365 DFVHPYYRREAYMATYGHVITPINGQKLWPTTNDPY----ILPPLYKRAPGRPKKLRRRD 420
D+ Y E+Y TY + PI + P + D + PP +R PGRP +R
Sbjct: 12 DYCSRYCTTESYRLTYSERVNPIPIMAV-PESKDSQPVVTVTPPFTRRPPGRP--ATKRY 182
Query: 421 PHEDTTDNQTRLRSRCGQFGHNSRSCR 447
++ + SRC GHN +C+
Sbjct: 183 VSQNIVKRELHC-SRCKGLGHNKSTCK 260
>TC17953 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17A9_9
{Arabidopsis thaliana;}, partial (6%)
Length = 527
Score = 36.6 bits (83), Expect = 0.009
Identities = 19/49 (38%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Frame = +3
Query: 401 ILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR-SRCGQFGHNSRSCRN 448
+ PP KR PGRPK P E + +L+ S+C + GHN ++C+N
Sbjct: 36 VTPPPTKRPPGRPKM----KPVESIDMIKRQLQCSKCKELGHNKKTCKN 170
>BP036178
Length = 561
Score = 36.2 bits (82), Expect = 0.011
Identities = 24/86 (27%), Positives = 32/86 (36%), Gaps = 11/86 (12%)
Frame = -2
Query: 376 YMATYGHVITPINGQKLWPTTN-----------DPYILPPLYKRAPGRPKKLRRRDPHED 424
Y Y +I PI + W D I PP +R PGRPKK R +
Sbjct: 560 YRMAYSQIIIPIPDRSQWREHGEGAEGVGGARVDIVIRPPKTRRPPGRPKKKVLRVENFK 381
Query: 425 TTDNQTRLRSRCGQFGHNSRSCRNPV 450
+ RC GH+ + C P+
Sbjct: 380 RPKRIVQC-GRCHMLGHSQKKCTMPI 306
>BP049972
Length = 496
Score = 35.4 bits (80), Expect = 0.019
Identities = 27/104 (25%), Positives = 34/104 (31%), Gaps = 19/104 (18%)
Frame = -1
Query: 366 FVHPYYRREAYMATYGHVITPINGQKLWPTTN-------------------DPYILPPLY 406
F + Y TY I PI + W + + I PP
Sbjct: 490 FTESCFTVATYRKTYSQTIHPIPDRSFWNELSIEDANANKAVEDANASQSIEIVINPPKS 311
Query: 407 KRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRNPV 450
R PGRP+K R R SRC Q GH +C P+
Sbjct: 310 LRPPGRPRKKRVRAEDRGRVKRVVHC-SRCNQTGHFRTTCAAPI 182
>CB827002
Length = 534
Score = 29.6 bits (65), Expect = 1.1
Identities = 18/50 (36%), Positives = 26/50 (52%)
Frame = +1
Query: 302 EMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRH 351
E++ Y VA + GED H +F V + +CSC +E G+ CRH
Sbjct: 400 EVITYHVA-KFGED-----HKAYYVKFNVLEMKASCSCQMFEFSGLLCRH 531
>TC15105 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein AtPOB1,
partial (31%)
Length = 831
Score = 29.6 bits (65), Expect = 1.1
Identities = 18/59 (30%), Positives = 31/59 (52%)
Frame = +1
Query: 240 LMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLE 298
L+ NL + ++ +L V + M D + Y+ G++ + L K+K EVM PL +E
Sbjct: 400 LLPNLPMAPDSPLLYVELPHTILMADAAKQYLAGQY----KDLTKFKEEVMALPLSGIE 564
>AV765466
Length = 599
Score = 29.3 bits (64), Expect = 1.4
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 2/39 (5%)
Frame = -2
Query: 337 CSCNFWELVGIPCRHAVAAISHS--SQRPEDFVHPYYRR 373
C C +E GI CRH++A +S + P+ +V +R+
Sbjct: 592 CDCRLFEFRGILCRHSLAILSQERVKEVPDRYVLDRWRK 476
>TC9034
Length = 583
Score = 28.9 bits (63), Expect = 1.8
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 8/62 (12%)
Frame = +1
Query: 394 PTTNDPY--ILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR------CGQFGHNSRS 445
P T+ Y I P KR PG + R+R E T + RS C +FGH+S
Sbjct: 7 PVTSGYYGKIDSPTLKRLPGWERINRKRGVTEGPTVSHEARRSHRVKCANCEEFGHDSWE 186
Query: 446 CR 447
C+
Sbjct: 187 CQ 192
>BP042653
Length = 452
Score = 28.5 bits (62), Expect = 2.4
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +1
Query: 393 WPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR 435
+P P + PPL ++P RP++ R P T N T+L R
Sbjct: 127 FPPFQSPPLHPPLLAQSPTRPRRCFHR-PQRANTKNFTKLHIR 252
>TC11134
Length = 376
Score = 28.5 bits (62), Expect = 2.4
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -3
Query: 384 ITPINGQKLWPTTNDPYILPPLYKR 408
ITPI Q L+PT +D + P +YK+
Sbjct: 101 ITPIQQQDLFPTASDHDLFPGVYKQ 27
>AV406521
Length = 267
Score = 27.3 bits (59), Expect = 5.3
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +3
Query: 155 DDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKAT 195
+D H FC+RHL + K+F ++ +L+ AA AT
Sbjct: 147 EDEFSNSSHTFCMRHLSESIGKEFKNSRLV-HLLWKAAYAT 266
>TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I7.2
{Arabidopsis thaliana;}, partial (13%)
Length = 640
Score = 26.9 bits (58), Expect = 6.9
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = +2
Query: 404 PLYKRAPGRPKKLRRRDP 421
P +R P PK+LRRRDP
Sbjct: 143 PQRRRLPPLPKRLRRRDP 196
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 26.9 bits (58), Expect = 6.9
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 430 TRLRSRCGQFGHNSRSC 446
+R S+CG GHNSR+C
Sbjct: 190 SRTCSQCGNNGHNSRTC 240
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,927,892
Number of Sequences: 28460
Number of extensions: 130830
Number of successful extensions: 992
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 981
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 988
length of query: 451
length of database: 4,897,600
effective HSP length: 93
effective length of query: 358
effective length of database: 2,250,820
effective search space: 805793560
effective search space used: 805793560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0213.7


