

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0207.3
(227 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
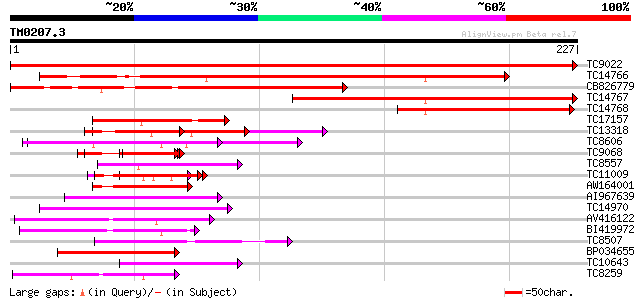
Score E
Sequences producing significant alignments: (bits) Value
TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Ara... 445 e-126
TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 213 2e-56
CB826779 187 2e-48
TC14767 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 150 2e-37
TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 75 7e-15
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 50 2e-07
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 49 5e-07
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 47 3e-06
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 46 5e-06
TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete 45 1e-05
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 44 2e-05
AW164001 44 3e-05
AI967639 43 5e-05
TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) prec... 42 9e-05
AV416122 42 1e-04
BI419972 42 1e-04
TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 40 3e-04
BP034655 40 4e-04
TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear phospho... 40 4e-04
TC8259 similar to UP|HMGL_SOYBN (P26585) HMG1/2-like protein (SB... 39 6e-04
>TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (75%)
Length = 1083
Score = 445 bits (1145), Expect = e-126
Identities = 227/227 (100%), Positives = 227/227 (100%)
Frame = +3
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD 60
MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD
Sbjct: 99 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD 278
Query: 61 DDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKP 120
DDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKP
Sbjct: 279 DDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKP 458
Query: 121 DVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEE 180
DVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEE
Sbjct: 459 DVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEE 638
Query: 181 EEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 227
EEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT
Sbjct: 639 EEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 779
>TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (58%)
Length = 600
Score = 213 bits (543), Expect = 2e-56
Identities = 116/190 (61%), Positives = 146/190 (76%), Gaps = 2/190 (1%)
Frame = +2
Query: 13 AEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
A++ E++L+A + Q DD P+VED DD +DD+DEDDDDEDDD +G
Sbjct: 65 AQSQEELLAAH-----LEQQKIHDDEPVVED---DD----EDDDDEDDDDEDDDNIEGQE 208
Query: 73 GGSEG-SKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETY 131
G + G SKQ+RSEKKSRKAMLKLG+KPVTGVSRVT+K++KNI+F ISKPDVFKSP S+TY
Sbjct: 209 GDASGRSKQTRSEKKSRKAMLKLGMKPVTGVSRVTVKKSKNIVFVISKPDVFKSPTSDTY 388
Query: 132 VIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAK-QDQGTAADGGQPEEEEEVDETGVEP 190
+IFGEAKIEDLSSQLQTQAA+QF+ P++G+V K + G A D +E +VDETGV+P
Sbjct: 389 IIFGEAKIEDLSSQLQTQAAEQFKAPNLGNVGLKPESSGVAQDEEDVDESVDVDETGVDP 568
Query: 191 HDIDLVMTQA 200
D++LVMTQA
Sbjct: 569 KDVELVMTQA 598
>CB826779
Length = 469
Score = 187 bits (474), Expect = 2e-48
Identities = 105/136 (77%), Positives = 115/136 (84%), Gaps = 1/136 (0%)
Frame = +1
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQE-DDAPIVEDVKDDDKDEADDDEDED 59
M+PG V DASS+ +Q+ SA + KKKPQPQE DDAP+VEDV D EAD+DED+D
Sbjct: 91 MSPGTVFDASSDDH--QQLPSATDD--KKKPQPQEYDDAPVVEDVHD----EADEDEDDD 246
Query: 60 DDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 119
+DDEDD EDGA G +EGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK
Sbjct: 247 EDDEDD--EDGAQGTTEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 420
Query: 120 PDVFKSPNSETYVIFG 135
PDVFKSPNSETYVIFG
Sbjct: 421 PDVFKSPNSETYVIFG 468
>TC14767 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (54%)
Length = 570
Score = 150 bits (379), Expect = 2e-37
Identities = 78/115 (67%), Positives = 93/115 (80%), Gaps = 1/115 (0%)
Frame = +1
Query: 114 LFFISKPDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAK-QDQGTAA 172
LF ISKPDVFKSP ++TY+IFGEAKIEDLSSQLQTQAA+QF+ P++ +V K + G A
Sbjct: 16 LFVISKPDVFKSPTTDTYIIFGEAKIEDLSSQLQTQAAEQFKAPNLSNVGLKPESSGVAQ 195
Query: 173 DGGQPEEEEEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 227
D +E+E+ +ETGV+P DI+LVMTQA VSR KAVKALK NGDIV AIMELTT
Sbjct: 196 DDEDEDEDEDENETGVDPKDIELVMTQATVSRSKAVKALKAANGDIVAAIMELTT 360
Score = 27.3 bits (59), Expect = 2.2
Identities = 13/42 (30%), Positives = 24/42 (56%)
Frame = +1
Query: 49 KDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKA 90
+D+ D+DEDED+++ D +D L ++ + K+ KA
Sbjct: 193 QDDEDEDEDEDENETGVDPKDIELVMTQATVSRSKAVKALKA 318
Score = 27.3 bits (59), Expect = 2.2
Identities = 9/23 (39%), Positives = 17/23 (73%)
Frame = +1
Query: 44 VKDDDKDEADDDEDEDDDDEDDD 66
+K + A DDEDED+D+++++
Sbjct: 166 LKPESSGVAQDDEDEDEDEDENE 234
>TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (21%)
Length = 1036
Score = 75.5 bits (184), Expect = 7e-15
Identities = 41/72 (56%), Positives = 51/72 (69%), Gaps = 1/72 (1%)
Frame = +2
Query: 156 MPDMGSVMAK-QDQGTAADGGQPEEEEEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTH 214
+ ++G+V K + G A D +E +VDETGV+P D++LVMTQA V R KAVKALK
Sbjct: 416 LANLGNVGLKPESSGVAQDEEDVDESVDVDETGVDPKDVELVMTQATVPRPKAVKALKAA 595
Query: 215 NGDIVGAIMELT 226
NGDIV AIMELT
Sbjct: 596 NGDIVAAIMELT 631
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 50.4 bits (119), Expect = 2e-07
Identities = 27/56 (48%), Positives = 37/56 (65%), Gaps = 1/56 (1%)
Frame = +2
Query: 34 QEDDAPIVEDVKDDDKDE-ADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSR 88
Q D I ED +D D+DE DDD+DE+++D+DDD ED G EG +S+S K+R
Sbjct: 752 QSDFEDIEEDDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEG--EGKSKSKSGSKAR 913
Score = 32.7 bits (73), Expect = 0.053
Identities = 15/64 (23%), Positives = 31/64 (48%)
Frame = +2
Query: 43 DVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGV 102
+ + D ++ ++D+++ D+DED+D +D + E+ K+ K G K G
Sbjct: 743 EAEQSDFEDIEEDDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSGSKARPGK 922
Query: 103 SRVT 106
+ T
Sbjct: 923 DQPT 934
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 49.3 bits (116), Expect = 5e-07
Identities = 26/68 (38%), Positives = 44/68 (64%), Gaps = 5/68 (7%)
Frame = +3
Query: 34 QEDDAPIVEDVKDDDKDEADDD-EDEDDDDEDDDKEDGA----LGGSEGSKQSRSEKKSR 88
+EDD ED DDD D+ DDD E+E+DDD+DD++E+G+ + G +G+K + ++
Sbjct: 276 EEDD----EDEDDDDDDDDDDDGEEEEDDDDDDEEEEGSEEVEVEGKDGAKGAAAQTTKA 443
Query: 89 KAMLKLGL 96
A + +G+
Sbjct: 444 AAAMCVGI 467
Score = 47.0 bits (110), Expect = 3e-06
Identities = 20/40 (50%), Positives = 29/40 (72%)
Frame = +3
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
P+P + D + +++D DE +DDEDEDDDD+DDD +DG
Sbjct: 228 PKPVQTD-----NEEEEDDDEEEDDEDEDDDDDDDDDDDG 332
Score = 44.7 bits (104), Expect = 1e-05
Identities = 26/97 (26%), Positives = 44/97 (44%)
Frame = +3
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKA 90
P+ D P + V+ D+++E DDDE+EDD+DEDDD +D E + + + +
Sbjct: 204 PEIYPDGPP--KPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEG 377
Query: 91 MLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPN 127
++ ++ G + TK F PN
Sbjct: 378 SEEVEVEGKDGAKGAAAQTTKAAAAMCVGIGSFSDPN 488
Score = 29.6 bits (65), Expect = 0.45
Identities = 19/72 (26%), Positives = 33/72 (45%)
Frame = +3
Query: 4 GPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDE 63
GP ++ E E E+ + +DD E+ DDD DD+E+E ++
Sbjct: 222 GPPKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEEEEDDDD----DDEEEEGSEEV 389
Query: 64 DDDKEDGALGGS 75
+ + +DGA G +
Sbjct: 390 EVEGKDGAKGAA 425
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 47.0 bits (110), Expect = 3e-06
Identities = 35/114 (30%), Positives = 57/114 (49%), Gaps = 4/114 (3%)
Frame = -2
Query: 8 DASSEAEALEQVLSAEETTLKKKPQ--PQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
D SSE A S ++ KK P+ P P E+ +D++ ++ +D ED+DDDDEDD
Sbjct: 545 DLSSEDGAGYGNNSNNKSNSKKAPEGGPGAGAGP-EENGEDEEDEDGEDQEDDDDDDEDD 369
Query: 66 DKED--GALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFI 117
D+ED G EG ++ +E + + L+P + T ++L F+
Sbjct: 368 DEEDDGGEDEEEEGVEEEDNEDEEEDEEDEEALQPPKKRKK*TTLAESSLLLFV 207
Score = 39.7 bits (91), Expect = 4e-04
Identities = 27/84 (32%), Positives = 43/84 (51%), Gaps = 4/84 (4%)
Frame = -2
Query: 6 VVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED----DD 61
VV A+ A L +L T K + + Q +D + +D + DE D+D+D D +
Sbjct: 731 VVKATLLAADLCGLLLINMATTKGETKSQSEDKQHDAEGEDGNDDEGDEDDDGDGPFGEG 552
Query: 62 DEDDDKEDGALGGSEGSKQSRSEK 85
+ED EDGA G+ + +S S+K
Sbjct: 551 EEDLSSEDGAGYGNNSNNKSNSKK 480
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 46.2 bits (108), Expect = 5e-06
Identities = 17/25 (68%), Positives = 22/25 (88%)
Frame = +2
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDG 70
DDD D+ DDD+D+DDDD+DDD +DG
Sbjct: 83 DDDDDDDDDDDDDDDDDDDDDDDDG 157
Score = 44.7 bits (104), Expect = 1e-05
Identities = 19/38 (50%), Positives = 26/38 (68%)
Frame = +2
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKE 68
P+P +DD DDD D+ DDD+D+DDDD+D DK+
Sbjct: 74 PKPDDDD-------DDDDDDDDDDDDDDDDDDDDGDKK 166
Score = 43.1 bits (100), Expect = 4e-05
Identities = 16/25 (64%), Positives = 21/25 (84%)
Frame = +2
Query: 45 KDDDKDEADDDEDEDDDDEDDDKED 69
K DD D+ DDD+D+DDDD+DDD +D
Sbjct: 77 KPDDDDDDDDDDDDDDDDDDDDDDD 151
Score = 42.7 bits (99), Expect = 5e-05
Identities = 20/42 (47%), Positives = 26/42 (61%)
Frame = +2
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
K+ P+ DD DDD D+ DDD+D+DDDD+DDD D
Sbjct: 59 KRLFSPKPDD--------DDDDDDDDDDDDDDDDDDDDDDGD 160
Score = 37.0 bits (84), Expect = 0.003
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +2
Query: 53 DDDEDEDDDDEDDDKEDGALGGSEGSKQ 80
DDD+D+DDDD+DDD +D +G K+
Sbjct: 83 DDDDDDDDDDDDDDDDDDDDDDDDGDKK 166
Score = 35.4 bits (80), Expect = 0.008
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +2
Query: 53 DDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
DDD+D+DDDD+DDD +D KQ S+
Sbjct: 86 DDDDDDDDDDDDDDDDDDDDDDDGDKKQPDSD 181
Score = 33.9 bits (76), Expect = 0.024
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +2
Query: 30 KPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDED 64
KP +DD +D DDD D+ DDD D+ D D
Sbjct: 77 KPDDDDDDDDDDDDDDDDDDDDDDDDGDKKQPDSD 181
Score = 26.6 bits (57), Expect = 3.8
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = +2
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD 62
K +DD +D DDD D+ D D+ + D D
Sbjct: 77 KPDDDDDDDDDDDDDDDDDDDDDDDDGDKKQPDSD 181
>TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete
Length = 1064
Score = 44.7 bits (104), Expect = 1e-05
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 1/59 (1%)
Frame = +3
Query: 36 DDAPIVEDVKDDDKD-EADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
+D + V DD D E DE+ DDDD+D+D + L E K+ R+E+K RK L+
Sbjct: 450 EDHIVARSVDADDSDVEVKSDEESDDDDDDEDDTEALLAELEQIKKERAEEKLRKERLE 626
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 43.9 bits (102), Expect = 2e-05
Identities = 22/47 (46%), Positives = 31/47 (65%), Gaps = 4/47 (8%)
Frame = +2
Query: 35 EDDAPIVEDVKDDDKDEADDDE----DEDDDDEDDDKEDGALGGSEG 77
EDD +D +DDD DE DDD+ +DDDDE+D++E+G + G G
Sbjct: 29 EDDD---DDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEEGGVEGGRG 160
Score = 40.0 bits (92), Expect = 3e-04
Identities = 20/57 (35%), Positives = 32/57 (56%), Gaps = 15/57 (26%)
Frame = +2
Query: 32 QPQEDDAPIVEDVKDDDKDEA---------------DDDEDEDDDDEDDDKEDGALG 73
+ +DDAP D DD++DE DDD+D+DDDD+++++E+ LG
Sbjct: 68 EDDDDDAPGGGDDDDDEEDEEEEGGVEGGRGGGGDPDDDDDDDDDDDEEEEEEEDLG 238
Score = 39.7 bits (91), Expect = 4e-04
Identities = 19/43 (44%), Positives = 26/43 (60%), Gaps = 8/43 (18%)
Frame = +2
Query: 45 KDDDKDEADDDEDEDDDDE--------DDDKEDGALGGSEGSK 79
+DDD D DDD DEDDDD+ DD++++ GG EG +
Sbjct: 29 EDDDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEEGGVEGGR 157
Score = 38.1 bits (87), Expect = 0.001
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +2
Query: 50 DEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKK 86
DE DDD+ EDDD ++DD +D GG + + E++
Sbjct: 26 DEDDDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEE 136
Score = 36.6 bits (83), Expect = 0.004
Identities = 18/46 (39%), Positives = 28/46 (60%), Gaps = 4/46 (8%)
Frame = +2
Query: 35 EDDAPIVEDVKDDDKDEA----DDDEDEDDDDEDDDKEDGALGGSE 76
+DD +D +DD D+A DDD+DE+D++E+ E G GG +
Sbjct: 35 DDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEEGGVEGGRGGGGD 172
Score = 33.5 bits (75), Expect = 0.031
Identities = 15/39 (38%), Positives = 26/39 (66%)
Frame = +2
Query: 47 DDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEK 85
+D++ + D E E+D+ +D+D +DG G SK+ RS+K
Sbjct: 275 EDEEASSDFEPEEDEGDDNDNDDGEKAGVP-SKRKRSDK 388
Score = 31.6 bits (70), Expect = 0.12
Identities = 14/41 (34%), Positives = 23/41 (55%)
Frame = +2
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKK 86
D+D D+ +D+D D+DD+DD A GG + E++
Sbjct: 26 DEDDDDDGEDDDGDEDDDDD-----APGGGDDDDDEEDEEE 133
Score = 31.6 bits (70), Expect = 0.12
Identities = 15/39 (38%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Frame = +2
Query: 48 DKDEADDDEDED-DDDEDDDKEDGALGGSEGSKQSRSEK 85
D+D+ DD ED+D D+D+DDD G G + ++ E+
Sbjct: 26 DEDDDDDGEDDDGDEDDDDDAPGG--GDDDDDEEDEEEE 136
Score = 30.0 bits (66), Expect = 0.34
Identities = 21/71 (29%), Positives = 31/71 (43%), Gaps = 11/71 (15%)
Frame = +2
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDD-----------APIVEDVKDDDKDEADDDE 56
D +E V +AE+ +P+ED+ A + K DKD + DD
Sbjct: 230 DLGTEYLVRRTVAAAEDEEASSDFEPEEDEGDDNDNDDGEKAGVPSKRKRSDKDGSGDD- 406
Query: 57 DEDDDDEDDDK 67
D DD EDD++
Sbjct: 407 DSDDGGEDDER 439
>AW164001
Length = 447
Score = 43.5 bits (101), Expect = 3e-05
Identities = 21/40 (52%), Positives = 26/40 (64%)
Frame = +1
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALG 73
Q DD E K++D DE DD+D+DDDD+DDD E LG
Sbjct: 43 QNDD----EVYKEEDVDEDFDDDDDDDDDDDDDDEPATLG 150
Score = 28.1 bits (61), Expect = 1.3
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = +1
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKED 69
+ V + DE +ED D+D +DDD +D
Sbjct: 28 DSVNKQNDDEVYKEEDVDEDFDDDDDDD 111
Score = 26.2 bits (56), Expect = 5.0
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +1
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
+ED +D DDD D+ DDDE D K++ +L
Sbjct: 67 EEDVDEDFDDDDDDDDDDDDDDEPATLGFVDKPKDEWSL 183
>AI967639
Length = 335
Score = 42.7 bits (99), Expect = 5e-05
Identities = 20/63 (31%), Positives = 37/63 (57%)
Frame = +1
Query: 23 EETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSR 82
+E LKK+ Q ++ + D +D+++D+ D+ ED++++DDD +D L SE S +
Sbjct: 43 DEEYLKKRKQRRDISSSSEGDEEDEEEDQGDEYNLEDEEEDDDDDDDDDLSISEDSDSDK 222
Query: 83 SEK 85
K
Sbjct: 223 PRK 231
Score = 26.6 bits (57), Expect = 3.8
Identities = 20/73 (27%), Positives = 36/73 (48%), Gaps = 3/73 (4%)
Frame = +1
Query: 19 VLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDED---EDDDDEDDDKEDGALGGS 75
+ S+ E + + + Q D+ + ++ +DDD DDD+D +D D D ++ L G
Sbjct: 82 ISSSSEGDEEDEEEDQGDEYNLEDEEEDDDD---DDDDDLSISEDSDSDKPRKVKQLPG- 249
Query: 76 EGSKQSRSEKKSR 88
++R E K R
Sbjct: 250 ----RTRRETKRR 276
>TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) precursor,
partial (51%)
Length = 1566
Score = 42.0 bits (97), Expect = 9e-05
Identities = 23/77 (29%), Positives = 39/77 (49%)
Frame = +1
Query: 13 AEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
AE L+QV + + T K+ P + IV+ DD + + DED+D+ED+D++
Sbjct: 373 AEFLQQVFNIDHDTAKQLQSPDDQRRQIVKVEGDDLSFISPESADEDEDEEDEDEDQEQH 552
Query: 73 GGSEGSKQSRSEKKSRK 89
G R+ + SR+
Sbjct: 553 GRHSRPSGRRTRRPSRE 603
>AV416122
Length = 414
Score = 41.6 bits (96), Expect = 1e-04
Identities = 22/81 (27%), Positives = 47/81 (57%), Gaps = 1/81 (1%)
Frame = +3
Query: 3 PGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDED-EDDD 61
P P D++ V + ++ + + K +D+ P DV+ + ++E DD+ED E++D
Sbjct: 168 PEPNPDSNDNNNNHPAVSATDDCSSQPKEAANDDNEPD-NDVQFEGEEEEDDEEDKEEED 344
Query: 62 DEDDDKEDGALGGSEGSKQSR 82
+E+DD++D + G +E ++ +
Sbjct: 345 EEEDDEDDDSNGEAEVDRKGK 407
Score = 25.8 bits (55), Expect = 6.5
Identities = 17/77 (22%), Positives = 30/77 (38%), Gaps = 17/77 (22%)
Frame = +3
Query: 30 KPQPQEDD-------APIVEDVKDDDKDEADDDED----------EDDDDEDDDKEDGAL 72
+P P +D +D K+ A+DD + E++DDE+D +E+
Sbjct: 171 EPNPDSNDNNNNHPAVSATDDCSSQPKEAANDDNEPDNDVQFEGEEEEDDEEDKEEEDEE 350
Query: 73 GGSEGSKQSRSEKKSRK 89
E + + RK
Sbjct: 351 EDDEDDDSNGEAEVDRK 401
>BI419972
Length = 498
Score = 41.6 bits (96), Expect = 1e-04
Identities = 29/80 (36%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Frame = +1
Query: 5 PVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED----- 59
P +++ +LE +L + L K EDD D +DDD D+ DDDE ED
Sbjct: 265 PKLNSKDGRFSLEDLLGRKPAYLANKDSETEDDED--GDDEDDDADDDDDDEGEDYSGDE 438
Query: 60 ---DDDEDDDKEDGALGGSE 76
D EDD + +GA GGS+
Sbjct: 439 GEEADPEDDPEANGA-GGSD 495
Score = 35.0 bits (79), Expect = 0.011
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +1
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
+DD+D D+D+D DDDD+D+ ++ G E + E
Sbjct: 355 EDDEDGDDEDDDADDDDDDEGEDYSGDEGEEADPEDDPE 471
>TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (40%)
Length = 620
Score = 40.0 bits (92), Expect = 3e-04
Identities = 26/79 (32%), Positives = 38/79 (47%)
Frame = +3
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKL 94
E+D ED +DDD DE DD D D D+ D D D + K + +K++ + LK
Sbjct: 6 EEDDSDDEDDEDDDSDEEMDDADSDSDESDSDDTDEE---TPVKKADQGKKRANDSALK- 173
Query: 95 GLKPVTGVSRVTIKRTKNI 113
+ V+ K+ KNI
Sbjct: 174 --------TPVSTKKAKNI 206
Score = 33.9 bits (76), Expect = 0.024
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = +3
Query: 48 DKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEK 85
D+++ DDED++DDD D++ +D E E+
Sbjct: 3 DEEDDSDDEDDEDDDSDEEMDDADSDSDESDSDDTDEE 116
>BP034655
Length = 517
Score = 39.7 bits (91), Expect = 4e-04
Identities = 14/49 (28%), Positives = 35/49 (70%)
Frame = +1
Query: 20 LSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKE 68
L+A + L + +E++ E+ ++++++E +++EDE+DD+ED+D++
Sbjct: 109 LTAADFGLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDED 255
Score = 37.4 bits (85), Expect = 0.002
Identities = 15/60 (25%), Positives = 37/60 (61%)
Frame = +1
Query: 10 SSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+S +E ++A T +D+ E+ ++++++E +++E+E+++DE+DD+ED
Sbjct: 64 ASPSEFHRNEIAAAHLTAADFGLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEED 243
Score = 36.2 bits (82), Expect = 0.005
Identities = 15/53 (28%), Positives = 35/53 (65%)
Frame = +1
Query: 9 ASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDD 61
A++ A + LS ++ +++ + +E++ E+ +++++DE DD+EDED+D
Sbjct: 97 AAAHLTAADFGLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDED 255
>TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear
phosphoprotein-like protein, partial (5%)
Length = 789
Score = 39.7 bits (91), Expect = 4e-04
Identities = 17/49 (34%), Positives = 29/49 (58%)
Frame = +2
Query: 45 KDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
+D+D++E +D E+ DDDED+ E G+ G ++ + + SRK K
Sbjct: 41 EDEDEEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDNGRSSRKRTRK 187
Score = 36.2 bits (82), Expect = 0.005
Identities = 17/46 (36%), Positives = 28/46 (59%), Gaps = 3/46 (6%)
Frame = +2
Query: 51 EADDDEDEDDDDE---DDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
E D+DE+E++D E DDD+++ A GGS G+ E + ++ K
Sbjct: 38 EEDEDEEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDNGRSSRK 175
Score = 28.5 bits (62), Expect = 1.00
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 10/55 (18%)
Frame = +2
Query: 22 AEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED----------DDDEDDD 66
AE + K +ED+ E+ ++D ++ +DDDEDE D+DE+D+
Sbjct: 2 AEGLRMMKLGSGEEDED---EEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDN 157
>TC8259 similar to UP|HMGL_SOYBN (P26585) HMG1/2-like protein (SB11
protein), partial (92%)
Length = 942
Score = 39.3 bits (90), Expect = 6e-04
Identities = 28/76 (36%), Positives = 37/76 (47%), Gaps = 9/76 (11%)
Frame = +2
Query: 2 APGPVVDASSEAEALEQVLSAE------ETTLKKKPQPQEDDAPIVEDVKDDDKDEA--- 52
A G + SEAE V AE E T+K + Q + P D ++ +K +
Sbjct: 407 AAGAKWKSMSEAEKAPYVAKAEKRKADYEKTMKAYNKKQAE-GPAAADEEESEKSVSEVN 583
Query: 53 DDDEDEDDDDEDDDKE 68
D DED+DD DEDDD E
Sbjct: 584 DQDEDDDDSDEDDDDE 631
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.305 0.127 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,409,239
Number of Sequences: 28460
Number of extensions: 28338
Number of successful extensions: 2058
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 517
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1117
length of query: 227
length of database: 4,897,600
effective HSP length: 87
effective length of query: 140
effective length of database: 2,421,580
effective search space: 339021200
effective search space used: 339021200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0207.3


