

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
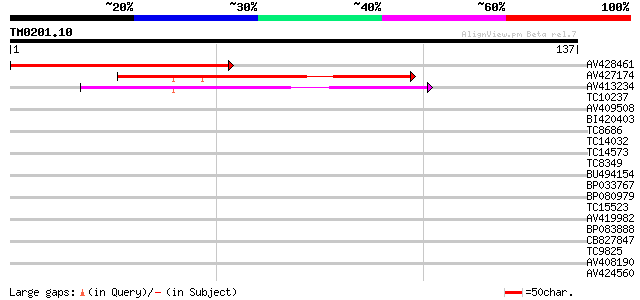
Score E
Sequences producing significant alignments: (bits) Value
AV428461 111 4e-26
AV427174 75 5e-15
AV413234 56 2e-09
TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precu... 30 0.15
AV409508 29 0.25
BI420403 28 0.56
TC8686 similar to UP|O23724 (O23724) GAI protein, partial (22%) 28 0.56
TC14032 homologue to UP|P93747 (P93747) At2g41990 protein, parti... 28 0.73
TC14573 homologue to UP|Q9MB42 (Q9MB42) Beta-amyrin synthase, co... 28 0.73
TC8349 27 0.95
BU494154 27 0.95
BP033767 27 0.95
BP080979 27 0.95
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 27 0.95
AV419982 27 1.2
BP083888 27 1.2
CB827847 27 1.2
TC9825 similar to UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (... 27 1.2
AV408190 27 1.2
AV424560 27 1.2
>AV428461
Length = 263
Score = 111 bits (278), Expect = 4e-26
Identities = 48/54 (88%), Positives = 51/54 (93%)
Frame = +2
Query: 1 MQEVLFAKQACICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNKVREWSE 54
MQEVLFAK+ACICYLIPC SSSETSLSWWERVRSPENKE WWA GW+KVR+WSE
Sbjct: 101 MQEVLFAKRACICYLIPCFSSSETSLSWWERVRSPENKERWWARGWSKVRDWSE 262
>AV427174
Length = 421
Score = 74.7 bits (182), Expect = 5e-15
Identities = 36/78 (46%), Positives = 48/78 (61%), Gaps = 6/78 (7%)
Frame = +3
Query: 27 SWWERVRSPENK---EWWWAHG---WNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASY 80
SWW+RVR+ + + WW+ G KVREWSEI+ GP+WKTFIRR + + S+
Sbjct: 204 SWWQRVRTSHSTVSGDRWWSRGIRALKKVREWSEILAGPRWKTFIRRLSHHR------SH 365
Query: 81 DKKGSFHYDSLSYALNFD 98
+ + YD SYALNFD
Sbjct: 366 KRMTKYQYDPFSYALNFD 419
>AV413234
Length = 378
Score = 56.2 bits (134), Expect = 2e-09
Identities = 32/94 (34%), Positives = 40/94 (42%), Gaps = 9/94 (9%)
Frame = +1
Query: 18 CSSSSETSLSWWERVRSPENK---------EWWWAHGWNKVREWSEIIVGPKWKTFIRRF 68
C +L WW+ + E W KV+E SE+I GPKWK FIR+
Sbjct: 109 CGCFKGFNLKWWQSHEEEGKRLIDQNGGGGEGWMVEKMKKVKEASEVIAGPKWKNFIRKI 288
Query: 69 NRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEE 102
+ KK F YD SYALNF+ E
Sbjct: 289 ---------SGQQKKKRFQYDEQSYALNFNSRAE 363
>TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75)
(NGM-75), partial (49%)
Length = 803
Score = 30.0 bits (66), Expect = 0.15
Identities = 13/31 (41%), Positives = 16/31 (50%), Gaps = 2/31 (6%)
Frame = -3
Query: 40 WWWAHGWNKVREWSEIIVG--PKWKTFIRRF 68
WWW W +R W I G +W F+RRF
Sbjct: 240 WWWFLIWRWIRRWWFPIWG*VHRWWFFLRRF 148
>AV409508
Length = 426
Score = 29.3 bits (64), Expect = 0.25
Identities = 11/26 (42%), Positives = 12/26 (45%), Gaps = 3/26 (11%)
Frame = -2
Query: 28 WWERVRSPENKEWWWAHG---WNKVR 50
WW R EWWW G W +VR
Sbjct: 92 WWWWFREKRGGEWWWECGGEEWREVR 15
>BI420403
Length = 425
Score = 28.1 bits (61), Expect = 0.56
Identities = 16/55 (29%), Positives = 22/55 (39%), Gaps = 14/55 (25%)
Frame = +2
Query: 6 FAKQACICYLIPCSSSSETSLSWW-ERVRSPENKE-------------WWWAHGW 46
F QA +C+ I S+E S WW E + ++ WWW GW
Sbjct: 227 FEDQA-LCFCIELGKSAE*SKPWW*ESMEGTQDNGACG*SSSNKWWWWWWWRGGW 388
>TC8686 similar to UP|O23724 (O23724) GAI protein, partial (22%)
Length = 786
Score = 28.1 bits (61), Expect = 0.56
Identities = 11/42 (26%), Positives = 18/42 (42%)
Frame = -3
Query: 11 CICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNKVREW 52
C+C+ + E L + E + N+ WWW W + W
Sbjct: 466 CVCWR---KGNQERRLLFPEWL*GRRNRRWWWLPEWGRNWNW 350
>TC14032 homologue to UP|P93747 (P93747) At2g41990 protein, partial (20%)
Length = 572
Score = 27.7 bits (60), Expect = 0.73
Identities = 14/34 (41%), Positives = 17/34 (49%), Gaps = 1/34 (2%)
Frame = -3
Query: 19 SSSSETSLSWWERVRSPENKEW-WWAHGWNKVRE 51
SSSS +S WW E+ W WW G +RE
Sbjct: 384 SSSSRSSSVWW*SFFHEESGWWFWWRRGRGFLRE 283
>TC14573 homologue to UP|Q9MB42 (Q9MB42) Beta-amyrin synthase, complete
Length = 2946
Score = 27.7 bits (60), Expect = 0.73
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = -2
Query: 5 LFAKQACICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNKVREW 52
LF+ IC+++ S+ S TSL +RVRS + +H + ++ W
Sbjct: 1340 LFSSS*WICFIVTSSAFSLTSLLKGQRVRSGSVNRYKLSHIKSCIKGW 1197
>TC8349
Length = 749
Score = 27.3 bits (59), Expect = 0.95
Identities = 11/30 (36%), Positives = 14/30 (46%), Gaps = 1/30 (3%)
Frame = +1
Query: 24 TSLSWWERVRSPENKEWWWAHGW-NKVREW 52
TS+ WW + KEWW W + R W
Sbjct: 133 TSVPWWIPWKW*HGKEWWGGLWWPDSTRSW 222
>BU494154
Length = 495
Score = 27.3 bits (59), Expect = 0.95
Identities = 9/21 (42%), Positives = 10/21 (46%)
Frame = +3
Query: 22 SETSLSWWERVRSPENKEWWW 42
SE WW SP+ WWW
Sbjct: 420 SERQQRWWGARASPDLTFWWW 482
>BP033767
Length = 541
Score = 27.3 bits (59), Expect = 0.95
Identities = 11/26 (42%), Positives = 13/26 (49%), Gaps = 7/26 (26%)
Frame = -2
Query: 40 WWWAHGWNK-------VREWSEIIVG 58
WWW GW K VRE ++VG
Sbjct: 270 WWWLWGWRKMGILGLEVREMVMVVVG 193
>BP080979
Length = 352
Score = 27.3 bits (59), Expect = 0.95
Identities = 10/30 (33%), Positives = 14/30 (46%)
Frame = -3
Query: 19 SSSSETSLSWWERVRSPENKEWWWAHGWNK 48
S + S SW E + +P WWW W +
Sbjct: 302 SQRARRSASWREDLSTPLRALWWWWWCWXR 213
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 27.3 bits (59), Expect = 0.95
Identities = 11/38 (28%), Positives = 18/38 (46%), Gaps = 3/38 (7%)
Frame = +1
Query: 28 WW---ERVRSPENKEWWWAHGWNKVREWSEIIVGPKWK 62
WW R+R + WWW ++ R W ++ +WK
Sbjct: 703 WW*RRRRIRRSS*RWWWWLWWQSRRRRWISLL--*RWK 810
>AV419982
Length = 415
Score = 26.9 bits (58), Expect = 1.2
Identities = 11/44 (25%), Positives = 16/44 (36%), Gaps = 9/44 (20%)
Frame = -1
Query: 14 YLIPCSSSSETSLSWWERVRSPENKE---------WWWAHGWNK 48
+L+PC L W + E +E WWW W +
Sbjct: 379 FLVPCFFQKMKGLMWRRKEEEEEEEEERGRGESKGWWWR*TWRE 248
>BP083888
Length = 296
Score = 26.9 bits (58), Expect = 1.2
Identities = 10/17 (58%), Positives = 10/17 (58%), Gaps = 2/17 (11%)
Frame = -1
Query: 38 KEWWWAHGWNKV--REW 52
KEWWW HG V R W
Sbjct: 53 KEWWWRHGHECVTARSW 3
>CB827847
Length = 503
Score = 26.9 bits (58), Expect = 1.2
Identities = 12/33 (36%), Positives = 17/33 (51%)
Frame = +3
Query: 15 LIPCSSSSETSLSWWERVRSPENKEWWWAHGWN 47
L+ SSSS +S S+ + + WWW WN
Sbjct: 15 LLRFSSSSPSSSSYSSSLNRFSHSWWWWC**WN 113
>TC9825 similar to UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (44%)
Length = 848
Score = 26.9 bits (58), Expect = 1.2
Identities = 13/40 (32%), Positives = 20/40 (49%), Gaps = 1/40 (2%)
Frame = +3
Query: 24 TSLSWWERVRSPENKEWWWAH-GWNKVREWSEIIVGPKWK 62
T SWW R ++ W W+H K+R+ + +V K K
Sbjct: 81 TQRSWWRT*RRRQSGTWMWSHQRKKKIRKRKKEVVRRKRK 200
Score = 25.0 bits (53), Expect = 4.7
Identities = 8/15 (53%), Positives = 9/15 (59%)
Frame = -2
Query: 31 RVRSPENKEWWWAHG 45
RVR + WWW HG
Sbjct: 511 RVRLHVERLWWWLHG 467
>AV408190
Length = 382
Score = 26.9 bits (58), Expect = 1.2
Identities = 12/33 (36%), Positives = 19/33 (57%), Gaps = 2/33 (6%)
Frame = -2
Query: 15 LIPCSSSSETSLSWWER--VRSPENKEWWWAHG 45
L P ++S + L WWER ++ +KEW+ G
Sbjct: 219 LRPVAASMKAVLPWWEREEMKVVTSKEWFGGRG 121
>AV424560
Length = 234
Score = 26.9 bits (58), Expect = 1.2
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 3/25 (12%)
Frame = -2
Query: 8 KQACIC---YLIPCSSSSETSLSWW 29
+ +C+C LIPC+SSS SL W
Sbjct: 92 RASCLCTSSMLIPCTSSSSKSLKPW 18
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,121,722
Number of Sequences: 28460
Number of extensions: 53165
Number of successful extensions: 642
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 617
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 634
length of query: 137
length of database: 4,897,600
effective HSP length: 81
effective length of query: 56
effective length of database: 2,592,340
effective search space: 145171040
effective search space used: 145171040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0201.10


