

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
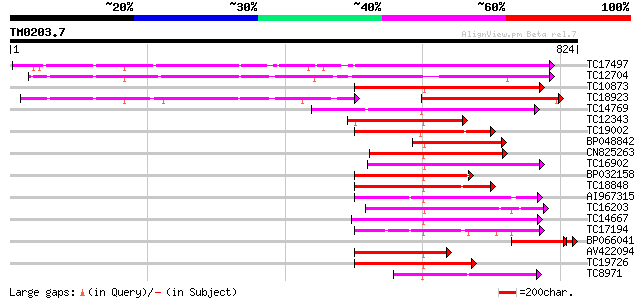
Query= TM0203.7
(824 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 511 e-145
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 495 e-140
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 234 5e-62
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 233 7e-62
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 205 2e-53
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 197 4e-51
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 193 1e-49
BP048842 192 1e-49
CN825263 189 2e-48
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 186 2e-47
BP032158 184 4e-47
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 180 7e-46
AI967315 178 4e-45
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 176 1e-44
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 173 1e-43
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 173 1e-43
BP066041 159 4e-43
AV422094 166 2e-41
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 156 1e-38
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 153 1e-37
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 511 bits (1316), Expect = e-145
Identities = 319/827 (38%), Positives = 459/827 (54%), Gaps = 38/827 (4%)
Frame = +1
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGG---------EDTLISAG-----GYFELGFF 49
+F F F++ +C+ ++T+ ++ G E +S G G FE GFF
Sbjct: 118 VFHSIIFSFSYVVCNGNGSLTLEDSSWKSGNYRCSSKKYEQCTLSQGMTVHDGTFEAGFF 297
Query: 50 TPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGG-AFSITEDGNLRVLDRTGKSF 108
+ + Y G+ Y ++P+T+VWVANRD PL +S +T G++ + D
Sbjct: 298 --HFENPQHHYFGVWYKSISPRTIVWVANRDAPLRNSTAPTLKVTHKGSILIRDGAKGVI 471
Query: 109 WGTNLERTSSSKNMTVKLLDSGNLIVSD-DKVKKILWQSFANPTDTFLPGMKMDDSIT-- 165
W TN R M +LLDSGNL+ D DK + ++W+SF P DTFL GMK+ ++
Sbjct: 472 WSTNTSRAKEQPFM--QLLDSGNLVAKDGDKGENVIWESFNYPGDTFLAGMKIKSNLAIG 645
Query: 166 ----LTSWRSHDDPAPGNFSFEQD-QGENQFVIWKRSMKYWKSS--VSGKFVGTGEMSSA 218
LTSWR+ +DPA G FS+ D +G Q V+ K + ++ KF +G
Sbjct: 646 PTSYLTSWRNSEDPASGEFSYHIDIRGFPQLVVTKGAAITLRAGPWTGNKF--SGAFGQV 819
Query: 219 ISYLLSNFTLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQYLKMDSMKM-WLMVWVEPRD 277
+ +L+ F ++ + T + TR V+T G +Q L W ++ P D
Sbjct: 820 LQKILTFFMQFTDQEISLEYETVNRSIITREVITPLGTIQRLLWSVRNQSWEIIATRPVD 999
Query: 278 RCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDT 337
+C+ + CG C++ + +C CL GF P W++ D++GGC + ++ D
Sbjct: 1000QCADYVFCGANSLCDTSKNPICDCLEGFMPQFQAKWNSLDWAGGCVSMEKLSCQNG--DG 1173
Query: 338 FLSLRMMKVGNPDAQFNAKNE--EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICW 395
F+ +K+ + + + KN +EC+ CL NC C AY A + VD ++C
Sbjct: 1174FMKHTGVKLPDTSSSWFGKNMSLDECRTLCLQNCSCTAY-------AGLDNDVD-RSVCL 1329
Query: 396 IWSQDLNNLEE--EYEGGCDLHVRVAFSDIEGDSYQTERN---LSSPTIILVTFTAVVFL 450
IW D+ ++ + + + G ++++RV S ++ + N L+ ++++ F V+F+
Sbjct: 1330IWFGDILDMSKHPDPDQGQEIYIRVVASKLDRTRNKKSINTKKLAGSLVVIIAF--VIFI 1503
Query: 451 IILS---STCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESI 507
IL STCI ++ ++ I + + R ++D I F +I
Sbjct: 1504TILGLAISTCIQRKKNKRGDEGEIGIINHWKDKRG--------DEDIDLATI--FDFSTI 1653
Query: 508 LDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQH 567
ATN+F+++NKLG+GGFGPVYKG GQEIAVKRLS+ SGQG+EEFKNE+ LIARLQH
Sbjct: 1654SSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKLIARLQH 1833
Query: 568 RNLVRLLGYCVEGDEKMLVYEYMPNR--SLDAFIFDWDMRFKIILGIARGLLYLHEDSRL 625
RNLV+L G V DE + M S + + DW+ R +II GIARGLLYLH+DSRL
Sbjct: 1834RNLVKLFGCSVHQDENSHANKKMKILLDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRL 2013
Query: 626 RIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHF 685
RIIHRDLK SNILLD+E NPKISDFGLARIF G T RV+GTYGYM PEYA+ G F
Sbjct: 2014RIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSF 2193
Query: 686 SVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCK 745
S+KSDVFSFGV+VLEIISGK+ FY P H L+LL +AW LW R L+ +D+ L
Sbjct: 2194SIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERPLELVDELLDDPVI 2373
Query: 746 AEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAF 792
E ++ ++V LLC+Q P RP M +++ ML E LP PR PAF
Sbjct: 2374PTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKE-LPKPRLPAF 2511
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 495 bits (1274), Expect = e-140
Identities = 314/809 (38%), Positives = 447/809 (54%), Gaps = 44/809 (5%)
Frame = +1
Query: 28 NLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSG 87
++QD ++TL+S G FE GFF S RRY GI Y ++P+T+VWVANRD P+ +S
Sbjct: 16 SIQD--DETLVSPEGTFEAGFFRFGNSL--RRYFGIWYKSISPRTIVWVANRDAPVQNST 183
Query: 88 GAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSD-DKVKKILWQS 146
+T+ GNL +LD W +N RT M +LLDSGN +V D DK + ++W+S
Sbjct: 184 ATLKLTDQGNLLILDGLKGIVWSSNASRTKDKPLM--QLLDSGNFVVKDGDKEENLIWES 357
Query: 147 FANPTDTFLPGMKMDDSIT------LTSWRSHDDPAPGNFSFEQD-QGENQFVIWKRSMK 199
F P DTFL GMK+ ++ LTSWR+ +DPA G FS+ D G Q V+ K +
Sbjct: 358 FDYPGDTFLAGMKIKSNLATGPTSYLTSWRNAEDPASGEFSYHIDTHGYPQLVVTKGATV 537
Query: 200 YWKSS--VSGKFVGTGEMSSAISYLLSNFTLRISPNN-TIPFLTSSLYSNTRLVMTYWGQ 256
++ + KF G + + +L+ F+++ + ++ + T++ TR V+T G
Sbjct: 538 TLRAGPWIGNKFSGASGLR--LQKILT-FSMQFTDKEVSLEYETANRSIITRTVITPSGT 708
Query: 257 LQYLKM-DSMKMWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSA 315
Q L D + W ++ P D+C+ + CG C++ + +C CL GF P W++
Sbjct: 709 TQRLLWSDRSQSWEIISTHPMDQCAYYAFCGANSMCDTSNNPICDCLEGFTPKFQAQWNS 888
Query: 316 GDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQF--NAKNEEECKLECLNNCQCYA 373
D++GGC N+ ++ D F ++ + + + N+K+ +EC CL NC C A
Sbjct: 889 LDWTGGCVPIKNLSCQNG--DGFPKHTGVQFPDTSSSWYGNSKSLDECGTICLQNCSCTA 1062
Query: 374 YSYEEYEKARQGDSVDPNAICWIWSQDLNNLEE--EYEGGCDLHVRVAFSDIEGDSYQTE 431
Y+Y D+V ++C W D+ ++ E + + G ++++RV S+++ +
Sbjct: 1063YAYL--------DNVGGRSVCLNWFGDILDMSEHPDPDQGQEIYLRVVASELDHRRNKKS 1218
Query: 432 RNLSSPTIIL---VTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGR 488
N+ L + F + ++ L++ R++ + + G G E + + + R
Sbjct: 1219INIKKLAGSLAGSIAFIICITILGLATVTCIRRKKNEREDEG----GIETRIINHWKDKR 1386
Query: 489 LQEDDAKAIDIPH-FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSC 547
ED ID+ F +I TN+F+ +NKLG+GGFGPVYKG GQEIAVKRLS+
Sbjct: 1387GDED----IDLATIFDFSTISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNT 1554
Query: 548 SGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFDWDMRFK 607
SGQG+EEFKNEV LIARLQHRNLV+LLG + DE +L+YE+M NRSLD FIFD
Sbjct: 1555SGQGMEEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLLIYEFMHNRSLDYFIFD------ 1716
Query: 608 IILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDR 667
SRLRIIHRDLK SNILLD E NPKISDFGLARIF G T R
Sbjct: 1717---------------SRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKR 1851
Query: 668 VVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGY----- 722
V+GTYGYMSPEYA+ G FSVKSDVFSFGV+VLEIISGK+ F P H +LL +
Sbjct: 1852VMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFA 2031
Query: 723 -------------------AWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQED 763
AW LW R L+ +D+ L E ++ +++ LLC+Q+
Sbjct: 2032VFLIKALRICMFENVKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLCVQQR 2211
Query: 764 PIERPTMSNVLFMLGSESNTLPSPREPAF 792
P RP M +V+ ML E LP P PAF
Sbjct: 2212PEYRPDMLSVVLMLNGEKE-LPKPSLPAF 2295
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 234 bits (596), Expect = 5e-62
Identities = 122/286 (42%), Positives = 181/286 (62%), Gaps = 10/286 (3%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F + DAT NF AN +G+GGFG VYKG+ G+ +AVK+LS QG +EF EV++
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD---------WDMRFKIILGI 612
++ L H NLVRL+GYC +GD+++LVYEYMP SL+ +F+ W R K+ +G
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
ARGL YLH + +I+RDLK++NILLD E NPK+SDFGLA++ D + RV+GTY
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVSTRVMGTY 954
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRT 732
GY +PEYA+ G ++KSD++SFGVV+LE+++G+R + E +L+ +A + R
Sbjct: 955 GYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRRR 1134
Query: 733 LDFIDQTLSQTCKAEECM-KCVNVGLLCLQEDPIERPTMSNVLFML 777
+ L Q C+ + + + +CLQE P RP +++++ L
Sbjct: 1135FGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIVVAL 1272
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 233 bits (595), Expect = 7e-62
Identities = 120/214 (56%), Positives = 145/214 (67%), Gaps = 7/214 (3%)
Frame = +1
Query: 599 IFDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGG 658
+ DW+ R +II GIARGLLYLH+DSRLRIIHRDLK SNILLD E NPKISDFGLARIF G
Sbjct: 1396 LLDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFGLARIFIG 1575
Query: 659 KDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELS 718
T RV+GTYGYM PEYA+ G FS+KSDVFSFGV+VLEIISGK+ FY P H L+
Sbjct: 1576 DQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFYDPHHHLN 1755
Query: 719 LLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLG 778
LL +AW LW G L+ +D+ + E ++ ++V LLC+Q P RP M +++ ML
Sbjct: 1756 LLSHAWRLWIEGSPLELVDKLFEDSIIPTEILRYIHVALLCVQRRPETRPDMLSIVLMLN 1935
Query: 779 SESNTLPSPREPAF-------VIRRCPSSRASTS 805
E LP P PAF ++ PS R STS
Sbjct: 1936 GEKE-LPKPSLPAFYTGKHDPILLESPSRRCSTS 2034
Score = 224 bits (570), Expect = 6e-59
Identities = 157/516 (30%), Positives = 261/516 (50%), Gaps = 23/516 (4%)
Frame = +1
Query: 16 LCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVW 75
L S+++ IT+ N ++TL+SA G FE GFF S R+Y GI Y ++P+T+VW
Sbjct: 7 LLSSQNYITVTQNQSIQDDETLVSAEGTFEAGFFGLGNSQ--RQYFGIWYKSISPRTIVW 180
Query: 76 VANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVS 135
VANRD P+ +S +T+ GNL +LD + W +N R + M +LLDSGNL+V
Sbjct: 181 VANRDAPVQNSTATIKLTDKGNLLILDGSKGIIWSSNGSRAAEKPYM--QLLDSGNLVVK 354
Query: 136 D--DKVKKILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHDDPAPGNFSFEQD-Q 186
D + K ++W+SF P DT L GMK+ ++ LTSWR+ +DPA G FS+ D +
Sbjct: 355 DGGKRKKNLIWESFDYPGDTLLAGMKIKSNLVKGPTSYLTSWRNTEDPASGEFSYLIDTR 534
Query: 187 GENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYL----LSNFTLRI-SPNNTIPFLTS 241
G Q VI + + Y+++ TG++ S S+L + F+++ S ++ + T+
Sbjct: 535 GFPQLVITRNATAYYRAG-----PWTGKLFSGSSWLRLRKILTFSMQFTSQEISLEYETA 699
Query: 242 SLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCK 300
+ TR V+ G Q L D + W ++ P D+C+ + CG C+ + +C
Sbjct: 700 NRSIITRAVINPSGTTQRLLWSDRSQSWEIISTHPTDQCTYYGLCGANSMCDISNNPICH 879
Query: 301 CLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQFNAKNE-- 358
CL GFRP W++ D+ GGC N+ ++ D FL +K+ + + + KN+
Sbjct: 880 CLEGFRPKFQAKWNSFDWPGGCVPMKNLSCQN--GDGFLKHTGVKLPDTSSSWYGKNKSL 1053
Query: 359 EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNL--EEEYEGGCDLHV 416
+EC CL NC C +Y+Y D+ + C IW D+ +L + G ++++
Sbjct: 1054DECGTLCLQNCSCTSYAYL--------DNDIGGSACLIWFGDILDLSIHPNPDQGQEIYI 1209
Query: 417 RVAFSDIE----GDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIK 472
+V S+++ S+ T++ S I+ ++ L + +STCI ++ +
Sbjct: 1210KVVASELDHRRNKKSFMTKKLAGSLAGIVALVICIIILGLATSTCIQRKKNERGD----- 1374
Query: 473 LYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESIL 508
G + L + RLQ D A + + H +S L
Sbjct: 1375--GDSTRSKLLDWNKRLQIIDGIARGLLYLHQDSRL 1476
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 205 bits (522), Expect = 2e-53
Identities = 133/343 (38%), Positives = 181/343 (51%), Gaps = 11/343 (3%)
Frame = +3
Query: 439 IILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDD--AKA 496
I++ T LI L+ +++ R RQ I G + + I +DD K+
Sbjct: 1938 IVIGAITCGSLLITLAFGVLFVCRYRQKLIPWEGFAGKKYPMETNIIFSLPSKDDFFIKS 2117
Query: 497 IDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFK 556
+ I F LE I AT + +G+GGFG VY+G GQE+AVK SS S QG EF
Sbjct: 2118 VSIQAFTLEYIEVATERYKTL--IGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFD 2291
Query: 557 NEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLD---------AFIFDWDMRFK 607
NE+ L++ +QH NLV LLGYC E D+++LVY +M N SL I DW R
Sbjct: 2292 NELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLS 2471
Query: 608 IILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDR 667
I LG ARGL YLH +IHRD+K+SNILLD K++DFG ++ + +
Sbjct: 2472 IALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLE 2651
Query: 668 VVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLW 727
V GT GY+ PEY S KSDVFSFGVV+LEI+SG+ +P E SL+ +A
Sbjct: 2652 VRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYI 2831
Query: 728 KVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTM 770
+ + + +D + AE + V V L CL+ RP+M
Sbjct: 2832 RGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSM 2960
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 197 bits (502), Expect = 4e-51
Identities = 106/186 (56%), Positives = 126/186 (66%), Gaps = 11/186 (5%)
Frame = +3
Query: 491 EDDAKAIDIPH---FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSC 547
EDD + I F E+++ AT NF NKLG+GGFGPV+KGK G+EIAVK+LS
Sbjct: 165 EDDIQNIAAKEHKTFSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRR 344
Query: 548 SGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF------- 600
S QG +F NE L+ R+QHRN+V L GYC G EK+LVYEY+P SLD +F
Sbjct: 345 SNQGRTQFINEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKEQ 524
Query: 601 -DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGK 659
DW RF II G+ARGLLYLHEDS IIHRD+KA+NILLDE+ PKI+DFGLARIF
Sbjct: 525 LDWKRRFDIISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFPED 704
Query: 660 DTVGNT 665
T NT
Sbjct: 705 QTHVNT 722
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 193 bits (490), Expect = 1e-49
Identities = 100/214 (46%), Positives = 139/214 (64%), Gaps = 9/214 (4%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F L+ + ATNNF NKLG+GGFG VY G+ G +IAVKRL S + EF EV +
Sbjct: 28 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 207
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSL---------DAFIFDWDMRFKIILGI 612
+AR++H+NL+ L GYC EG E+++VY+YMPN SL + DW+ R I +G
Sbjct: 208 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGS 387
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
A G++YLH + IIHRD+KASN+LLD + +++DFG A++ T T RV GT
Sbjct: 388 AEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGAT-HVTTRVKGTL 564
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
GY++PEYA+ G + DVFSFG+++LE+ SGK+
Sbjct: 565 GYLAPEYAMLGKANECCDVFSFGILLLELASGKK 666
>BP048842
Length = 524
Score = 192 bits (489), Expect = 1e-49
Identities = 98/145 (67%), Positives = 112/145 (76%), Gaps = 8/145 (5%)
Frame = -3
Query: 586 VYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNI 637
+YE+M NRSL+ FIFD W+ R +II GIARGLLYLH+DSRLRIIHRDLK SNI
Sbjct: 522 IYEFMHNRSLNYFIFDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNI 343
Query: 638 LLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVV 697
LLD+E NPKISDFGLARIF G T RV+GTYGYM PEYA+ G FS+KSDVFSFGV+
Sbjct: 342 LLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVI 163
Query: 698 VLEIISGKRNTGFYQPEHELSLLGY 722
VLEIISGK+ FY P H L+LL +
Sbjct: 162 VLEIISGKKIGRFYDPHHHLNLLSH 88
>CN825263
Length = 663
Score = 189 bits (480), Expect = 2e-48
Identities = 97/209 (46%), Positives = 138/209 (65%), Gaps = 9/209 (4%)
Frame = +1
Query: 524 GFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEK 583
GFG VYKG G+++AVK L +G EF EV +++RL HRNLV+L+G C+E +
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 584 MLVYEYMPNRSLDAFI---------FDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKA 634
L+YE +PN S+++ + DW+ R KI LG ARGL YLHEDS +IHRD K+
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 635 SNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSF 694
SNILL+ + PK+SDFGLAR + + V+GT+GY++PEYA+ GH VKSDV+S+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 695 GVVVLEIISGKRNTGFYQPEHELSLLGYA 723
GVV+LE+++G + QP + +L+ +A
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLVTWA 627
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 186 bits (471), Expect = 2e-47
Identities = 102/265 (38%), Positives = 154/265 (57%), Gaps = 7/265 (2%)
Frame = +3
Query: 520 LGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVE 579
LG GGFG VY G+ GG ++A+KR + S QG+ EF+ E+ ++++L+HR+LV L+GYC E
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 580 GDEKMLVYEYMPNRSLDAFIFD-------WDMRFKIILGIARGLLYLHEDSRLRIIHRDL 632
E +LVY++M +L ++ W R +I +G ARGL YLH ++ IIHRD+
Sbjct: 195 NTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRDV 374
Query: 633 KASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVF 692
K +NILLDE+ K+SDFGL++ D + V G++GY+ PEY + KSDV+
Sbjct: 375 KTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDVY 554
Query: 693 SFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKC 752
SFGVV+ EI+ + + ++SL +A H + G +D L E K
Sbjct: 555 SFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFKKF 734
Query: 753 VNVGLLCLQEDPIERPTMSNVLFML 777
+ C+ + IERP+M +VL+ L
Sbjct: 735 AETAMKCVSDQGIERPSMGDVLWNL 809
>BP032158
Length = 555
Score = 184 bits (468), Expect = 4e-47
Identities = 98/182 (53%), Positives = 128/182 (69%), Gaps = 9/182 (4%)
Frame = +2
Query: 501 HFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVV 560
+F L I ATNNF ANK+G+GGFGPVYKG G IAVK+LSS S QG EF NE+
Sbjct: 11 YFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIG 190
Query: 561 LIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF---------DWDMRFKIILG 611
+I+ LQH NLV+L G C+EG++ +LVYEYM N SL +F +W R KI +G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 612 IARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGT 671
IA+GL YLHE+SRL+I+HRD+KA+N+LLD++ KISDFGLA++ ++T +T R+ GT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHIST-RIAGT 547
Query: 672 YG 673
G
Sbjct: 548 IG 553
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 180 bits (457), Expect = 7e-46
Identities = 91/214 (42%), Positives = 139/214 (64%), Gaps = 9/214 (4%)
Frame = +2
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F + + AT F+ NKLG+GGFG VY G+ G +IAVK+L + + + EF EV +
Sbjct: 266 FTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEV 445
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI---------FDWDMRFKIILGI 612
+ R++H+NL+ L GYCV D++++VY+YMPN SL + + +W R KI +G
Sbjct: 446 LGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGS 625
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
A G+LYLH + IIHRD+KASN+LL+ + P ++DFG A++ + T RV GT
Sbjct: 626 AEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLI-PEGVSHMTTRVKGTL 802
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
GY++PEYA+ G S DV+SFG+++LE+++G++
Sbjct: 803 GYLAPEYAMWGKVSESCDVYSFGILLLELVTGRK 904
>AI967315
Length = 1308
Score = 178 bits (451), Expect = 4e-45
Identities = 111/284 (39%), Positives = 161/284 (56%), Gaps = 11/284 (3%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLS-SCSGQGLE-EFKNEV 559
F E + ATN F+ N +G+GG+ VYKG+ G EIAVKRL+ +C + E EF E+
Sbjct: 112 FSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEI 291
Query: 560 VLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI-------FDWDMRFKIILGI 612
I + H N++ LLG C++ LV+E S+ + I DW R+KI+LG
Sbjct: 292 GTIGHVCHSNVMPLLGCCIDNG-LYLVFELSTVGSVASLIHDEKMAPLDWKTRYKIVLGT 468
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
ARGL YLH+ + RIIHRD+KASNILL E+ P+ISDFGLA+ + T + + GT+
Sbjct: 469 ARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGTF 648
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNT-GFYQPEHELSL-LGYAWHLWKVG 730
G+++PEY + G K+DVF+FGV +LE+ISG++ G +Q H + + W + K+
Sbjct: 649 GHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVDGSHQSLHTWAKPILSKWEIEKL- 825
Query: 731 RTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVL 774
+D L + + LC++ RPTMS VL
Sbjct: 826 -----VDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVL 942
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 176 bits (446), Expect = 1e-44
Identities = 104/283 (36%), Positives = 159/283 (55%), Gaps = 17/283 (6%)
Frame = +2
Query: 518 NKLGQGGFGPVYKGKFPGGQEIAVKRL-SSCSGQGLEEFKNEVVLIARLQHRNLVRLLGY 576
N +G+GG G VY+G P G ++A+KRL SG+ F+ E+ + +++HRN++RLLGY
Sbjct: 2204 NIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGY 2383
Query: 577 CVEGDEKMLVYEYMPNRSLDAFIFD-------WDMRFKIILGIARGLLYLHEDSRLRIIH 629
D +L+YEYMPN SL ++ W+MR+KI + ARGL Y+H D IIH
Sbjct: 2384 VSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAVEAARGLCYMHHDCSPLIIH 2563
Query: 630 RDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKS 689
RD+K++NILLD + ++DFGLA+ + + G+YGY++PEYA KS
Sbjct: 2564 RDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEKS 2743
Query: 690 DVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWK-------VGRTLDFIDQTLSQ 742
DV+SFGVV+LE+I G++ G + + ++G+ L +D LS
Sbjct: 2744 DVYSFGVVLLELIIGRKPVGEF--GDGVDIVGWVNKTMSELSQPSDTALVLAVVDPRLS- 2914
Query: 743 TCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGS--ESNT 783
+ N+ ++C++E RPTM V+ ML + +SNT
Sbjct: 2915 GYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHMLTNPPQSNT 3043
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 173 bits (438), Expect = 1e-43
Identities = 101/293 (34%), Positives = 158/293 (53%), Gaps = 15/293 (5%)
Frame = +2
Query: 497 IDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLS-SCSGQGLEEF 555
I++P L+ + + T+NF +G+G +G VY G +AVK+L S + EF
Sbjct: 308 IEVPALSLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEF 487
Query: 556 KNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI--------------FD 601
+V +++RL++ N V L GYCVEG+ ++L YE+ SL + D
Sbjct: 488 LTQVSMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLD 667
Query: 602 WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDT 661
W R +I + ARGL YLHE + IIHRD+++SN+L+ E+ KI+DF L+
Sbjct: 668 WIQRVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAA 847
Query: 662 VGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLG 721
++ RV+GT+GY +PEYA+ G + KSDV+SFGVV+LE+++G++ P + SL+
Sbjct: 848 RLHSTRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVT 1027
Query: 722 YAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVL 774
+A + +D L + K V LC+Q + RP MS V+
Sbjct: 1028WATPRLSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVV 1186
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 173 bits (438), Expect = 1e-43
Identities = 108/292 (36%), Positives = 164/292 (55%), Gaps = 16/292 (5%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F + + ATNNF++ NK+GQGGFG VY + G ++ A+K++ Q EF E+ +
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELRG-KKTAIKKMDV---QASTEFLCELKV 1218
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI-------FDWDMRFKIILGIAR 614
+ + H NLVRL+GYCVEG LVYE++ N +L ++ W R +I L AR
Sbjct: 1219 LTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDAAR 1395
Query: 615 GLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNT---DRVVGT 671
GL Y+HE + IHRD+K++NIL+D+ K++DFGL ++ VGN+ R+VGT
Sbjct: 1396 GLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLI----EVGNSTLQTRLVGT 1563
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR---NTGFYQPEHELSLLGYAWHLWK 728
+GYM PEYA G S K DV++FGVV+ E+IS K TG E + + + L K
Sbjct: 1564 FGYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNK 1743
Query: 729 ---VGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFML 777
+D L + + +K +G C +++P+ RP+M +++ L
Sbjct: 1744 SDPCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLVVAL 1899
>BP066041
Length = 507
Score = 159 bits (401), Expect(2) = 4e-43
Identities = 77/82 (93%), Positives = 79/82 (95%)
Frame = -2
Query: 730 GRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPRE 789
GRTLDFID TLS TC+AEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPRE
Sbjct: 506 GRTLDFIDPTLSPTCQAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPRE 327
Query: 790 PAFVIRRCPSSRASTSSKMETF 811
PAFVIRRCPSSRASTSS+ME F
Sbjct: 326 PAFVIRRCPSSRASTSSQMELF 261
Score = 33.9 bits (76), Expect(2) = 4e-43
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = -1
Query: 810 TFSRNDLTVTLENGR 824
TFSRNDLTVTLENGR
Sbjct: 267 TFSRNDLTVTLENGR 223
>AV422094
Length = 462
Score = 166 bits (419), Expect = 2e-41
Identities = 83/147 (56%), Positives = 106/147 (71%), Gaps = 6/147 (4%)
Frame = +2
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F + +ATN+F I NKLG+GGFGPVYKG G IAVK+LS S QG +F E+
Sbjct: 14 FSYSELKNATNDFNIDNKLGEGGFGPVYKGILNDGTVIAVKQLSLGSHQGKSQFIAEIAT 193
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF------DWDMRFKIILGIARG 615
I+ +QHRNLV+L G C+EG +++LVYEY+ N+SLD +F +W R+ I LG+ARG
Sbjct: 194 ISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQALFGKALSLNWSTRYDICLGVARG 373
Query: 616 LLYLHEDSRLRIIHRDLKASNILLDEE 642
L YLHE+SRLRI+HRD+KASNILLD E
Sbjct: 374 LTYLHEESRLRIVHRDVKASNILLDHE 454
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 156 bits (395), Expect = 1e-38
Identities = 85/190 (44%), Positives = 121/190 (62%), Gaps = 13/190 (6%)
Frame = +3
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQ-EIAVKRLSSCSGQGLEEFKNEVV 560
F+ + ATNNF +KLGQGG+G VY+G P + E+AVK S + ++F +E++
Sbjct: 132 FNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELI 311
Query: 561 LIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF-----------DWDMRFKII 609
+I RL+H++LVRL G+C + +LVY+YMPN SLD+ IF W +R+KII
Sbjct: 312 IINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKII 491
Query: 610 LGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLAR-IFGGKDTVGNTDRV 668
G+A L YLH + +++HRDLKASNI+LD E N K+ DFGLAR + K + + V
Sbjct: 492 SGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEGV 671
Query: 669 VGTYGYMSPE 678
GT GY++PE
Sbjct: 672 HGTMGYIAPE 701
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 153 bits (386), Expect = 1e-37
Identities = 84/225 (37%), Positives = 134/225 (59%), Gaps = 9/225 (4%)
Frame = +3
Query: 558 EVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAF-------IFDWDMRFKIIL 610
EV L+A++ HRNLV+LLG+ +G+E++L+ EY+PN +L I D++ R +I +
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 185
Query: 611 GIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARI--FGGKDTVGNTDRV 668
+A GL YLH + +IIHRD+K+SNILL E K++DFG AR+ G T +T +V
Sbjct: 186 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIST-KV 362
Query: 669 VGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWK 728
GT GY+ PEY + KSDV+SFG+++LEI++G+R + E L +A+ +
Sbjct: 363 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYN 542
Query: 729 VGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNV 773
G ++ +D + + A+ MK +++ C +RP M +V
Sbjct: 543 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 677
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,631,878
Number of Sequences: 28460
Number of extensions: 240869
Number of successful extensions: 1650
Number of sequences better than 10.0: 328
Number of HSP's better than 10.0 without gapping: 1432
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1447
length of query: 824
length of database: 4,897,600
effective HSP length: 98
effective length of query: 726
effective length of database: 2,108,520
effective search space: 1530785520
effective search space used: 1530785520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0203.7


