

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.9
(529 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
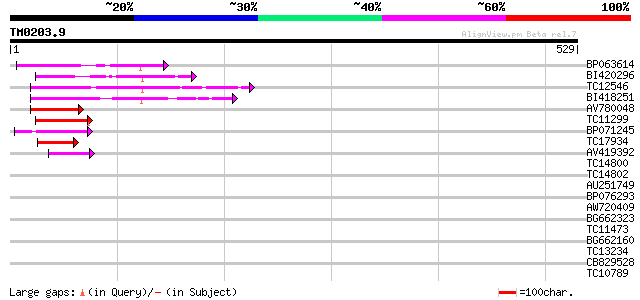
Score E
Sequences producing significant alignments: (bits) Value
BP063614 73 1e-13
BI420296 70 1e-12
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 66 2e-11
BI418251 60 1e-09
AV780048 59 3e-09
TC11299 57 6e-09
BP071245 55 3e-08
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 47 6e-06
AV419392 41 4e-04
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 36 0.014
TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (... 36 0.018
AU251749 30 0.023
BP076293 34 0.052
AW720409 32 0.26
BG662323 30 0.75
TC11473 30 0.98
BG662160 30 1.3
TC13234 29 2.2
CB829528 29 2.2
TC10789 28 2.8
>BP063614
Length = 508
Score = 72.8 bits (177), Expect = 1e-13
Identities = 50/144 (34%), Positives = 69/144 (47%), Gaps = 2/144 (1%)
Frame = +2
Query: 7 KRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPD 66
K K+ Q + DR+SDLPD +L HILS + K A QT ILS RWK LW P
Sbjct: 113 KAKRGRQSLNENENEDNKDRLSDLPDCILLHILSFVKAKAAVQTCILSTRWKDLWKRLPS 292
Query: 67 LDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFR--ARLS 124
L L SS+F T S F++++L+ RD + + L F +
Sbjct: 293 L-------------ILLSSNFWT----YKSFTKFVSRLLTLRDGSTALHGLDFEHDGHIQ 421
Query: 125 FSRLNSLIRSAVRHNVKELDIEVT 148
L +++ AV HNV+ L + VT
Sbjct: 422 PHLLKRIVKYAVSHNVQRLGLSVT 493
>BI420296
Length = 592
Score = 69.7 bits (169), Expect = 1e-12
Identities = 53/152 (34%), Positives = 76/152 (49%), Gaps = 2/152 (1%)
Frame = +2
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
DR+SDLPD VL H+L + K A QT +LS RWK LW ++ L S
Sbjct: 194 DRLSDLPDLVLLHVLKFMSTKQAVQTCVLSTRWKDLW-------------KGVTTLALNS 334
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRAR--LSFSRLNSLIRSAVRHNVKE 142
SDF T+ P S +F++ VLS R+ + L R + + LN ++ AV H+V+
Sbjct: 335 SDFATA--PRFS--EFLSCVLSHRNDSVSLHNLDLRRKGCVEPELLNRVMSYAVSHDVQS 502
Query: 143 LDIEVTPEVRTDEYFNFPRCLIASKTLRVLKL 174
L IE ++ F C+ + +TL LKL
Sbjct: 503 LTIEFNLYLKLG--FKLHPCIFSCRTLTYLKL 592
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 65.9 bits (159), Expect = 2e-11
Identities = 62/216 (28%), Positives = 95/216 (43%), Gaps = 7/216 (3%)
Frame = +2
Query: 20 KAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSS 79
K D IS LPDA+L HILS L K A TS+LS+RW LW + P L F+ + H
Sbjct: 134 KMKMEDGISTLPDALLCHILSFLTSKEAVATSVLSRRWIPLWRSVPTLHFKDANYHT--- 304
Query: 80 RNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRAR------LSFSRLNSLIR 133
++ +D + + + + V+ +RD I+ R + ++ +
Sbjct: 305 -DIGHADHDIVKDVRSRHVQSVYAVILSRDFQLPIKKFYLRLNDVCQPFYDPANVSVWVN 481
Query: 134 SAVRHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQSL 193
+ V+ ++ LDI + + + N + + +TL VLKLR G L F L
Sbjct: 482 AVVQRQLEHLDISLPYPMLSTPRANL-SSIFSCRTLVVLKLRGGLEL--KRFPSVHFPCL 652
Query: 194 HTLSLTLGPVDHQAP-LSDLFTESAFPFLRNLHLQL 228
L L + H P L++L S P L NL L
Sbjct: 653 KVLHLQGALLLHDVPYLAELL--SGCPVLENLKQSL 754
>BI418251
Length = 546
Score = 59.7 bits (143), Expect = 1e-09
Identities = 58/196 (29%), Positives = 85/196 (42%), Gaps = 3/196 (1%)
Frame = +1
Query: 20 KAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSS 79
+A DRIS LPD VL ILS LP + A TS+L KRW LW + P LDF
Sbjct: 22 RAPMADRISKLPDEVLCQILSFLPTEDAVATSVLCKRWSSLWLSVPTLDFDDYRYLKGKK 201
Query: 80 RNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRA---RLSFSRLNSLIRSAV 136
LQS S ++F+ ++ +R I+ + + + +A+
Sbjct: 202 LKLQS-----------SFINFVYAIILSRALHQPIKNFTLSVISEECPYPDVKVWLNAAM 348
Query: 137 RHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQSLHTL 196
+ V+ LDI ++ P +++ TL VLKL S + S + SL TL
Sbjct: 349 QRQVENLDICLS-------LSTLPCSILSCTTLVVLKL-SDVKFHVFSCV--DLPSLKTL 498
Query: 197 SLTLGPVDHQAPLSDL 212
L L + + L DL
Sbjct: 499 HLELVVILNPQSLMDL 546
>AV780048
Length = 501
Score = 58.5 bits (140), Expect = 3e-09
Identities = 30/50 (60%), Positives = 32/50 (64%)
Frame = +2
Query: 20 KAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF 69
K DR S LPD +L HILSLL K A TSILSKRW LW + P LDF
Sbjct: 236 KMKMEDRFSSLPDPILCHILSLLTTKEAVTTSILSKRWIPLWRSVPTLDF 385
>TC11299
Length = 498
Score = 57.4 bits (137), Expect = 6e-09
Identities = 28/58 (48%), Positives = 36/58 (61%), Gaps = 5/58 (8%)
Frame = +1
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTI-----DPHAF 77
DRIS LPD ++ HILS LP + A T++LSKRW LW + LDF ++ D H F
Sbjct: 85 DRISALPDEIICHILSFLPTENAVATAVLSKRWTHLWRSVSALDFSSVRMYEPDDHRF 258
>BP071245
Length = 552
Score = 55.1 bits (131), Expect = 3e-08
Identities = 34/74 (45%), Positives = 41/74 (54%), Gaps = 1/74 (1%)
Frame = +1
Query: 5 SAKRKKMAQI-MENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTT 63
+AKR + ++ ENE DR+SDLPD VL HILS + K A QT LS RWK LW
Sbjct: 328 NAKRGRESESESENEENK---DRLSDLPDCVLLHILSFVNAKYAVQTCTLSTRWKDLWKR 498
Query: 64 FPDLDFRTIDPHAF 77
P L + D F
Sbjct: 499 LPSLILHSSDFSTF 540
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 47.4 bits (111), Expect = 6e-06
Identities = 20/38 (52%), Positives = 29/38 (75%)
Frame = -1
Query: 27 ISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTF 64
I+ LPD + ILS LPI AA+TSILS++W+++WT +
Sbjct: 564 INQLPDGIPGAILSKLPINEAAKTSILSRKWRYMWTFY 451
>AV419392
Length = 318
Score = 41.2 bits (95), Expect = 4e-04
Identities = 20/43 (46%), Positives = 26/43 (59%)
Frame = +3
Query: 37 HILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSS 79
HILS + K A Q S+LSKRW+ LW P L+F + +SS
Sbjct: 3 HILSYVETKDAMQCSVLSKRWRSLWRALPVLNFDSASFTHYSS 131
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5, TAC
clone:K14B20, partial (7%)
Length = 782
Score = 36.2 bits (82), Expect = 0.014
Identities = 27/84 (32%), Positives = 42/84 (49%), Gaps = 1/84 (1%)
Frame = +3
Query: 30 LPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQSSDFE- 88
LPD ++ ILS LP+K+ Q ++SK WK + D + + H + +++DFE
Sbjct: 48 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFIS-----DPQFVKLHLLHRLSFRNADFEH 212
Query: 89 TSGHPLASTMDFITQVLSTRDKDS 112
TS T DF +S+R S
Sbjct: 213 TSLLIKCHTDDFGRPYISSRTVSS 284
>TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (SLF)-S4
protein, partial (10%)
Length = 551
Score = 35.8 bits (81), Expect = 0.018
Identities = 26/80 (32%), Positives = 41/80 (50%), Gaps = 1/80 (1%)
Frame = +3
Query: 30 LPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQSSDFE- 88
LPD ++ ILS LP+K+ Q ++SK WK + D + + H + +++DFE
Sbjct: 21 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFIS-----DPQFVKLHLLHRLSFRNADFEH 185
Query: 89 TSGHPLASTMDFITQVLSTR 108
TS T DF +S+R
Sbjct: 186 TSLLIKCHTDDFGRPYISSR 245
>AU251749
Length = 362
Score = 30.4 bits (67), Expect(2) = 0.023
Identities = 15/27 (55%), Positives = 17/27 (62%)
Frame = +2
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTS 51
D IS LPD +L+ ILS LP K TS
Sbjct: 137 DWISALPDTILNFILSFLPTKQVVATS 217
Score = 23.9 bits (50), Expect(2) = 0.023
Identities = 10/20 (50%), Positives = 11/20 (55%)
Frame = +1
Query: 49 QTSILSKRWKFLWTTFPDLD 68
Q S KRWK LW + P D
Sbjct: 196 QASRCHKRWKPLWRSVPTRD 255
>BP076293
Length = 485
Score = 34.3 bits (77), Expect = 0.052
Identities = 14/47 (29%), Positives = 26/47 (54%)
Frame = -2
Query: 446 WESEIPTLKPFLQHLKVVEIHGFLDCENEVTLAKFLLKNGKALEEMV 492
W+ +P + HLK ++ + E E A+++L+NG+ LE M+
Sbjct: 463 WQCSLPGPECIQLHLKRCYLNDYRGTEGEFQFARYILQNGRVLESMI 323
>AW720409
Length = 575
Score = 32.0 bits (71), Expect = 0.26
Identities = 18/43 (41%), Positives = 25/43 (57%)
Frame = +3
Query: 30 LPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTI 72
LP + ILS LP+KT + S +SK WK L + D DF+ +
Sbjct: 93 LPFELWIEILSWLPVKTLMRFSCVSKSWKSL--IYQDRDFKKL 215
>BG662323
Length = 375
Score = 30.4 bits (67), Expect = 0.75
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = +1
Query: 16 ENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWT 62
E E + D + LP +L HIL LPI + + + KRW+ + T
Sbjct: 40 EKEGTIVSVDEV--LPGDLLEHILMYLPIASIFRACCVCKRWREIVT 174
>TC11473
Length = 435
Score = 30.0 bits (66), Expect = 0.98
Identities = 12/46 (26%), Positives = 26/46 (56%)
Frame = +2
Query: 446 WESEIPTLKPFLQHLKVVEIHGFLDCENEVTLAKFLLKNGKALEEM 491
W+ K L HLKV ++ + + E+ A++++++G+ L+ M
Sbjct: 62 WQDPPSVPKCILLHLKVCYLNDYRGTKGELQFARYIMRHGRFLKRM 199
>BG662160
Length = 445
Score = 29.6 bits (65), Expect = 1.3
Identities = 20/62 (32%), Positives = 31/62 (49%)
Frame = +2
Query: 26 RISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQSS 85
R LP+ + ILS LP+K+ + +SK WK + D F + H SS +++
Sbjct: 11 RAQFLPEELRLEILSWLPVKSLVRFRCVSKSWK---SIISDSQFIKLHLHR-SSSTTRNT 178
Query: 86 DF 87
DF
Sbjct: 179DF 184
>TC13234
Length = 609
Score = 28.9 bits (63), Expect = 2.2
Identities = 12/46 (26%), Positives = 25/46 (54%)
Frame = +3
Query: 446 WESEIPTLKPFLQHLKVVEIHGFLDCENEVTLAKFLLKNGKALEEM 491
W+ K L HLK ++ + + E A+++++NG+ L++M
Sbjct: 231 WQYPSSVPKCILFHLKECYLNNYRGTKGEFKFARYIMRNGRFLKKM 368
>CB829528
Length = 188
Score = 28.9 bits (63), Expect = 2.2
Identities = 22/48 (45%), Positives = 26/48 (53%), Gaps = 3/48 (6%)
Frame = +2
Query: 209 LSDLFTESAFPFLRNLHLQLCFGLRVLR--VGCLA-LRDLSLEKCINL 253
L++L SA PFLRNL L C L + VG L LR LS + C L
Sbjct: 5 LTELPNLSAAPFLRNLSLDNCPSLVTIHESVGFLENLRSLSAKGCTQL 148
>TC10789
Length = 584
Score = 28.5 bits (62), Expect = 2.8
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Frame = +3
Query: 227 QLCFGLRVLRVGCLALRDL---SLEKCINLHGLDVVTPKLERLR 267
Q CF R L L +R L +C+N H D+ PK++RLR
Sbjct: 195 QHCFFSRTLLCMLLTVRSLLGLRCSQCLNGHPRDLT*PKMKRLR 326
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.134 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,705,504
Number of Sequences: 28460
Number of extensions: 139297
Number of successful extensions: 774
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 772
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 774
length of query: 529
length of database: 4,897,600
effective HSP length: 94
effective length of query: 435
effective length of database: 2,222,360
effective search space: 966726600
effective search space used: 966726600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0203.9


