

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.11
(579 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
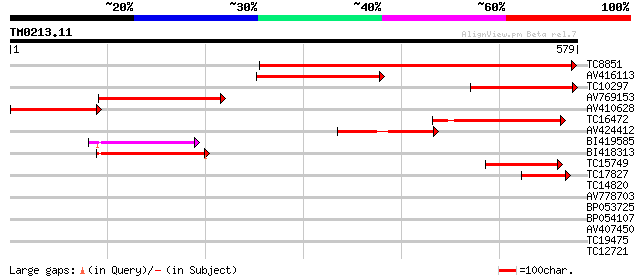
Score E
Sequences producing significant alignments: (bits) Value
TC8851 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-phos... 581 e-166
AV416113 261 2e-70
TC10297 similar to UP|G6PC_SOLTU (Q43839) Glucose-6-phosphate 1-... 221 2e-58
AV769153 200 5e-52
AV410628 194 3e-50
TC16472 similar to UP|G6PD_SOLTU (P37830) Glucose-6-phosphate 1-... 119 1e-27
AV424412 110 8e-25
BI419585 84 5e-17
BI418313 82 2e-16
TC15749 homologue to UP|Q9ZSR1 (Q9ZSR1) Cytoplasmic glucose-6-ph... 74 6e-14
TC17827 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate 1-... 68 5e-12
TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate p... 33 0.17
AV778703 32 0.28
BP053725 30 1.4
BP054107 28 4.1
AV407450 28 5.3
TC19475 28 5.3
TC12721 similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferas... 28 5.3
>TC8851 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-phosphate
1-dehydrogenase {Arabidopsis thaliana;} , partial (54%)
Length = 1291
Score = 581 bits (1497), Expect = e-166
Identities = 279/323 (86%), Positives = 303/323 (93%)
Frame = +1
Query: 256 LTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYF 315
LTEDQIFRIDHYLGKELVENLSVLRFSNL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYF
Sbjct: 13 LTEDQIFRIDHYLGKELVENLSVLRFSNLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYF 192
Query: 316 DNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKG 375
DNYGIIRDIMQNHLLQILALFAME PVSLDAEDIRNEKVKVLRSMRPL+LE+++ GQYK
Sbjct: 193 DNYGIIRDIMQNHLLQILALFAMETPVSLDAEDIRNEKVKVLRSMRPLRLEDMIIGQYKS 372
Query: 376 HSKGGKSYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRV 435
H++GG +YP Y DD +VPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRV
Sbjct: 373 HTRGGVTYPAYTDDKTVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRV 552
Query: 436 QFRHVPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYR 495
QFRHVPGN+Y RNFG D+D+ATNELV+RVQPDEAIYLK+NNK+PGL MRLDRS+LNL Y
Sbjct: 553 QFRHVPGNVYNRNFGTDLDRATNELVIRVQPDEAIYLKINNKVPGLEMRLDRSNLNLHYA 732
Query: 496 ARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGS 555
ARY EIPDAYERLLLDAIEGERRLFIRSDELDAAW+LFTPLL E+E+KK+ PE YP+GS
Sbjct: 733 ARYAKEIPDAYERLLLDAIEGERRLFIRSDELDAAWSLFTPLLNELEEKKVIPEYYPYGS 912
Query: 556 RGPIGAHYLAAKHNVRWGDTGND 578
RGP+GAHYLAA++NVRWGD G D
Sbjct: 913 RGPVGAHYLAARYNVRWGDLGID 981
>AV416113
Length = 392
Score = 261 bits (667), Expect = 2e-70
Identities = 129/130 (99%), Positives = 130/130 (99%)
Frame = +2
Query: 253 KQYLTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRG 312
KQYLTEDQIFRIDHYLGKELVENLSVLRFS+LVFEPLWSRAYIRNVQFIFSEDFGTEGRG
Sbjct: 2 KQYLTEDQIFRIDHYLGKELVENLSVLRFSHLVFEPLWSRAYIRNVQFIFSEDFGTEGRG 181
Query: 313 GYFDNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQ 372
GYFDNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQ
Sbjct: 182 GYFDNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQ 361
Query: 373 YKGHSKGGKS 382
YKGHSKGGKS
Sbjct: 362 YKGHSKGGKS 391
>TC10297 similar to UP|G6PC_SOLTU (Q43839) Glucose-6-phosphate
1-dehydrogenase, chloroplast precursor (G6PD) , partial
(19%)
Length = 547
Score = 221 bits (564), Expect = 2e-58
Identities = 106/109 (97%), Positives = 108/109 (98%)
Frame = +3
Query: 471 YLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAA 530
YLKVNN+IPGLGMRLD SDL+LLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAA
Sbjct: 3 YLKVNNQIPGLGMRLDSSDLHLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAA 182
Query: 531 WALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGNDD 579
WALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGNDD
Sbjct: 183 WALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGNDD 329
>AV769153
Length = 435
Score = 200 bits (509), Expect = 5e-52
Identities = 93/130 (71%), Positives = 114/130 (87%)
Frame = +1
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S++SITVVGASGDLAKKKIFPALFAL+YE CLP++F + G+AR+KMT ELRNM+S+TLT
Sbjct: 46 SSVSITVVGASGDLAKKKIFPALFALYYEGCLPKHFTICGYARSKMTDAELRNMVSKTLT 225
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CRIDKR NC +KM+Q L+RCFYHSG Y+SE + + LD K+KE EGG+ SNRLFYLSIPPN
Sbjct: 226 CRIDKRENCSEKMEQXLERCFYHSGQYDSEDNXAALDKKLKEHEGGRTSNRLFYLSIPPN 405
Query: 211 IFVNVVRCAS 220
IF++ V+CAS
Sbjct: 406 IFIDAVKCAS 435
>AV410628
Length = 290
Score = 194 bits (494), Expect = 3e-50
Identities = 93/93 (100%), Positives = 93/93 (100%)
Frame = +3
Query: 1 MACVLSSSSTIVASYALRNEPQLYPVWFSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHD 60
MACVLSSSSTIVASYALRNEPQLYPVWFSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHD
Sbjct: 12 MACVLSSSSTIVASYALRNEPQLYPVWFSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHD 191
Query: 61 GLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNL 93
GLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNL
Sbjct: 192 GLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNL 290
>TC16472 similar to UP|G6PD_SOLTU (P37830) Glucose-6-phosphate
1-dehydrogenase, cytoplasmic isoform (G6PD) , partial
(29%)
Length = 673
Score = 119 bits (299), Expect = 1e-27
Identities = 64/137 (46%), Positives = 91/137 (65%), Gaps = 1/137 (0%)
Frame = +3
Query: 432 EIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLN 491
EIR+QF+ VPG++++ + NELV+R+QP EAIY+K+ K PGL M +S+L+
Sbjct: 3 EIRIQFKDVPGDIFRCK-----KQGRNELVIRLQPLEAIYMKLTVKQPGLEMSTVQSELD 167
Query: 492 LLYRARYPTEI-PDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPEL 550
L Y+ RY I P+AYERL+LD I G+++ F+R DEL A+W +FTPLL +++ + P
Sbjct: 168 LSYKQRYQGMIIPEAYERLILDTIRGDQQHFVRRDELKASWKIFTPLLHKIDRGEFKPIP 347
Query: 551 YPHGSRGPIGAHYLAAK 567
Y GSRGP A L K
Sbjct: 348 YKPGSRGPAEADQLLEK 398
>AV424412
Length = 287
Score = 110 bits (274), Expect = 8e-25
Identities = 57/104 (54%), Positives = 76/104 (72%)
Frame = +2
Query: 335 LFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPK 394
L AME PVSL+ E IR+EKVKVL+S+ PL+ ++VV GQY+G Y DD +VP
Sbjct: 5 LVAMEKPVSLNPEHIRDEKVKVLQSVLPLKDDDVVLGQYEG----------YRDDSTVPD 154
Query: 395 GSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFR 438
S TPTFA L + N RW+GVPF++KAGKAL++++A+IRVQF+
Sbjct: 155 HSNTPTFATVILRVHNERWEGVPFILKAGKALNSRKADIRVQFK 286
>BI419585
Length = 523
Score = 84.3 bits (207), Expect = 5e-17
Identities = 54/118 (45%), Positives = 70/118 (58%), Gaps = 4/118 (3%)
Frame = +1
Query: 81 FLSSDSEC---TGSNLSITVVGASGDLAKKKIFPALFALFYEDCLP-ENFLVFGFARTKM 136
F++ D E TGS LSI V+GASGDLAKKK FPALF L+ + LP + +FG+ARTK+
Sbjct: 169 FINEDLESAPETGS-LSIIVLGASGDLAKKKTFPALFNLYRQGFLPADEICIFGYARTKI 345
Query: 137 TSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKE 194
+ +ELRN + L D + +FL Y SG Y+SE F LD K+ E E
Sbjct: 346 SDDELRNRLHGFLVKEKDASPEQLQDVSKFLHLIKYVSGSYDSENDFHLLDKKISEHE 519
>BI418313
Length = 574
Score = 82.0 bits (201), Expect = 2e-16
Identities = 53/122 (43%), Positives = 74/122 (60%), Gaps = 6/122 (4%)
Frame = +3
Query: 89 TGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFL-VFGFARTKMTSEELRNMISR 147
TGS LSI V+GASGDLAKKK FPALF L+ + LP++ + +FG+ARTK++ ++LR+ +
Sbjct: 210 TGS-LSIVVLGASGDLAKKKTFPALFHLYLQGFLPQDEVHIFGYARTKISDDDLRDRLRG 386
Query: 148 TLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKR-----SNRL 202
L D + + +FL+ Y SG Y+S F LD ++ E E K S RL
Sbjct: 387 YLIPDKDASPKQLEDVSKFLQLIKYVSGSYDSGDGFILLDKEISEHESSKNSIEGSSRRL 566
Query: 203 FY 204
FY
Sbjct: 567 FY 572
>TC15749 homologue to UP|Q9ZSR1 (Q9ZSR1) Cytoplasmic glucose-6-phosphate
1-dehydrogenase (G6PD) , partial (19%)
Length = 676
Score = 73.9 bits (180), Expect = 6e-14
Identities = 37/79 (46%), Positives = 53/79 (66%), Gaps = 1/79 (1%)
Frame = +3
Query: 487 RSDLNLLYRARYP-TEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKK 545
+S+L+L YR RY IP+AYERL+LD I+G+++ F+R DEL A+W +FTPLL ++ +
Sbjct: 3 QSELDLSYRQRYQGVTIPEAYERLILDTIKGDQQHFVRRDELKASWEIFTPLLHSIDKGE 182
Query: 546 IAPELYPHGSRGPIGAHYL 564
Y G+RGP A L
Sbjct: 183 FKSIPYQPGTRGPAEADEL 239
>TC17827 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate
1-dehydrogenase 4, chloroplast precursor (G6PD4)
(G6PDH4) , partial (8%)
Length = 534
Score = 67.8 bits (164), Expect = 5e-12
Identities = 28/50 (56%), Positives = 37/50 (74%)
Frame = +2
Query: 523 RSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRW 572
RSDEL AAW + TP+L E++ ++ ELY G RGP+GA+YL AKH +RW
Sbjct: 2 RSDELAAAWNILTPILNEIDKDNVSVELYELGGRGPVGAYYLWAKHGIRW 151
>TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate protein 80,
partial (53%)
Length = 1180
Score = 32.7 bits (73), Expect = 0.17
Identities = 18/41 (43%), Positives = 23/41 (55%)
Frame = -1
Query: 6 SSSSTIVASYALRNEPQLYPVWFSCIRPGNLPRNHFQLKSS 46
SSSS+ + S +LR P LYP + P +LPR H Q S
Sbjct: 826 SSSSSFLPSPSLRQHPPLYP-----LHPSSLPRPHHQRSPS 719
>AV778703
Length = 603
Score = 32.0 bits (71), Expect = 0.28
Identities = 16/62 (25%), Positives = 27/62 (42%)
Frame = -1
Query: 410 NARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQPDEA 469
N WD P+ + HTK + H+ N + FG + ++ ++LRV P
Sbjct: 186 NVSWDTFPYHIHE----HTKSLLMECAASHLRHNKFTSTFGTHLTSSSRRILLRVLPSTE 19
Query: 470 IY 471
+Y
Sbjct: 18 LY 13
>BP053725
Length = 517
Score = 29.6 bits (65), Expect = 1.4
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = -1
Query: 49 HPLNAVSSHSHDGLAGSSLTKEDSKPQP 76
H L H +GL+ + KEDSKP P
Sbjct: 475 HHLRPXQKHEKEGLSANGPNKEDSKPDP 392
>BP054107
Length = 327
Score = 28.1 bits (61), Expect = 4.1
Identities = 12/20 (60%), Positives = 13/20 (65%), Gaps = 1/20 (5%)
Frame = -3
Query: 556 RGPIGAHYLAAKH-NVRWGD 574
RGP+GAHY VRWGD
Sbjct: 274 RGPVGAHYSCRPDTTVRWGD 215
>AV407450
Length = 337
Score = 27.7 bits (60), Expect = 5.3
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +3
Query: 9 STIVASYALRNEPQLYPVWFSCIRPGNLPRNHFQLKSSNGHP 50
S++ SY+ +N +P+ P +LP HFQ KSS+ P
Sbjct: 102 SSLPPSYSPQNHAHPHPL----TTPSHLPSLHFQFKSSSKTP 215
>TC19475
Length = 524
Score = 27.7 bits (60), Expect = 5.3
Identities = 17/41 (41%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Frame = +1
Query: 182 SFSDLDCKMKEKEGGKRSNRL--FYLSIPPNIFVNVVRCAS 220
+FSD+ MKE + G RS+ L F L + PN V+ AS
Sbjct: 148 TFSDVGWNMKESDTGDRSSMLLPFLLGLLPNTRALVIFSAS 270
>TC12721 similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase 85A8,
partial (4%)
Length = 573
Score = 27.7 bits (60), Expect = 5.3
Identities = 13/21 (61%), Positives = 16/21 (75%)
Frame = +1
Query: 341 PVSLDAEDIRNEKVKVLRSMR 361
P LD ED RN++VKVL S+R
Sbjct: 412 PDGLDPEDDRNDQVKVLLSIR 474
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,869,389
Number of Sequences: 28460
Number of extensions: 135751
Number of successful extensions: 622
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 616
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 617
length of query: 579
length of database: 4,897,600
effective HSP length: 95
effective length of query: 484
effective length of database: 2,193,900
effective search space: 1061847600
effective search space used: 1061847600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0213.11


