

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
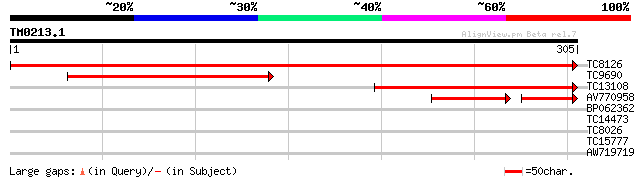
Query= TM0213.1
(305 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8126 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase... 647 0.0
TC9690 similar to UP|AAP59548 (AAP59548) Myo-inositol oxygenase,... 171 2e-43
TC13108 similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase (Myo... 170 2e-43
AV770958 89 2e-27
BP062362 35 0.021
TC14473 similar to UP|Q8H1Y2 (Q8H1Y2) 20S proteasome alpha 6 sub... 28 2.5
TC8026 27 3.3
TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing... 26 7.3
AW719719 26 7.3
>TC8126 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase
(Myo-inositol oxygenase) (Aldehyde reductase-like 6) ,
partial (39%)
Length = 1368
Score = 647 bits (1670), Expect = 0.0
Identities = 305/305 (100%), Positives = 305/305 (100%)
Frame = +2
Query: 1 MTIIIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVEN 60
MTIIIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVEN
Sbjct: 200 MTIIIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVEN 379
Query: 61 FYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQT 120
FYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQT
Sbjct: 380 FYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQT 559
Query: 121 AEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKH 180
AEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKH
Sbjct: 560 AEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKH 739
Query: 181 FKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYH 240
FKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYH
Sbjct: 740 FKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYH 919
Query: 241 SFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFP 300
SFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFP
Sbjct: 920 SFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFP 1099
Query: 301 AKLKW 305
AKLKW
Sbjct: 1100AKLKW 1114
>TC9690 similar to UP|AAP59548 (AAP59548) Myo-inositol oxygenase, partial
(35%)
Length = 519
Score = 171 bits (432), Expect = 2e-43
Identities = 77/111 (69%), Positives = 90/111 (80%)
Frame = +1
Query: 32 GFVMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSM 91
GF+ P NS+G +FRDY ESERQ+ VE YR HI Q+ +F K MREEYGKLN+ EM +
Sbjct: 187 GFIAPDINSYGKSFRDYYGESERQKSVEELYRLQHINQTYEFAKTMREEYGKLNKGEMGI 366
Query: 92 WECCELLNEVVDESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHD 142
WECCELL+E+VD SDPDL+E QI+H LQ+AEAIRKDYPNEDWLHLT LIHD
Sbjct: 367 WECCELLDEIVDASDPDLEESQIKHALQSAEAIRKDYPNEDWLHLTALIHD 519
>TC13108 similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase (Myo-inositol
oxygenase) (Aldehyde reductase-like 6) , partial (6%)
Length = 592
Score = 170 bits (431), Expect = 2e-43
Identities = 77/109 (70%), Positives = 93/109 (84%)
Frame = +3
Query: 197 GMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYKHLMN 256
G+Y+E CGL+NV++SWGHD+YM+LVAK N TLPS ALFIIRYHSF+ L+ GAYKHL +
Sbjct: 6 GIYAEGCGLDNVIISWGHDEYMHLVAKGNSNTLPSVALFIIRYHSFHPLYEAGAYKHLYS 185
Query: 257 EEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
EED +NL+WL IF+ YDLYS+S V IDVE VKPYY+SLIEKYFP KL+W
Sbjct: 186 EEDVQNLKWLEIFH*YDLYSQSHVPIDVEPVKPYYMSLIEKYFPPKLQW 332
>AV770958
Length = 459
Score = 88.6 bits (218), Expect(2) = 2e-27
Identities = 39/42 (92%), Positives = 41/42 (96%)
Frame = -1
Query: 228 TLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIF 269
TLPSAALFIIRYHSFYALHREGAY HLMNEE+FENLEWLH+F
Sbjct: 459 TLPSAALFIIRYHSFYALHREGAYNHLMNEEEFENLEWLHLF 334
Score = 50.1 bits (118), Expect(2) = 2e-27
Identities = 20/30 (66%), Positives = 26/30 (86%)
Frame = -3
Query: 276 SKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
+++K + EKV+PYYLSLI+KYFPAKLKW
Sbjct: 313 ARAKFELSAEKVQPYYLSLIDKYFPAKLKW 224
>BP062362
Length = 387
Score = 34.7 bits (78), Expect = 0.021
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 6/39 (15%)
Frame = -3
Query: 92 WECCELLNEVVDESDP------DLDEPQIEHLLQTAEAI 124
W CC ++ V SD + +EPQ++HLL TAE +
Sbjct: 346 WRCCHTSSQTVPASDSSSTEDEEPEEPQLQHLLSTAEGV 230
>TC14473 similar to UP|Q8H1Y2 (Q8H1Y2) 20S proteasome alpha 6 subunit,
partial (86%)
Length = 1302
Score = 27.7 bits (60), Expect = 2.5
Identities = 15/60 (25%), Positives = 28/60 (46%)
Frame = -2
Query: 241 SFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFP 300
SF LH + HL+N +N+EWL N + +++ ++ + + +K FP
Sbjct: 866 SFILLHNLKSINHLLNSFLIQNVEWLTNSNHSNSTNRTPELFTLQGLPGCNECIFDKIFP 687
>TC8026
Length = 667
Score = 27.3 bits (59), Expect = 3.3
Identities = 16/61 (26%), Positives = 32/61 (52%)
Frame = +2
Query: 124 IRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKE 183
+ +DY + ++ GL+++ +V+ + V +++V PV C FD S + HF E
Sbjct: 368 LSRDYSSC*FVK*LGLVNEFEEVVFIKVV----EFSVSKLLSPVSCNFDFSSSSYTHFSE 535
Query: 184 N 184
+
Sbjct: 536 H 538
>TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (34%)
Length = 566
Score = 26.2 bits (56), Expect = 7.3
Identities = 14/43 (32%), Positives = 20/43 (45%)
Frame = +3
Query: 166 PVGCRFDESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNV 208
P F S H +H +P Y+ A N + + +K GLN V
Sbjct: 39 PTKASFRASTPHPQHTTPSPSYHIQATNREHIVRQKKNGLNAV 167
>AW719719
Length = 512
Score = 26.2 bits (56), Expect = 7.3
Identities = 12/49 (24%), Positives = 24/49 (48%)
Frame = +1
Query: 49 NAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCEL 97
N+E +E + N ++ + ++G+L+ MS+W CC+L
Sbjct: 10 NSERTINTNIEKLFLCNDACHYLNSMFFFLPDFGRLSVSFMSVWLCCDL 156
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,931,348
Number of Sequences: 28460
Number of extensions: 90164
Number of successful extensions: 364
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 363
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 364
length of query: 305
length of database: 4,897,600
effective HSP length: 90
effective length of query: 215
effective length of database: 2,336,200
effective search space: 502283000
effective search space used: 502283000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0213.1


