

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.3
(475 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
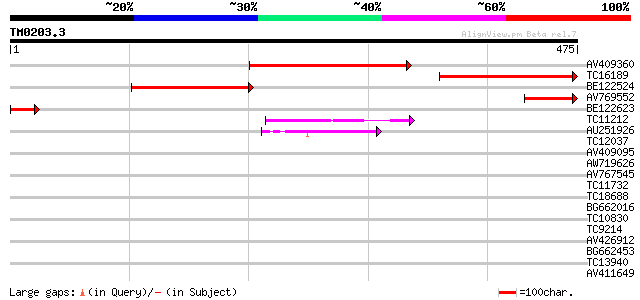
Score E
Sequences producing significant alignments: (bits) Value
AV409360 260 4e-70
TC16189 227 4e-60
BE122524 209 1e-54
AV769552 80 6e-16
BE122623 55 3e-08
TC11212 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (38%) 50 1e-06
AU251926 42 2e-04
TC12037 weakly similar to PIR|T00914|T00914 leucine-rich repeat ... 37 0.007
AV409095 37 0.007
AW719626 35 0.035
AV767545 34 0.059
TC11732 34 0.059
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 32 0.17
BG662016 31 0.50
TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, ... 31 0.50
TC9214 30 1.1
AV426912 28 2.5
BG662453 28 3.3
TC13940 28 3.3
AV411649 28 4.3
>AV409360
Length = 408
Score = 260 bits (664), Expect = 4e-70
Identities = 135/135 (100%), Positives = 135/135 (100%)
Frame = +1
Query: 202 VVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDL 261
VVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDL
Sbjct: 4 VVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDL 183
Query: 262 SYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLA 321
SYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLA
Sbjct: 184 SYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLA 363
Query: 322 ANYNSLQAINLDGNP 336
ANYNSLQAINLDGNP
Sbjct: 364 ANYNSLQAINLDGNP 408
>TC16189
Length = 622
Score = 227 bits (578), Expect = 4e-60
Identities = 115/115 (100%), Positives = 115/115 (100%)
Frame = +1
Query: 361 NRQAMKVSTLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTV 420
NRQAMKVSTLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTV
Sbjct: 1 NRQAMKVSTLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTV 180
Query: 421 ESPKLSKGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL 475
ESPKLSKGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL
Sbjct: 181 ESPKLSKGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL 345
>BE122524
Length = 334
Score = 209 bits (531), Expect = 1e-54
Identities = 101/102 (99%), Positives = 102/102 (99%)
Frame = +2
Query: 103 KTLQGDNSVDCFGEFPNKDFKVKRIEDWVVDLQHCGPPVEEINELPESVDPVVDINTING 162
+TLQGDNSVDCFGEFPNKDFKVKRIEDWVVDLQHCGPPVEEINELPESVDPVVDINTING
Sbjct: 14 ETLQGDNSVDCFGEFPNKDFKVKRIEDWVVDLQHCGPPVEEINELPESVDPVVDINTING 193
Query: 163 VTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVP 204
VTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVP
Sbjct: 194 VTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVP 319
>AV769552
Length = 358
Score = 80.5 bits (197), Expect = 6e-16
Identities = 39/44 (88%), Positives = 42/44 (94%)
Frame = -1
Query: 432 HLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL 475
HLPPI+TKVSTQSRHYL+AQ+KVLNLTSGHSM KS SEGTLGAL
Sbjct: 358 HLPPIKTKVSTQSRHYLNAQNKVLNLTSGHSMPKSPSEGTLGAL 227
>BE122623
Length = 370
Score = 54.7 bits (130), Expect = 3e-08
Identities = 25/25 (100%), Positives = 25/25 (100%)
Frame = +3
Query: 1 MKSRSLPNIKASTLSPENHAFKHPS 25
MKSRSLPNIKASTLSPENHAFKHPS
Sbjct: 294 MKSRSLPNIKASTLSPENHAFKHPS 368
>TC11212 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (38%)
Length = 518
Score = 49.7 bits (117), Expect = 1e-06
Identities = 36/125 (28%), Positives = 56/125 (44%)
Frame = +1
Query: 215 LNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLA 274
L+L N + +T L L LS N I+ +EGL L LRVLD+S N++ + +
Sbjct: 1 LSLQSNRLTSMTGLEGCIALEELYLSHNGITKMEGLSSLVNLRVLDVSSNKLTSV-DDIQ 177
Query: 275 SCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDG 334
+ + L++L+L N+I +EG +A + L I L+
Sbjct: 178 NLTQLEDLWLNDNQIDSLEGFDE---------------------AVAGSTEKLTTIYLEN 294
Query: 335 NPCQK 339
NPC K
Sbjct: 295 NPCAK 309
>AU251926
Length = 335
Score = 42.0 bits (97), Expect = 2e-04
Identities = 34/105 (32%), Positives = 55/105 (52%), Gaps = 5/105 (4%)
Frame = +2
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIE----GLRELTRLRVLDLSYNRIL 267
L ++ LA S++ ++A +G L L + ++ L +L+ L LDLS NRI+
Sbjct: 8 LSLIKLA--SLIEVSA---KKGTRDLKLQNKLLDQVDWLPDSLGKLSSLVTLDLSENRIV 172
Query: 268 RIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKI 311
+ + SSL L L N+I E+ + + LL L LDLR N++
Sbjct: 173 ALPSTIGGLSSLTRLDLHTNRIQELPDSIGNLLNLVYLDLRGNQL 307
>TC12037 weakly similar to PIR|T00914|T00914 leucine-rich repeat protein
F21B7.28 - Arabidopsis thaliana {Arabidopsis thaliana;}
, partial (37%)
Length = 573
Score = 37.0 bits (84), Expect = 0.007
Identities = 30/74 (40%), Positives = 45/74 (60%), Gaps = 4/74 (5%)
Frame = +3
Query: 234 LHSLNLSRNNIS-SIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNK-IGE 291
L SL+L RN + S++ L E+T ++ +DLSYN+ G +S++ LYL NK IG+
Sbjct: 54 LTSLHLERNQFTGSVQPLNEVT-IQTVDLSYNQF--SGEISPMLASVQHLYLNNNKFIGQ 224
Query: 292 VEG--LHRLLKLSI 303
V G + RL+ SI
Sbjct: 225 VPGSFVERLMTASI 266
>AV409095
Length = 414
Score = 37.0 bits (84), Expect = 0.007
Identities = 30/92 (32%), Positives = 47/92 (50%), Gaps = 3/92 (3%)
Frame = +2
Query: 204 PFLSAFVSLKVLNLAGNSIVRI-TAGALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDL 261
P + SL+ L+L+ N I + T GL S+ ++ N + + + L+RL LDL
Sbjct: 17 PEIGCLKSLEYLDLSFNKIKTLPTEITYLIGLISMKVANNKLVELPSAMTSLSRLECLDL 196
Query: 262 SYNRILRIGH-GLASCSSLKELYLAGNKIGEV 292
S NR+ +G LAS L+ L L NK+ +
Sbjct: 197 SNNRLTSLGSLELASMHRLQNLNLQYNKLPSI 292
Score = 35.0 bits (79), Expect = 0.027
Identities = 28/85 (32%), Positives = 45/85 (52%), Gaps = 2/85 (2%)
Frame = +2
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKI 311
L L LDLS+N+I + + L + +A NK+ E+ + L +L LDL N++
Sbjct: 32 LKSLEYLDLSFNKIKTLPTEITYLIGLISMKVANNKLVELPSAMTSLSRLECLDLSNNRL 211
Query: 312 STAKCLGQL-AANYNSLQAINLDGN 335
++ LG L A+ + LQ +NL N
Sbjct: 212 TS---LGSLELASMHRLQNLNLQYN 277
>AW719626
Length = 598
Score = 34.7 bits (78), Expect = 0.035
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 1/68 (1%)
Frame = +3
Query: 216 NLAGNSIVRITAGALP-RGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLA 274
++A NS G++P L SLNLS N +++ L + L+ LDLS+N + + G
Sbjct: 390 SIALNSKPTSQNGSIPISSLQSLNLSHNRFTNLLHLSAFSNLKSLDLSHNNLGTLPSGFQ 569
Query: 275 SCSSLKEL 282
+ + L L
Sbjct: 570 NLTKLHHL 593
Score = 31.2 bits (69), Expect = 0.38
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Frame = +3
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI-EGLRELTRLRVLD 260
SL+ LNL+ N + + L SL+LS NN+ ++ G + LT+L LD
Sbjct: 444 SLQSLNLSHNRFTNLLHLSAFSNLKSLDLSHNNLGTLPSGFQNLTKLHHLD 596
Score = 30.0 bits (66), Expect = 0.86
Identities = 34/144 (23%), Positives = 59/144 (40%), Gaps = 14/144 (9%)
Frame = +3
Query: 184 SSLTANASAAQLAN--------HGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPR--- 232
SS ++N S++Q+ + G + +L L++L+L+GN + G +P
Sbjct: 171 SSSSSNCSSSQIRSIELPSKNLRGSISWKYLKNMTKLEILDLSGNYL----QGQIPNWFW 338
Query: 233 ---GLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI 289
L +NLS+N + ++ N +G SSL+ L L+ N+
Sbjct: 339 ESSSLLVVNLSKNWLGG-------------SIALNSKPTSQNGSIPISSLQSLNLSHNRF 479
Query: 290 GEVEGLHRLLKLSILDLRFNKIST 313
+ L L LDL N + T
Sbjct: 480 TNLLHLSAFSNLKSLDLSHNNLGT 551
>AV767545
Length = 562
Score = 33.9 bits (76), Expect = 0.059
Identities = 31/92 (33%), Positives = 47/92 (50%), Gaps = 10/92 (10%)
Frame = -1
Query: 211 SLKVLNLAGNSIVRITAGALPRG------LHSLNLSRNNISSI--EGLRELTRLRVLDLS 262
SLKVLNL N+I +G +P L L++SRN+I L +L L+ LD+S
Sbjct: 538 SLKVLNLGSNNI----SGPIPVSISNLIDLERLDISRNHILGAIPSSLGQLLELQWLDVS 371
Query: 263 YNRIL-RIGHGLASCSSLKELYLAGNKI-GEV 292
N + I L+ ++LK N++ GE+
Sbjct: 370 INSLAGSIPSSLSQITNLKHASFRANRLCGEI 275
>TC11732
Length = 530
Score = 33.9 bits (76), Expect = 0.059
Identities = 27/82 (32%), Positives = 46/82 (55%), Gaps = 2/82 (2%)
Frame = +2
Query: 256 LRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE-GLHRLLKLSILDLRFNKI-ST 313
+R LDL++NRI+ I ++ +++ L LA N I + L +L L +++L N+I S
Sbjct: 254 VRTLDLTHNRIVDIPLEISKLVNVQRLVLADNLIERLPVNLGKLQSLKLVNLDGNRITSL 433
Query: 314 AKCLGQLAANYNSLQAINLDGN 335
LGQL L+ +++ GN
Sbjct: 434 PDELGQLV----RLERLSISGN 487
Score = 32.3 bits (72), Expect = 0.17
Identities = 26/86 (30%), Positives = 47/86 (54%), Gaps = 2/86 (2%)
Frame = +2
Query: 230 LPRGLHSLNLSRNNISSIE-GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNK 288
L R + +L+L+ N I I + +L ++ L L+ N I R+ L SLK + L GN+
Sbjct: 242 LDRSVRTLDLTHNRIVDIPLEISKLVNVQRLVLADNLIERLPVNLGKLQSLKLVNLDGNR 421
Query: 289 IGEV-EGLHRLLKLSILDLRFNKIST 313
I + + L +L++L L + N +++
Sbjct: 422 ITSLPDELGQLVRLERLSISGNLLTS 499
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 32.3 bits (72), Expect = 0.17
Identities = 26/63 (41%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Frame = +2
Query: 253 LTRLRVLDLSYNRILR-IGHGLASCSSLKELYLAGNKI-GEV-EGLHRLLKLSILDLRFN 309
L LR L L +N + + LA+CSSL+ LYL N + GE+ L RL L L+L N
Sbjct: 320 LPHLRTLSLRFNALSGPLPSDLAACSSLRNLYLQQNLLSGELPPALSRLTGLVRLNLASN 499
Query: 310 KIS 312
S
Sbjct: 500 NFS 508
>BG662016
Length = 459
Score = 30.8 bits (68), Expect = 0.50
Identities = 34/126 (26%), Positives = 61/126 (47%), Gaps = 12/126 (9%)
Frame = +1
Query: 240 SRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGL--H 296
S N ++S+ + L++L++L++S N I + + +CS+L+EL NK+ ++
Sbjct: 13 SSNQLTSLPNSIGCLSKLKLLNVSGNFIESLPKTIENCSALEELNANFNKLSKLPDTIGF 192
Query: 297 RLLKLSILDLRFNKI-------STAKCLGQLAANYNSLQAI--NLDGNPCQKNVGDEQLK 347
L+ L L + NK+ S L L A N L+A+ NL+ + + Q
Sbjct: 193 ELINLKKLAVNSNKLVLLPSSTSHLTALTVLDARLNCLRALPENLENLINLETLNVSQNF 372
Query: 348 KYLQGL 353
+YL L
Sbjct: 373 RYLDTL 390
>TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, partial
(24%)
Length = 702
Score = 30.8 bits (68), Expect = 0.50
Identities = 31/92 (33%), Positives = 44/92 (47%), Gaps = 8/92 (8%)
Frame = +2
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPR-----GLHSLNLSRNNISSI--EGLRELTRLRV 258
L++ L+VL+L N G +P G+ L LS NN S L LTRL
Sbjct: 374 LTSLTQLRVLSLKNNRF----HGPIPNLSNLTGMRLLFLSHNNFSGDFPVTLTSLTRLYR 541
Query: 259 LDLSYNRIL-RIGHGLASCSSLKELYLAGNKI 289
LDLS+N + I + + + L L L GN++
Sbjct: 542 LDLSHNSLSGEIPAAVNNFTRLLTLRLDGNQL 637
>TC9214
Length = 587
Score = 29.6 bits (65), Expect = 1.1
Identities = 17/42 (40%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Frame = -1
Query: 398 SHGVAAARRPPSTSTHSHRSQTVESP----KLSKGKQSHLPP 435
SH A+ PP+ S+ H SQ+ ESP K S S PP
Sbjct: 191 SHSKETAQNPPTHSSP*HTSQSAESPSSAHKNSSSPSSSTPP 66
>AV426912
Length = 429
Score = 28.5 bits (62), Expect = 2.5
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = -1
Query: 4 RSLPNIKASTLSPENHAFKHPSS 26
R LP+ ST+SP H F HP S
Sbjct: 156 RRLPSTPFSTISPVPHGFSHPPS 88
>BG662453
Length = 460
Score = 28.1 bits (61), Expect = 3.3
Identities = 18/61 (29%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Frame = +2
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKI 311
L + VLD+ N++ + + + S LK L ++GN I + + + L L+ FNK+
Sbjct: 35 LLNMVVLDVHSNQLRSLPNSVGCLSKLKVLNVSGNLIEYLPKSIENCRALEELNANFNKL 214
Query: 312 S 312
S
Sbjct: 215 S 217
>TC13940
Length = 808
Score = 28.1 bits (61), Expect = 3.3
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = -1
Query: 408 PSTSTHSHRSQTVESPKLSKGKQSHLPP 435
P+ STH S ++ KL GK S LPP
Sbjct: 646 PAPSTHLSASPVFDTIKLGSGKGSSLPP 563
>AV411649
Length = 358
Score = 27.7 bits (60), Expect = 4.3
Identities = 16/56 (28%), Positives = 33/56 (58%)
Frame = +2
Query: 203 VPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRV 258
+PF+ SLKV++L+ +I G LP+G ++ S +++++ L++L L +
Sbjct: 173 IPFIGMMPSLKVVDLSKTNI----TGFLPQGDRDISFS---LTALQDLKQLESLNM 319
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,290,493
Number of Sequences: 28460
Number of extensions: 92462
Number of successful extensions: 476
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 465
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 471
length of query: 475
length of database: 4,897,600
effective HSP length: 94
effective length of query: 381
effective length of database: 2,222,360
effective search space: 846719160
effective search space used: 846719160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0203.3


