

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.4
(342 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
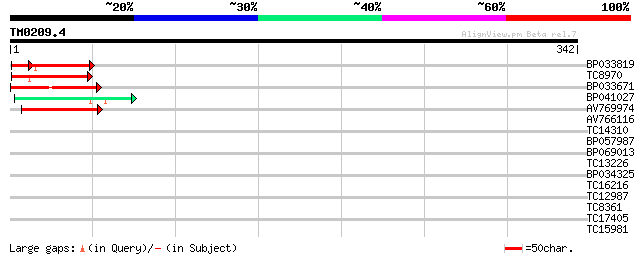
Score E
Sequences producing significant alignments: (bits) Value
BP033819 63 2e-11
TC8970 64 3e-11
BP033671 49 9e-07
BP041027 43 7e-05
AV769974 41 3e-04
AV766116 31 0.34
TC14310 homologue to UP|Q9M3H6 (Q9M3H6) Histone H2B, partial (92%) 31 0.34
BP057987 28 2.2
BP069013 28 2.2
TC13226 similar to UP|HMGB_SOYBN (Q10370) HMG-Y related protein ... 27 6.5
BP034325 27 6.5
TC16216 similar to UP|RK9_PEA (P11894) 50S ribosomal protein L9,... 27 6.5
TC12987 similar to UP|Q9SM91 (Q9SM91) Zwh18.1, partial (5%) 27 6.5
TC8361 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial... 26 8.5
TC17405 similar to UP|Q940D1 (Q940D1) At1g64110/F22C12_22, parti... 26 8.5
TC15981 similar to UP|ERF3_ARATH (O80339) Ethylene responsive el... 26 8.5
>BP033819
Length = 554
Score = 63.2 bits (152), Expect(2) = 2e-11
Identities = 30/43 (69%), Positives = 35/43 (80%), Gaps = 3/43 (6%)
Frame = +1
Query: 12 RPP--YVRQT-PCDPHGREPQKKLIVGDLPCPKMRNDNQSCDT 51
RPP ++R T PCDPHGREP +KLIVG+L CP MRND+QS DT
Sbjct: 334 RPPKDFLRPTTPCDPHGREPHEKLIVGNLSCPNMRNDSQSYDT 462
Score = 21.9 bits (45), Expect(2) = 2e-11
Identities = 10/13 (76%), Positives = 10/13 (76%)
Frame = +3
Query: 2 KVSQPKMASMRPP 14
KVSQPK AS R P
Sbjct: 315 KVSQPKTASKRLP 353
>TC8970
Length = 609
Score = 64.3 bits (155), Expect = 3e-11
Identities = 32/54 (59%), Positives = 36/54 (66%), Gaps = 5/54 (9%)
Frame = -1
Query: 2 KVSQPKMAS-----MRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMRNDNQSCD 50
K S PK S + Y RQTPCDPHGRE KKLIVG+LPCPK++ NQS D
Sbjct: 405 KYSPPKSKSAQNGFQKTSYARQTPCDPHGRESHKKLIVGNLPCPKIQLMNQSYD 244
>BP033671
Length = 482
Score = 49.3 bits (116), Expect = 9e-07
Identities = 26/55 (47%), Positives = 34/55 (61%)
Frame = -3
Query: 1 MKVSQPKMASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMRNDNQSCDTTPTL 55
+++SQP+ RPP Q P P ++P KKL VG+ P PK ND S D+TPTL
Sbjct: 168 LRISQPRWPKTRPPKSNQLPVIPV-KKPHKKL*VGNYPVPKGTNDVPS*DSTPTL 7
>BP041027
Length = 527
Score = 43.1 bits (100), Expect = 7e-05
Identities = 35/100 (35%), Positives = 39/100 (39%), Gaps = 27/100 (27%)
Frame = +3
Query: 4 SQPKMASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMRNDNQ-----SCDTTPTLT-- 56
S PK + P VRQT CD REP K G P PK+R C T P T
Sbjct: 162 SAPKWPKQKTPIVRQTLCDSSSREPHKSHASGTYPVPKVRRIRALHKI*RCLTYPPSTAH 341
Query: 57 --------------------QSESRTINSLFTSMTSLLTV 76
+SES I S+ S TSLLTV
Sbjct: 342 SSSSSPSSLGE*LSTSLSESESESIVIISVSKSTTSLLTV 461
>AV769974
Length = 406
Score = 41.2 bits (95), Expect = 3e-04
Identities = 22/50 (44%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Frame = +1
Query: 8 MASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMR-NDNQSCDTTPTLT 56
MA + P V+ T CDP REP +K VG+LP PK + N + + + TP T
Sbjct: 190 MA*AKTPMVQLTHCDPSVREPHEKSCVGNLPDPKSKTNQSSTQNMTPAQT 339
>AV766116
Length = 357
Score = 30.8 bits (68), Expect = 0.34
Identities = 13/18 (72%), Positives = 14/18 (77%)
Frame = +2
Query: 26 REPQKKLIVGDLPCPKMR 43
REPQ+KL GDLPCP R
Sbjct: 194 REPQEKLRGGDLPCPNER 247
>TC14310 homologue to UP|Q9M3H6 (Q9M3H6) Histone H2B, partial (92%)
Length = 502
Score = 30.8 bits (68), Expect = 0.34
Identities = 21/43 (48%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Frame = +1
Query: 197 SARTILVSTLGILAKVPSLG---GRIRPLGVGGALPIWRSGLA 236
S+R+ L S LG LA SL GR +PL V G L WRS L+
Sbjct: 307 SSRSSLRSLLGSLATTRSLRSPLGRFKPLCVWGFLENWRSTLS 435
>BP057987
Length = 337
Score = 28.1 bits (61), Expect = 2.2
Identities = 16/45 (35%), Positives = 21/45 (46%)
Frame = +2
Query: 6 PKMASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMRNDNQSCD 50
PK R P + P EP +K VG+ PK++ NQS D
Sbjct: 203 PKGLPKRLPTSDYSRRSPSSGEPHRKQPVGNYRVPKVQRTNQSYD 337
>BP069013
Length = 470
Score = 28.1 bits (61), Expect = 2.2
Identities = 14/27 (51%), Positives = 18/27 (65%), Gaps = 1/27 (3%)
Frame = +3
Query: 40 PKMRNDNQSCDTTPTLTQSE-SRTINS 65
PK++ D QS D TPTL S SR++ S
Sbjct: 273 PKLKRDGQSIDPTPTLIPSRVSRSVAS 353
>TC13226 similar to UP|HMGB_SOYBN (Q10370) HMG-Y related protein B (SB16B
protein) (Fragment), partial (60%)
Length = 474
Score = 26.6 bits (57), Expect = 6.5
Identities = 15/33 (45%), Positives = 20/33 (60%)
Frame = -3
Query: 204 STLGILAKVPSLGGRIRPLGVGGALPIWRSGLA 236
+TLG + + S GGR PLG+GGA + R A
Sbjct: 145 TTLGFGSALGSFGGR--PLGLGGATTVSRGRAA 53
>BP034325
Length = 417
Score = 26.6 bits (57), Expect = 6.5
Identities = 17/52 (32%), Positives = 23/52 (43%)
Frame = -2
Query: 227 ALPIWRSGLASSSFATAIRAIMASSVCGVGGVEPLEPVAVDLVSTILLQCHT 278
+LP WR +SSSF + A CG GV + V V +L H+
Sbjct: 404 SLPFWRRNPSSSSFKSGGGG*FAVVGCGGDGVRLVWFAGVSSVGVVLGCAHS 249
>TC16216 similar to UP|RK9_PEA (P11894) 50S ribosomal protein L9,
chloroplast precursor (CL13), partial (96%)
Length = 944
Score = 26.6 bits (57), Expect = 6.5
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +3
Query: 34 VGDLPCPKMRNDNQSCDTTPTLTQSESRTINSLFTSMTSLLT 75
VGD K+ N P+L+ + S ++ FT+ LLT
Sbjct: 12 VGDTDTTKLANTTTMASAFPSLSSTSSSFLHHCFTANPKLLT 137
>TC12987 similar to UP|Q9SM91 (Q9SM91) Zwh18.1, partial (5%)
Length = 552
Score = 26.6 bits (57), Expect = 6.5
Identities = 14/40 (35%), Positives = 17/40 (42%)
Frame = -3
Query: 2 KVSQPKMASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPK 41
KV K + P +RQT CDP + K P PK
Sbjct: 352 KVGTQKGPERKTPILRQTLCDPPKKNHTKSHTSEPYPVPK 233
>TC8361 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial (92%)
Length = 1523
Score = 26.2 bits (56), Expect = 8.5
Identities = 45/165 (27%), Positives = 65/165 (39%), Gaps = 10/165 (6%)
Frame = +3
Query: 124 PSGRPRHP--TRVGRDIELTSDKTKWQDAMADRFSRKTKLAWIESQRQDLQVVGIKDRDI 181
P G P VG+D L + +W +A A T W + DI
Sbjct: 636 PHGMPERKMGAAVGQDCALLEEVCRWINAKA------TVPVWAKMTPNIT--------DI 773
Query: 182 SQPACVDIASHRVDVSARTILVSTLGI----LAKVPSLGGRIRPLGVGGALPIWRSGLAS 237
S+PA V ++S V+A ++S +GI L P + G P G A + LA
Sbjct: 774 SEPARVALSSGC*GVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGY-SAKAVHPIALAK 950
Query: 238 -SSFATAIRAIMAS---SVCGVGGVEPLEPVAVDLVSTILLQCHT 278
S A +++ S S+ +GGVE D ILL +T
Sbjct: 951 VMSIAKMMKSEFDSEDYSLSAIGGVE----TGGDAAEFILLGANT 1073
>TC17405 similar to UP|Q940D1 (Q940D1) At1g64110/F22C12_22, partial (10%)
Length = 522
Score = 26.2 bits (56), Expect = 8.5
Identities = 17/60 (28%), Positives = 24/60 (39%)
Frame = -2
Query: 3 VSQPKMASMRPPYVRQTPCDPHGREPQKKLIVGDLPCPKMRNDNQSCDTTPTLTQSESRT 62
+ P+ +S PP T C P P P + DN++C + T T ES T
Sbjct: 365 ICSPETSSEFPPTHFCTACPELNPSPTPN------PTPTLNADNRTCFCSITTTHLESET 204
>TC15981 similar to UP|ERF3_ARATH (O80339) Ethylene responsive element
binding factor 3 (AtERF3), partial (45%)
Length = 671
Score = 26.2 bits (56), Expect = 8.5
Identities = 10/24 (41%), Positives = 18/24 (74%)
Frame = +2
Query: 55 LTQSESRTINSLFTSMTSLLTVRV 78
LT+S+SRT+ ++ T + L T+R+
Sbjct: 599 LTRSQSRTVTAIATPLPPLSTIRI 670
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,919,164
Number of Sequences: 28460
Number of extensions: 77773
Number of successful extensions: 354
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 345
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 352
length of query: 342
length of database: 4,897,600
effective HSP length: 91
effective length of query: 251
effective length of database: 2,307,740
effective search space: 579242740
effective search space used: 579242740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0209.4


