

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
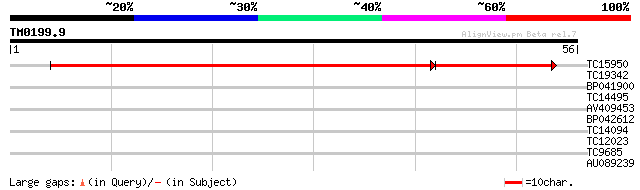
Query= TM0199.9
(56 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC15950 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase ... 82 2e-17
TC19342 26 1.4
BP041900 26 1.4
TC14495 UP|O22618 (O22618) Aspartate aminotransferase , complete 26 1.8
AV409453 25 2.4
BP042612 24 5.3
TC14094 homologue to emb|X02559.1|MIOBRN26 Oenothera berteriana ... 24 6.9
TC12023 24 6.9
TC9685 23 9.1
AU089239 23 9.1
>TC15950 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase subunit C
(V-ATPase C subunit) (Vacuolar proton pump C subunit)
, partial (48%)
Length = 562
Score = 82.0 bits (201), Expect(2) = 2e-17
Identities = 36/38 (94%), Positives = 38/38 (99%)
Frame = +1
Query: 5 WWVVSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIP 42
+WVVSLPVHNSASELWN+LQQNVSKHSFDTPLYRFNIP
Sbjct: 31 YWVVSLPVHNSASELWNQLQQNVSKHSFDTPLYRFNIP 144
Score = 20.4 bits (41), Expect(2) = 2e-17
Identities = 9/12 (75%), Positives = 11/12 (91%)
Frame = +2
Query: 43 ISGSEP*TLSAM 54
IS SEP*TLS++
Sbjct: 146 ISASEP*TLSSL 181
>TC19342
Length = 548
Score = 26.2 bits (56), Expect = 1.4
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = +2
Query: 12 VHNSASELWNRLQQNVSKHS 31
+HN+ E WNRLQ + K S
Sbjct: 380 IHNNRDEGWNRLQGPIHKRS 439
>BP041900
Length = 282
Score = 26.2 bits (56), Expect = 1.4
Identities = 13/45 (28%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Frame = -2
Query: 1 GAVVWWVVSLPVHNSASELWNRLQ--QNVSKHSFDTPLYRFNIPI 43
GA+ WW+ + VH + ++ RL+ +K D ++ F +PI
Sbjct: 227 GAIKWWIGNAAVHGKFATVFARLRLPNYDAKGVSDMGVHAFIVPI 93
>TC14495 UP|O22618 (O22618) Aspartate aminotransferase , complete
Length = 2030
Score = 25.8 bits (55), Expect = 1.8
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = -3
Query: 8 VSLPVHNSASELWNRLQQNVSKHSFDTPLY 37
+S P +NSA L N Q ++ ++S +PL+
Sbjct: 489 LSAPSNNSAVALLNAAQPSIGRYSLFSPLF 400
>AV409453
Length = 402
Score = 25.4 bits (54), Expect = 2.4
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +3
Query: 13 HNSASELWNRLQQNVSKHSFDTPLYR 38
H+S+S+L L + S SFD+PL+R
Sbjct: 69 HSSSSKLSISLSGSPSSASFDSPLHR 146
>BP042612
Length = 351
Score = 24.3 bits (51), Expect = 5.3
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +3
Query: 22 RLQQNVSKHSFDT 34
RL QN+SKH +DT
Sbjct: 207 RLHQNLSKHMYDT 245
>TC14094 homologue to emb|X02559.1|MIOBRN26 Oenothera berteriana
mitochondrial gene for 26S ribosomal RNA, partial (83%)
Length = 3140
Score = 23.9 bits (50), Expect = 6.9
Identities = 7/27 (25%), Positives = 14/27 (50%)
Frame = -2
Query: 6 WVVSLPVHNSASELWNRLQQNVSKHSF 32
W P+HN ++E W + ++ + F
Sbjct: 361 WFFHHPIHN*SNETWRKRSEHFAAEEF 281
>TC12023
Length = 553
Score = 23.9 bits (50), Expect = 6.9
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +1
Query: 9 SLPVHNSASELWNRLQQNVSKHSFDTPL 36
+LPV + + L N L NV++ S TPL
Sbjct: 466 TLPVSHMTAMLCNALIPNVNRFSAPTPL 549
>TC9685
Length = 919
Score = 23.5 bits (49), Expect = 9.1
Identities = 12/33 (36%), Positives = 15/33 (45%)
Frame = +3
Query: 13 HNSASELWNRLQQNVSKHSFDTPLYRFNIPISG 45
H+S++ L L S H F PL F SG
Sbjct: 21 HSSSTRLLLML*NQSSSHEFHNPLAPFTFACSG 119
>AU089239
Length = 500
Score = 23.5 bits (49), Expect = 9.1
Identities = 6/16 (37%), Positives = 13/16 (80%)
Frame = -1
Query: 6 WVVSLPVHNSASELWN 21
W+++LP+H A++ W+
Sbjct: 95 WILNLPIHFVANQPWS 48
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.135 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,156,342
Number of Sequences: 28460
Number of extensions: 12461
Number of successful extensions: 104
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 104
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 104
length of query: 56
length of database: 4,897,600
effective HSP length: 32
effective length of query: 24
effective length of database: 3,986,880
effective search space: 95685120
effective search space used: 95685120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0199.9


