

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.5
(852 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
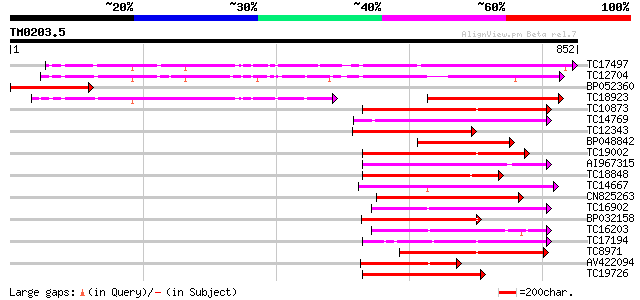
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 458 e-129
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 419 e-117
BP052360 264 4e-71
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 244 3e-65
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 227 7e-60
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 214 6e-56
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 206 1e-53
BP048842 202 2e-52
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 196 1e-50
AI967315 192 2e-49
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 190 7e-49
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 189 2e-48
CN825263 188 3e-48
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 186 2e-47
BP032158 186 2e-47
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 185 3e-47
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 178 4e-45
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 170 8e-43
AV422094 166 1e-41
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 152 2e-37
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 458 bits (1179), Expect = e-129
Identities = 308/826 (37%), Positives = 444/826 (53%), Gaps = 27/826 (3%)
Frame = +1
Query: 54 FELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADD 111
FE GFF + + Y G+WY +SP +VWVANRD P+ + + ++
Sbjct: 280 FEAGFF----HFENPQHHYFGVWY-----KSISPRTIVWVANRDAPLRNSTAPTLKVTHK 432
Query: 112 GNLVVLDTSGIRYWSASNLTNSSSATNRS-VKLMDSGNLVLLDEHVGMK-LWESFEHPTD 169
G++++ D + WS TN+S A + ++L+DSGNLV D G +WESF +P D
Sbjct: 433 GSILIRDGAKGVIWS----TNTSRAKEQPFMQLLDSGNLVAKDGDKGENVIWESFNYPGD 600
Query: 170 TFLPGMKMDKTLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEE 223
TFL GMK+ L LT W++ DP G F++ +D + + + + ++
Sbjct: 601 TFLAGMKIKSNLAIGPTSYLTSWRNSEDPASGEFSYHIDIRGFPQLVVTKGAAITLRA-- 774
Query: 224 QGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVS-SYDN------TRLLLNSTGVI-KV 275
G + +LT F + ++ +S Y+ TR ++ G I ++
Sbjct: 775 ---GPWTGNKFSGAFGQVLQKILTFFMQFTDQEISLEYETVNRSIITREVITPLGTIQRL 945
Query: 276 LYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRR--SPLNDYTVG 333
L+ V QS + +P C Y CG S C+ + +C CL GF + + N
Sbjct: 946 LWSVRNQSWEIIATRPVDQCADYVFCGANSLCDTSKNPICDCLEGFMPQFQAKWNSLDWA 1125
Query: 334 GDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQA 393
G S+ + SC N + F+ T +K+ PD S +N C ++C Q C
Sbjct: 1126 GGCVSM----EKLSC-QNGDGFMKHTGVKL--PDTSSSWFGKNMSLDECRTLCLQN-CSC 1281
Query: 394 CSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRK---GN 450
+Y + ++ S C IW ++ + + +++++RV S ++ K
Sbjct: 1282 TAYAGL---DNDVDRSVCLIWFGDILDMSKHPDPDQGQEIYIRVVASKLDRTRNKKSINT 1452
Query: 451 PKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKD 510
K SL++ IA V+ + + + C ++ K E E + +
Sbjct: 1453 KKLAGSLVVIIAF--VIFITILGLAISTCIQRKKNKRGDEGEIGIINHWK---------- 1596
Query: 511 LIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVK 570
DK G E+ D I FDF +I AT++FS +NKLG GG+GPVYKG L G+EIAVK
Sbjct: 1597 --DKRGDEDIDLATI----FDFSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVK 1758
Query: 571 RLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDP 630
RLS+ S QG++EFKNE+ LIA+LQHRNLV+L+G + DE NK + + D
Sbjct: 1759 RLSNTSGQGMEEFKNEIKLIARLQHRNLVKLFGCSVHQDENS-----HANKKMK-ILLDS 1920
Query: 631 TKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLAR 690
T+S L+DW R I+ GIARGLLYLHQDSRLR+IHRDLKTSNILLD EM PKISDFGLAR
Sbjct: 1921 TRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLAR 2100
Query: 691 IFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYK 750
IF G + EA T+RV+GTYGYM PEYA+ G FS KSD+FSFGV++LEIISGKK FY
Sbjct: 2101 IFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPH 2280
Query: 751 GTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVV 810
L+LL +AW+LW E + L+L+D L + + +R HV LLCVQ P++RP+M ++V
Sbjct: 2281 HHLNLLSHAWRLWIEERPLELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIV 2460
Query: 811 IMLDSETATLPTPKQPTFFTRKD----LSSTASSSLQFDSSIVEGR 852
+ML+ E LP P+ P F+T K L S + S S++E R
Sbjct: 2461 LMLNGE-KELPKPRLPAFYTGKHDPIWLGSPSRCSTSITISLLEAR 2595
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 419 bits (1076), Expect = e-117
Identities = 306/837 (36%), Positives = 425/837 (50%), Gaps = 50/837 (5%)
Frame = +1
Query: 47 LVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIG 104
LVS FE GFF RY GIWY +SP +VWVANRD PV +S
Sbjct: 37 LVSPEGTFEAGFF----RFGNSLRRYFGIWY-----KSISPRTIVWVANRDAPV-QNSTA 186
Query: 105 VFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSV-KLMDSGNLVLLDEHVGMKL-WE 162
++ D GNL++LD WS+ N+S ++ + +L+DSGN V+ D L WE
Sbjct: 187 TLKLTDQGNLLILDGLKGIVWSS----NASRTKDKPLMQLLDSGNFVVKDGDKEENLIWE 354
Query: 163 SFEHPTDTFLPGMKMDKTLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQ 216
SF++P DTFL GMK+ L LT W++ DP G F++ +D + +
Sbjct: 355 SFDYPGDTFLAGMKIKSNLATGPTSYLTSWRNAEDPASGEFSYHIDTHGYPQLVVTKGAT 534
Query: 217 LYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVS-SYDN------TRLLLNS 269
+ ++ G + N S + + +LT + +K VS Y+ TR ++
Sbjct: 535 VTLRA---GPWIGNKFSGASGLR--LQKILTFSMQFTDKEVSLEYETANRSIITRTVITP 699
Query: 270 TGVI-KVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLN 328
+G ++L+ QS + P C Y CG S C+ N+ +C CL GF +
Sbjct: 700 SGTTQRLLWSDRSQSWEIISTHPMDQCAYYAFCGANSMCDTSNNPICDCLEGFTPKFQAQ 879
Query: 329 DYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVS----AQDENECKFRCIS 384
++ D + K+ SC N + F T ++ PD S ++ +EC C+
Sbjct: 880 WNSL--DWTGGCVPIKNLSC-QNGDGFPKHTGVQF--PDTSSSWYGNSKSLDECGTICLQ 1044
Query: 385 MCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEE 444
CS C A +Y+ V R S C W ++ + E +++++RV S+++
Sbjct: 1045NCS---CTAYAYLD-NVGGR----SVCLNWFGDILDMSEHPDPDQGQEIYLRVVASELDH 1200
Query: 445 PTRKGNPKSTLSLILGIALPGVVILACICILAYVC----RRKIALKLKQESESILRQRGR 500
R+ + + G + + CI IL RRK K ++E E + R
Sbjct: 1201--RRNKKSINIKKLAGSLAGSIAFIICITILGLATVTCIRRK---KNEREDEGGIETR-- 1359
Query: 501 FYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGK 560
+ DK G E+ D I FDF +I T++FS++NKLG GG+GPVYKG
Sbjct: 1360------IINHWKDKRGDEDIDLATI----FDFSTISSTTNHFSESNKLGEGGFGPVYKGV 1509
Query: 561 LQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPN 620
L G+EIAVKRLS+ S QG++EFKNEV LIA+LQHRNLV+L G I DE +LIYE+M N
Sbjct: 1510LANGQEIAVKRLSNTSGQGMEEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLLIYEFMHN 1689
Query: 621 KSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQ 680
+SLD F+F DSRLR+IHRDLKTSNILLD EM
Sbjct: 1690RSLDYFIF-----------------------------DSRLRIIHRDLKTSNILLDSEMN 1782
Query: 681 PKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISG 740
PKISDFGLARIF G + EA T+RV+GTYGYMSPEYA+ G FS KSD+FSFGV++LEIISG
Sbjct: 1783PKISDFGLARIFTGDQVEAKTKRVMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLEIISG 1962
Query: 741 KKNTGFYQYKGTLSLLGY------------------------AWKLWTENKLLDLMDLSL 776
KK F +LL + AW+LW E + L+L+D L
Sbjct: 1963KKIGRFCDPHHHRNLLSHSSNFAVFLIKALRICMFENVKNRKAWRLWIEERPLELVDELL 2142
Query: 777 GEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKD 833
+ +R H+ LLCVQ P+ RP+M +VV+ML+ E LP P P F+T D
Sbjct: 2143DGLAIPTEILRYIHIALLCVQQRPEYRPDMLSVVLMLNGE-KELPKPSLPAFYTGND 2310
>BP052360
Length = 519
Score = 264 bits (675), Expect = 4e-71
Identities = 124/125 (99%), Positives = 124/125 (99%)
Frame = +3
Query: 1 MFFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFS 60
MFFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFS
Sbjct: 144 MFFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFS 323
Query: 61 PDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTS 120
PDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDS GVFRIADDGNLVVLDTS
Sbjct: 324 PDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSXGVFRIADDGNLVVLDTS 503
Query: 121 GIRYW 125
GIRYW
Sbjct: 504 GIRYW 518
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 244 bits (624), Expect = 3e-65
Identities = 123/204 (60%), Positives = 149/204 (72%)
Frame = +1
Query: 629 DPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGL 688
D T+S LLDW R I+ GIARGLLYLHQDSRLR+IHRDLKTSNILLD EM PKISDFGL
Sbjct: 1378 DSTRSKLLDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFGL 1557
Query: 689 ARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQ 748
ARIF G + EA T+RV+GTYGYM PEYA+ G FS KSD+FSFGV++LEIISGKK FY
Sbjct: 1558 ARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFYD 1737
Query: 749 YKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSN 808
L+LL +AW+LW E L+L+D ++ + +R HV LLCVQ P+ RP+M +
Sbjct: 1738 PHHHLNLLSHAWRLWIEGSPLELVDKLFEDSIIPTEILRYIHVALLCVQRRPETRPDMLS 1917
Query: 809 VVIMLDSETATLPTPKQPTFFTRK 832
+V+ML+ E LP P P F+T K
Sbjct: 1918 IVLMLNGE-KELPKPSLPAFYTGK 1986
Score = 154 bits (389), Expect = 6e-38
Identities = 137/476 (28%), Positives = 219/476 (45%), Gaps = 16/476 (3%)
Frame = +1
Query: 33 TLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSP--VVW 90
T+ Q I + T LVSA FE GFF + + +Y GIWY +SP +VW
Sbjct: 31 TVTQNQSIQDDET-LVSAEGTFEAGFFG----LGNSQRQYFGIWY-----KSISPRTIVW 180
Query: 91 VANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLV 150
VANRD PV +S ++ D GNL++LD S WS++ S +A ++L+DSGNLV
Sbjct: 181 VANRDAPV-QNSTATIKLTDKGNLLILDGSKGIIWSSNG---SRAAEKPYMQLLDSGNLV 348
Query: 151 LLDEHVGMK--LWESFEHPTDTFLPGMKMDKTLE------LTCWKSLSDPGRGNFTFKMD 202
+ D K +WESF++P DT L GMK+ L LT W++ DP G F++ +D
Sbjct: 349 VKDGGKRKKNLIWESFDYPGDTLLAGMKIKSNLVKGPTSYLTSWRNTEDPASGEFSYLID 528
Query: 203 KKWENRFAILNQGQLYWQSEEQGDGVMNPES--NPDDISNDVYNLLTNFKELKNKTVSSY 260
+ + I Y+++ + + S I + L+ +T +
Sbjct: 529 TRGFPQLVITRNATAYYRAGPWTGKLFSGSSWLRLRKILTFSMQFTSQEISLEYETANRS 708
Query: 261 DNTRLLLNSTGVI-KVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLP 319
TR ++N +G ++L+ QS + P C Y +CG S C+ N+ +C CL
Sbjct: 709 IITRAVINPSGTTQRLLWSDRSQSWEIISTHPTDQCTYYGLCGANSMCDISNNPICHCLE 888
Query: 320 GFGRR--SPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDENE 377
GF + + N + G + K+ SC N + FL T +K+ PD S +N+
Sbjct: 889 GFRPKFQAKWNSFDWPGGCVPM----KNLSC-QNGDGFLKHTGVKL--PDTSSSWYGKNK 1047
Query: 378 CKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRV 437
C ++C Q C SY + + S C IW ++ L ++++++V
Sbjct: 1048SLDECGTLCLQ-NCSCTSYAYL---DNDIGGSACLIWFGDILDLSIHPNPDQGQEIYIKV 1215
Query: 438 AKSDIEEPTRKGNPKSTLSLILGIALPGVVILA-CICILAYVCRRKIALKLKQESE 492
S+++ + N KS ++ L +L G+V L CI IL I K + +
Sbjct: 1216VASELD---HRRNKKSFMTKKLAGSLAGIVALVICIIILGLATSTCIQRKKNERGD 1374
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 227 bits (578), Expect = 7e-60
Identities = 121/288 (42%), Positives = 186/288 (64%), Gaps = 3/288 (1%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F F + AT F +AN +G GG+G VYKG+L G +AVK+LS QG QEF EV++
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRFDILLGI 648
++ L H NLVRL GYC GD+++L+YEYMP SL+ +F+ + L+W R + +G
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG-GKETEANTQRVVGT 707
ARGL YLH + VI+RDLK++NILLD E PK+SDFGLA++ G T +T RV+GT
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVST-RVMGT 951
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK 767
YGY +PEYA+ G+ + KSDI+SFGVVLLE+++G++ + G +L+ +A +++ +
Sbjct: 952 YGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRR 1131
Query: 768 LL-DLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
++D L + + + + +C+Q++P RP ++++V+ L+
Sbjct: 1132RFGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIVVALE 1275
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 214 bits (544), Expect = 6e-56
Identities = 124/302 (41%), Positives = 177/302 (58%), Gaps = 4/302 (1%)
Frame = +3
Query: 517 LEEKDN---EGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLS 573
L KD+ + + + F E I VAT+ + +G GG+G VY+G L G+E+AVK S
Sbjct: 2085 LPSKDDFFIKSVSIQAFTLEYIEVATERYKTL--IGEGGFGSVYRGTLNDGQEVAVKVRS 2258
Query: 574 SVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSL-DAFVFDPTK 632
S S+QG +EF NE+ L++ +QH NLV L GYC + D++IL+Y +M N SL D +P K
Sbjct: 2259 STSTQGTREFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAK 2438
Query: 633 SALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIF 692
+LDW R I LG ARGL YLH VIHRD+K+SNILLD M K++DFG ++
Sbjct: 2439 RKILDWPTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYA 2618
Query: 693 GGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGT 752
+ + V GT GY+ PEY Q S KSD+FSFGVVLLEI+SG++ + +
Sbjct: 2619 PQEGDSYVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTE 2798
Query: 753 LSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIM 812
SL+ +A +K+ +++D + Y+A R V L C++ RP+M +V
Sbjct: 2799 WSLVEWATPYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRE 2978
Query: 813 LD 814
L+
Sbjct: 2979 LE 2984
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 206 bits (524), Expect = 1e-53
Identities = 103/190 (54%), Positives = 133/190 (69%), Gaps = 3/190 (1%)
Frame = +3
Query: 515 EGLEEKDNEGI---EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKR 571
E E D + I E F +E+++ AT F NKLG GG+GPV+KGKL GREIAVK+
Sbjct: 153 EAQNEDDIQNIAAKEHKTFSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKK 332
Query: 572 LSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPT 631
LS S+QG +F NE L+ ++QHRN+V L+GYC G EK+L+YEY+P +SLD +F
Sbjct: 333 LSRRSNQGRTQFINEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQ 512
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
K LDW+ RFDI+ G+ARGLLYLH+DS +IHRD+K +NILLD + PKI+DFGLARI
Sbjct: 513 KKEQLDWKRRFDIISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARI 692
Query: 692 FGGKETEANT 701
F +T NT
Sbjct: 693 FPEDQTHVNT 722
>BP048842
Length = 524
Score = 202 bits (513), Expect = 2e-52
Identities = 100/145 (68%), Positives = 117/145 (79%)
Frame = -3
Query: 614 IYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNI 673
IYE+M N+SL+ F+FD T+S L+DW R I+ GIARGLLYLHQDSRLR+IHRDLKTSNI
Sbjct: 522 IYEFMHNRSLNYFIFDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNI 343
Query: 674 LLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVV 733
LLD EM PKISDFGLARIF G + EA T+RV+GTYGYM PEYA+ G FS KSD+FSFGV+
Sbjct: 342 LLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVI 163
Query: 734 LLEIISGKKNTGFYQYKGTLSLLGY 758
+LEIISGKK FY L+LL +
Sbjct: 162 VLEIISGKKIGRFYDPHHHLNLLSH 88
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 196 bits (498), Expect = 1e-50
Identities = 104/252 (41%), Positives = 154/252 (60%), Gaps = 1/252 (0%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F + + AT+ F+ NKLG GG+G VY G+L G +IAVKRL S++ EF EV +
Sbjct: 28 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 207
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS-ALLDWQMRFDILLGI 648
+A+++H+NL+ L GYC +G E++++Y+YMPN SL + + S LLDW R +I +G
Sbjct: 208 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGS 387
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
A G++YLH + +IHRD+K SN+LLD + Q +++DFG A++ T T RV GT
Sbjct: 388 AEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTT-RVKGTL 564
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GY++PEYA+ G+ + D+FSFG++LLE+ SGKK S+ +A L K
Sbjct: 565 GYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAKKF 744
Query: 769 LDLMDLSLGEAY 780
+ D L Y
Sbjct: 745 TEFADPRLNGEY 780
>AI967315
Length = 1308
Score = 192 bits (487), Expect = 2e-49
Identities = 113/288 (39%), Positives = 171/288 (59%), Gaps = 3/288 (1%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSV--SSQGIQEFKNEV 587
F +E + AT+ FS N +G+GGY VYKG+L+ G EIAVKRL+ + +EF E+
Sbjct: 112 FSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEI 291
Query: 588 VLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLG 647
I + H N++ L G CI L++E S+ + + D K A LDW+ R+ I+LG
Sbjct: 292 GTIGHVCHSNVMPLLGCCIDNG-LYLVFELSTVGSVASLIHDE-KMAPLDWKTRYKIVLG 465
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
ARGL YLH+ + R+IHRD+K SNILL + +P+ISDFGLA+ + T + + GT
Sbjct: 466 TARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGT 645
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNT-GFYQYKGTLSLLGYAWKLWTEN 766
+G+++PEY + G K+D+F+FGV LLE+ISG+K G +Q SL +A + ++
Sbjct: 646 FGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVDGSHQ-----SLHTWAKPILSKW 810
Query: 767 KLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
++ L+D L Y+ QF R LC++ RP MS V+ +++
Sbjct: 811 EIEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVME 954
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 190 bits (483), Expect = 7e-49
Identities = 90/214 (42%), Positives = 144/214 (67%), Gaps = 1/214 (0%)
Frame = +2
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F ++ + AT FSD NKLG GG+G VY G+ G +IAVK+L +++S+ EF EV +
Sbjct: 266 FTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEV 445
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDP-TKSALLDWQMRFDILLGI 648
+ +++H+NL+ L GYC+ D+++++Y+YMPN SL + + L+WQ R I +G
Sbjct: 446 LGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGS 625
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
A G+LYLH + +IHRD+K SN+LL+ + +P ++DFG A++ + T RV GT
Sbjct: 626 AEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLI-PEGVSHMTTRVKGTL 802
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKK 742
GY++PEYA+ G+ S D++SFG++LLE+++G+K
Sbjct: 803 GYLAPEYAMWGKVSESCDVYSFGILLLELVTGRK 904
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 189 bits (479), Expect = 2e-48
Identities = 112/310 (36%), Positives = 167/310 (53%), Gaps = 10/310 (3%)
Frame = +2
Query: 525 IEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLS-SVSSQGIQEF 583
IEVP + + TD F +G G YG VY L G +AVK+L S + EF
Sbjct: 308 IEVPALSLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEF 487
Query: 584 KNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAF------VFDPTKSALLD 637
+V ++++L++ N V L GYC++G+ ++L YE+ SL V LD
Sbjct: 488 LTQVSMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLD 667
Query: 638 WQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKET 697
W R I + ARGL YLH+ + +IHRD+++SN+L+ + + KI+DF L+
Sbjct: 668 WIQRVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAA 847
Query: 698 EANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLG 757
++ RV+GT+GY +PEYA+ GQ + KSD++SFGVVLLE+++G+K +G SL+
Sbjct: 848 RLHSTRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVT 1027
Query: 758 YAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD--- 814
+A +E+K+ +D L Y + V LCVQ E + RPNMS VV L
Sbjct: 1028WATPRLSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLL 1207
Query: 815 SETATLPTPK 824
A P P+
Sbjct: 1208KTPAAAPAPE 1237
>CN825263
Length = 663
Score = 188 bits (478), Expect = 3e-48
Identities = 100/221 (45%), Positives = 143/221 (64%), Gaps = 1/221 (0%)
Frame = +1
Query: 552 GYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEK 611
G+G VYKG L GR++AVK L +G +EF EV ++++L HRNLV+L G CI+ +
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 612 ILIYEYMPNKSLDAFVFDPTK-SALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKT 670
LIYE +PN S+++ + K + LDW R I LG ARGL YLH+DS VIHRD K+
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 671 SNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSF 730
SNILL+ + PK+SDFGLAR + + + V+GT+GY++PEYA+ G KSD++S+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 731 GVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
GVVLLE+++G K Q G +L+ +A + T + L +
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLVTWARPILTSKEGLQM 663
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 186 bits (471), Expect = 2e-47
Identities = 101/271 (37%), Positives = 162/271 (59%)
Frame = +3
Query: 544 DANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWG 603
+A LG GG+G VY G++ GG ++A+KR + +S QG+ EF+ E+ +++KL+HR+LV L G
Sbjct: 3 EALLLGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIG 182
Query: 604 YCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRV 663
YC + E IL+Y++M +L ++ T+ L W+ R +I +G ARGL YLH ++ +
Sbjct: 183 YCEENTEMILVYDHMAYGTLREHLY-KTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTI 359
Query: 664 IHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFST 723
IHRD+KT+NILLD + K+SDFGL++ + + V G++GY+ PEY Q +
Sbjct: 360 IHRDVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTD 539
Query: 724 KSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNAN 783
KSD++SFGVVL EI+ + K +SL +A + + L ++D L
Sbjct: 540 KSDVYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPE 719
Query: 784 QFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
F + + CV D+ +RP+M +V+ L+
Sbjct: 720 CFKKFAETAMKCVSDQGIERPSMGDVLWNLE 812
>BP032158
Length = 555
Score = 186 bits (471), Expect = 2e-47
Identities = 93/182 (51%), Positives = 130/182 (71%), Gaps = 1/182 (0%)
Frame = +2
Query: 529 YFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVV 588
YF I AT+ F ANK+G GG+GPVYKG L G IAVK+LSS S QG +EF NE+
Sbjct: 11 YFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIG 190
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRFDILLG 647
+I+ LQH NLV+L+G CI+G++ +L+YEYM N SL +F + L L+W+ R I +G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
IA+GL YLH++SRL+++HRD+K +N+LLD +++ KISDFGLA++ + T +T R+ GT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHIST-RIAGT 547
Query: 708 YG 709
G
Sbjct: 548 IG 553
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 185 bits (469), Expect = 3e-47
Identities = 102/278 (36%), Positives = 161/278 (57%), Gaps = 8/278 (2%)
Frame = +2
Query: 544 DANKLGRGGYGPVYKGKLQGGREIAVKRL-SSVSSQGIQEFKNEVVLIAKLQHRNLVRLW 602
+ N +G+GG G VY+G + G ++A+KRL S + F+ E+ + K++HRN++RL
Sbjct: 2198 EENIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLL 2377
Query: 603 GYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLR 662
GY D +L+YEYMPN SL ++ K L W+MR+ I + ARGL Y+H D
Sbjct: 2378 GYVSNKDTNLLLYEYMPNGSLGEWLHG-AKGGHLRWEMRYKIAVEAARGLCYMHHDCSPL 2554
Query: 663 VIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFS 722
+IHRD+K++NILLD + + ++DFGLA+ + + G+YGY++PEYA +
Sbjct: 2555 IIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVD 2734
Query: 723 TKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK-------LLDLMDLS 775
KSD++SFGVVLLE+I G+K G ++ + ++G+ K +E +L ++D
Sbjct: 2735 EKSDVYSFGVVLLELIIGRKPVG--EFGDGVDIVGWVNKTMSELSQPSDTALVLAVVDPR 2908
Query: 776 LGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
L Y I ++ ++CV++ RP M VV ML
Sbjct: 2909 L-SGYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHML 3019
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 178 bits (451), Expect = 4e-45
Identities = 110/290 (37%), Positives = 171/290 (58%), Gaps = 6/290 (2%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F ++ + AT+ FS NK+G+GG+G VY +L+G ++ A+K++ +S EF E+ +
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELRG-KKTAIKKMDVQAST---EFLCELKV 1218
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
+ + H NLVRL GYC++G L+YE++ N +L ++ K L W R I L A
Sbjct: 1219 LTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGNLGQYLHGSGKEPL-PWSSRVQIALDAA 1392
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
RGL Y+H+ + IHRD+K++NIL+D ++ K++DFGL ++ + T R+VGT+G
Sbjct: 1393 RGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQT-RLVGTFG 1569
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKK---NTG--FYQYKGTLSLLGYAW-KLW 763
YM PEYA G S K D+++FGVVL E+IS K TG + KG ++L A K
Sbjct: 1570 YMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKSD 1749
Query: 764 TENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
+ L L+D LGE Y + ++ +G C +D P RP+M ++V+ L
Sbjct: 1750 PCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLVVAL 1899
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 170 bits (431), Expect = 8e-43
Identities = 91/226 (40%), Positives = 142/226 (62%), Gaps = 2/226 (0%)
Frame = +3
Query: 586 EVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDIL 645
EV L+AK+ HRNLV+L G+ KG+E+ILI EY+PN +L + D + +LD+ R +I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL-DGLRGKILDFNQRLEIA 182
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI--FGGKETEANTQR 703
+ +A GL YLH + ++IHRD+K+SNILL M+ K++DFG AR+ G +T +T +
Sbjct: 183 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIST-K 359
Query: 704 VVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLW 763
V GT GY+ PEY Q + KSD++SFG++LLEI++G++ + L +A++ +
Sbjct: 360 VKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY 539
Query: 764 TENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNV 809
E +++LMD + EA NA+ ++ + C DRPNM +V
Sbjct: 540 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 677
>AV422094
Length = 462
Score = 166 bits (420), Expect = 1e-41
Identities = 83/153 (54%), Positives = 112/153 (72%), Gaps = 1/153 (0%)
Frame = +2
Query: 528 PY-FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNE 586
PY F + + AT+ F+ NKLG GG+GPVYKG L G IAVK+LS S QG +F E
Sbjct: 5 PYTFSYSELKNATNDFNIDNKLGEGGFGPVYKGILNDGTVIAVKQLSLGSHQGKSQFIAE 184
Query: 587 VVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILL 646
+ I+ +QHRNLV+L+G CI+G +++L+YEY+ NKSLD +F K+ L+W R+DI L
Sbjct: 185 IATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQALFG--KALSLNWSTRYDICL 358
Query: 647 GIARGLLYLHQDSRLRVIHRDLKTSNILLDGEM 679
G+ARGL YLH++SRLR++HRD+K SNILLD E+
Sbjct: 359 GVARGLTYLHEESRLRIVHRDVKASNILLDHEL 457
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 152 bits (384), Expect = 2e-37
Identities = 80/190 (42%), Positives = 121/190 (63%), Gaps = 5/190 (2%)
Frame = +3
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGR-EIAVKRLSSVSSQGIQEFKNEVV 588
F++ + AT+ F + +KLG+GGYG VY+G L + E+AVK S + +F +E++
Sbjct: 132 FNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELI 311
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF---DPTKSALLDWQMRFDIL 645
+I +L+H++LVRL G+C K +L+Y+YMPN SLD+ +F T + L W +R+ I+
Sbjct: 312 IINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKII 491
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLAR-IFGGKETEANTQRV 704
G+A L YLH + +V+HRDLK SNI+LD E K+ DFGLAR + K + + V
Sbjct: 492 SGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEGV 671
Query: 705 VGTYGYMSPE 714
GT GY++PE
Sbjct: 672 HGTMGYIAPE 701
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,597,684
Number of Sequences: 28460
Number of extensions: 234400
Number of successful extensions: 1745
Number of sequences better than 10.0: 316
Number of HSP's better than 10.0 without gapping: 1531
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1545
length of query: 852
length of database: 4,897,600
effective HSP length: 98
effective length of query: 754
effective length of database: 2,108,520
effective search space: 1589824080
effective search space used: 1589824080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0203.5


