

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.3
(1547 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
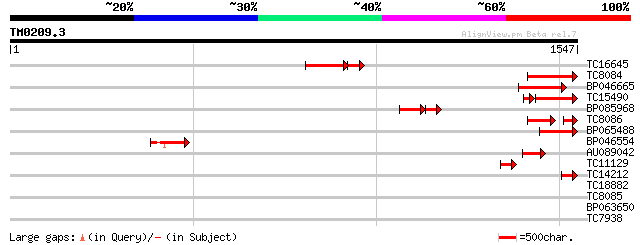
Score E
Sequences producing significant alignments: (bits) Value
TC16645 177 1e-57
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 183 2e-46
BP046665 171 8e-43
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 145 1e-40
BP085968 99 5e-37
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 93 5e-28
BP065488 96 3e-20
BP046554 86 4e-17
AU089042 61 1e-09
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 57 3e-08
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 47 2e-05
TC18882 31 1.7
TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meioti... 30 2.2
BP063650 27 4.2
TC7938 homologue to UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein,... 28 8.5
>TC16645
Length = 596
Score = 177 bits (450), Expect(2) = 1e-57
Identities = 81/119 (68%), Positives = 98/119 (82%)
Frame = +1
Query: 806 MKANKRYPEGRNLTYAEFPSKFVYREGAHEWVPRKRGLSLGRVQYIAPGMGEVYYMRVLL 865
M ANK+Y EGRNLTYAEFPS+F+Y + +WVP+++ SLGR+ +IAPGMGE YYMRVLL
Sbjct: 7 MNANKKYQEGRNLTYAEFPSEFLYHQHTTKWVPKQKRFSLGRLPFIAPGMGENYYMRVLL 186
Query: 866 TRQRGCDSFESIRTVKGIAYPTFHDACEAMRLMDDDREYIDGIGDVSVLGSGAYLRNLF 924
T Q+GCDSF+SI+TVKG+ YPTFHDACEAM L++DDREY+DGI S GSG LR LF
Sbjct: 187 TMQKGCDSFKSIKTVKGVVYPTFHDACEAMGLLEDDREYVDGISISSEFGSGTQLRKLF 363
Score = 64.7 bits (156), Expect(2) = 1e-57
Identities = 28/45 (62%), Positives = 35/45 (77%)
Frame = +2
Query: 922 NLFTRILTTNAMSRPKEVWEKTWRILSDGILYERRKALNIPGVLS 966
N FTR+L TN +SRP+EVW K+W +L DGILY+RRK LNI + S
Sbjct: 356 NFFTRMLMTNLISRPQEVWRKSWSLLCDGILYDRRKTLNISDIQS 490
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 183 bits (465), Expect = 2e-46
Identities = 89/135 (65%), Positives = 109/135 (79%)
Frame = +1
Query: 1413 IDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPSDSGLPFKFSRRQF 1472
I + LCNGTRL V L Y+I ATV++G +G +IPR+ + PSDSG PFKF RR F
Sbjct: 43 IQKFENLCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 222
Query: 1473 PVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVS 1532
P++LCFAMTINKSQGQSLSHV LYL RPVFTHGQLYVALSRV+SRKGLK+L++ E+ V+
Sbjct: 223 PISLCFAMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVT 402
Query: 1533 TSTRNVVYKEVFENV 1547
+T+NVVY+EVFEN+
Sbjct: 403 NTTKNVVYREVFENI 447
>BP046665
Length = 524
Score = 171 bits (433), Expect = 8e-43
Identities = 86/131 (65%), Positives = 101/131 (76%), Gaps = 1/131 (0%)
Frame = -1
Query: 1388 DVQCSGIPNHRLILKERVPIMLLRNIDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGK 1447
D+ C GIPNH++ LKE PIMLLRNI QA G CNGTRL V L +I ATV++ +G
Sbjct: 524 DITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGD 345
Query: 1448 TEYIPRITLTPSDSGLPFKFSRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQL 1507
+IPR+ + PSDSG PFKF RRQFP++LC AMTINKSQGQSLSHVGLYL R VFTHGQL
Sbjct: 344 DIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQL 165
Query: 1508 YVAL-SRVKSR 1517
YVAL +++K R
Sbjct: 164 YVALHAKIKKR 132
Score = 38.1 bits (87), Expect = 0.011
Identities = 18/45 (40%), Positives = 31/45 (68%)
Frame = -2
Query: 1496 YLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVY 1540
Y+ +++H LS ++SRKGLK+L++ E+ V+ +T+NVVY
Sbjct: 202 YIFLAMYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 145 bits (365), Expect(2) = 1e-40
Identities = 70/113 (61%), Positives = 90/113 (78%)
Frame = +2
Query: 1435 IVATVLSGIRLGKTEYIPRITLTPSDSGLPFKFSRRQFPVTLCFAMTINKSQGQSLSHVG 1494
I TV++G +G IPR+ + PSDS PFKF RRQ P++LCFAMTINKSQG+SLSHVG
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVG 316
Query: 1495 LYLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
LYL RPV THG LYVAL RV+SRK LK+L++ E+ ++ +T+NVVY+E+FEN+
Sbjct: 317 LYLSRPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFENI 475
Score = 40.0 bits (92), Expect(2) = 1e-40
Identities = 21/29 (72%), Positives = 21/29 (72%)
Frame = +1
Query: 1402 KERVPIMLLRNIDQAGGLCNGTRLRVTHL 1430
KE IMLLRNI QA GLCNGTRL V L
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIVVDL 129
>BP085968
Length = 341
Score = 98.6 bits (244), Expect(2) = 5e-37
Identities = 46/71 (64%), Positives = 58/71 (80%)
Frame = +2
Query: 1064 RSLTGEQKAAYKKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLIVLNVASS 1123
+S T +Q YK+++ V S+ GGF FLYGFGG+GKTFVWNT S+ +RS+GL+VLNVASS
Sbjct: 2 KSWTAKQSKVYKQIMSSVWSDDGGFYFLYGFGGSGKTFVWNTWSSGLRSQGLMVLNVASS 181
Query: 1124 GIASLLLPGGR 1134
GIAS LLPGG+
Sbjct: 182 GIASWLLPGGK 214
Score = 74.7 bits (182), Expect(2) = 5e-37
Identities = 35/43 (81%), Positives = 39/43 (90%)
Frame = +3
Query: 1134 RTAHSRFSILISINEVSTCNLRQGSPKAELLQKASLIIWDETP 1176
RTAHSRFSI ISIN++STCN++QGS KAELLQKAS IIWDE P
Sbjct: 213 RTAHSRFSIHISINDISTCNIKQGSQKAELLQKASSIIWDEAP 341
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 93.2 bits (230), Expect(2) = 5e-28
Identities = 44/75 (58%), Positives = 55/75 (72%)
Frame = +1
Query: 1413 IDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPSDSGLPFKFSRRQF 1472
I + LC+GTRL V L Y+I ATV++G +G +IPR+ + PSDSG PFKF RR F
Sbjct: 151 IQKFENLCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 330
Query: 1473 PVTLCFAMTINKSQG 1487
P++LCFAMTINKSQG
Sbjct: 331 PISLCFAMTINKSQG 375
Score = 50.1 bits (118), Expect(2) = 5e-28
Identities = 22/36 (61%), Positives = 32/36 (88%)
Frame = +3
Query: 1512 SRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
SRV+SRKGLK+L++ E+ V+ +T+NVVY+EVFEN+
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 476
>BP065488
Length = 439
Score = 96.3 bits (238), Expect = 3e-20
Identities = 53/104 (50%), Positives = 72/104 (68%), Gaps = 2/104 (1%)
Frame = -1
Query: 1446 GKTEYIPRITLTPSDSGLPFKFSRRQFPVTLCFAMTINKSQGQ-SLSHVGLYLPRPVFTH 1504
G +IPR+ + PS SG P KF R QFP++LCFAMTINKSQ V L ++H
Sbjct: 379 GDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQXSVHYPTVALISFLACYSH 200
Query: 1505 -GQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
+LYVALS V+SRKGLK+L++ E+ V+ +T+N VY+EVF+N+
Sbjct: 199 MDKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKNI 68
>BP046554
Length = 556
Score = 85.9 bits (211), Expect = 4e-17
Identities = 61/113 (53%), Positives = 70/113 (60%), Gaps = 7/113 (6%)
Frame = -1
Query: 384 RKPSTRGRQTPHLLESELFCHRLLPGDIAICLTIARMQ-------WPYARNLATRTSS*H 436
+KPSTR R+ LLE+ F I T +Q +RNL T+TSS*H
Sbjct: 391 KKPSTRERRILLLLEASYFA------PIFYWWTSLYVQ*FQGCNGHASSRNLGTQTSS*H 230
Query: 437 SLAIQLGVKYKGFYNLGITSRRTDPTFVSVFSK*SLIDSCLI*EKGKYSVLLM 489
LAIQLGVK+KG NLGI +T TFV VFSK*S IDS LI*+KGKYSVLLM
Sbjct: 229 LLAIQLGVKFKGLCNLGI*M*KTVQTFVFVFSK*SSIDSYLI*KKGKYSVLLM 71
>AU089042
Length = 191
Score = 61.2 bits (147), Expect = 1e-09
Identities = 30/61 (49%), Positives = 40/61 (65%)
Frame = +3
Query: 1400 ILKERVPIMLLRNIDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPS 1459
+LK VP+ML+ N+ + GLCNGTRL V +L +I AT+LSG +G YI + L PS
Sbjct: 6 VLKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPS 185
Query: 1460 D 1460
D
Sbjct: 186 D 188
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 56.6 bits (135), Expect = 3e-08
Identities = 26/46 (56%), Positives = 33/46 (71%)
Frame = +2
Query: 1338 PTLESVVHVNNYILSKLPGVEREYLSYDTPCRSDEDSEVHAEWFTS 1383
PTLESV VN ++L LPG EYLS DT C+ DED+E+ + WFT+
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WFTT 595
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 47.4 bits (111), Expect = 2e-05
Identities = 22/41 (53%), Positives = 31/41 (74%)
Frame = +2
Query: 1507 LYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
LYVA+SRVKS+ GLK+LI + +T+N+VYKEVF+ +
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVFQKI 124
>TC18882
Length = 339
Score = 30.8 bits (68), Expect = 1.7
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +1
Query: 150 QVFFFLPVSIYPSFLNYISSNIC 172
QVF FL +SIY + +Y SSN+C
Sbjct: 124 QVFLFLYLSIYHTLTHYTSSNLC 192
>TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meiotic
recombination protein)-like protein {Arabidopsis
thaliana;} , partial (8%)
Length = 500
Score = 30.4 bits (67), Expect = 2.2
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +1
Query: 933 MSRPKEVWEKTWRILSDGILYERRKALNIP 962
M+ P VW TW++L++ I +E R L P
Sbjct: 115 MTNPDHVWNITWKLLANDIQHEYRGTLTQP 204
>BP063650
Length = 497
Score = 26.9 bits (58), Expect(2) = 4.2
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +1
Query: 1219 QILPVISKGSRADMVGSTVTSSYLWKYCKVM 1249
+ L V+ KG D++ + V +SYL +C+V+
Sbjct: 58 KFLTVVYKGRMQDIIHAIVNASYL*DHCQVL 150
Score = 20.8 bits (42), Expect(2) = 4.2
Identities = 9/10 (90%), Positives = 9/10 (90%)
Frame = +3
Query: 1211 VVLGGDFRQI 1220
VVL GDFRQI
Sbjct: 33 VVLQGDFRQI 62
>TC7938 homologue to UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein, complete
Length = 1142
Score = 28.5 bits (62), Expect = 8.5
Identities = 26/116 (22%), Positives = 44/116 (37%), Gaps = 7/116 (6%)
Frame = +2
Query: 32 AAFSLCCLK----GKVDLPYLPKPPKLLRHLWTDEDPRSNNFKDNIRAYNSMFAFTSMGG 87
A F LC ++ G+ +PYL D R+ + D + N
Sbjct: 404 AKFKLCKVRSVQFGQKGIPYL-----------NTYDGRTIRYPDPVIRANDTIKLDLQEN 550
Query: 88 KVQNSINDGGGPPKFILSGQNYHRIGSLVPREGQTPKFAQLYIYD---HEMKI*IG 140
K+ + I G + G+N R+G + R+ Q F +++ D HE +G
Sbjct: 551 KIVDFIKFDVGNVVMVTGGRNRGRVGVIKSRDRQKGSFDTIHVQDATGHEFATRLG 718
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.145 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,798,206
Number of Sequences: 28460
Number of extensions: 405643
Number of successful extensions: 2156
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 2116
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2154
length of query: 1547
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1445
effective length of database: 1,994,680
effective search space: 2882312600
effective search space used: 2882312600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0209.3


