

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.8
(242 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
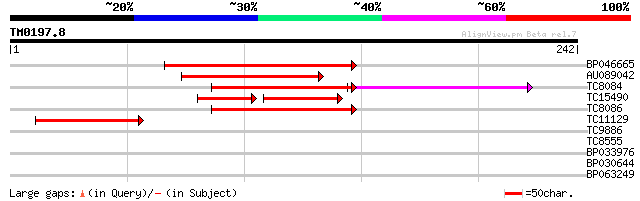
Score E
Sequences producing significant alignments: (bits) Value
BP046665 87 2e-18
AU089042 69 9e-13
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 60 8e-12
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 46 2e-10
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 58 1e-09
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 42 1e-04
TC9886 similar to UP|Q94K08 (Q94K08) Nucleotide pyrophosphatase-... 38 0.001
TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin ... 29 0.83
BP033976 27 2.4
BP030644 25 9.2
BP063249 25 9.2
>BP046665
Length = 524
Score = 87.4 bits (215), Expect = 2e-18
Identities = 45/82 (54%), Positives = 52/82 (62%)
Frame = -1
Query: 67 GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHIS 126
G+ NHK+ LK PIMLLRNI QA G CNGTRLIV +LG I ATVIT TN D + I
Sbjct: 509 GIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGDDIFIP 330
Query: 127 IMDLVPSDPNFQLNSEEDSFQL 148
MD+VPSD + E F +
Sbjct: 329 RMDMVPSDSGYPFKFERRQFPI 264
>AU089042
Length = 191
Score = 68.6 bits (166), Expect = 9e-13
Identities = 32/61 (52%), Positives = 45/61 (73%)
Frame = +3
Query: 74 LLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPS 133
+LK +P+ML+ N+ + GLCNGTRLIV LG IGAT+++GT+ V+IS+M+L PS
Sbjct: 6 VLKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPS 185
Query: 134 D 134
D
Sbjct: 186 D 188
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 60.1 bits (144), Expect(2) = 8e-12
Identities = 31/62 (50%), Positives = 38/62 (61%)
Frame = +1
Query: 87 IDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPNFQLNSEEDSF 146
I + LCNGTRLIVV+LG I ATVITGTN D + I +D+VPSD + E F
Sbjct: 43 IQKFENLCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 222
Query: 147 QL 148
+
Sbjct: 223 PI 228
Score = 25.4 bits (54), Expect(2) = 8e-12
Identities = 24/79 (30%), Positives = 37/79 (46%)
Frame = +2
Query: 145 SFQLQYALR*P*TKANDKHYLRWGSSFRDPSSLMVSYM*LSLEYAQERVLNFEYWNSQAN 204
SF+ AL+* * K HY F S MVS M LS QE+ N+ W +
Sbjct: 218 SFRSVCALQ*L*IKVKVSHYPTLAYIFLALYSHMVSCMLLSPG*DQEKDSNY*CWMRKKK 397
Query: 205 NRPVL*M*CLRKFLEMFDV 223
+ + * K+L+++++
Sbjct: 398 *QTLQRT*FTEKYLKIYEI 454
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 46.2 bits (108), Expect(2) = 2e-10
Identities = 22/25 (88%), Positives = 24/25 (96%)
Frame = +1
Query: 81 IMLLRNIDQAAGLCNGTRLIVVELG 105
IMLLRNI QA+GLCNGTRLIVV+LG
Sbjct: 58 IMLLRNIVQASGLCNGTRLIVVDLG 132
Score = 34.3 bits (77), Expect(2) = 2e-10
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = +2
Query: 109 IGATVITGTNANDKVHISIMDLVPSDPNFQLNSE 142
I TVITGT+ D + I MD+VPSD ++ E
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFE 238
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 58.2 bits (139), Expect = 1e-09
Identities = 30/62 (48%), Positives = 38/62 (60%)
Frame = +1
Query: 87 IDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPNFQLNSEEDSF 146
I + LC+GTRLIVV+LG I ATVITGTN D + I +D+VPSD + E F
Sbjct: 151 IQKFENLCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 330
Query: 147 QL 148
+
Sbjct: 331 PI 336
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 41.6 bits (96), Expect = 1e-04
Identities = 19/46 (41%), Positives = 30/46 (64%)
Frame = +2
Query: 12 PTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT 57
PTL++VE +N +ML ++ EYLSS++ + E +E+ WFTT
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WFTT 595
>TC9886 similar to UP|Q94K08 (Q94K08) Nucleotide pyrophosphatase-like
protein, partial (50%)
Length = 685
Score = 38.1 bits (87), Expect = 0.001
Identities = 17/20 (85%), Positives = 17/20 (85%)
Frame = -2
Query: 223 VYNTISFIYFIFLLIYYYTI 242
VYNTISFIY IFL IYYY I
Sbjct: 684 VYNTISFIYSIFLFIYYYMI 625
>TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} , partial
(83%)
Length = 1776
Score = 28.9 bits (63), Expect = 0.83
Identities = 23/70 (32%), Positives = 33/70 (46%)
Frame = +1
Query: 102 VELGERYIGATVITGTNANDKVHISIMDLVPSDPNFQLNSEEDSFQLQYALR*P*TKAND 161
V+L + IT TN N +++ S +LV PNF ++ + F QY P ND
Sbjct: 976 VKLNYKEYWRVKITITNFNYRMNYSQWNLVVQHPNF--DNLTEIFSFQYKSLTPYEGLND 1149
Query: 162 KHYLRWGSSF 171
L WG+ F
Sbjct: 1150 TGML-WGTKF 1176
>BP033976
Length = 551
Score = 27.3 bits (59), Expect = 2.4
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +1
Query: 120 NDKVHISIMDLVPSDPNFQLNSEEDSFQLQYALR 153
N+K HI+I +P D N EE + + QY R
Sbjct: 25 NEKNHINIAHQIPQDKT*TTNQEESN*KYQYHCR 126
>BP030644
Length = 288
Score = 25.4 bits (54), Expect = 9.2
Identities = 7/18 (38%), Positives = 14/18 (76%)
Frame = -3
Query: 223 VYNTISFIYFIFLLIYYY 240
V+ S++YF+FL ++Y+
Sbjct: 55 VFGMTSYLYFLFLFLFYF 2
>BP063249
Length = 515
Score = 25.4 bits (54), Expect = 9.2
Identities = 12/46 (26%), Positives = 26/46 (56%)
Frame = -3
Query: 193 VLNFEYWNSQANNRPVL*M*CLRKFLEMFDVYNTISFIYFIFLLIY 238
V+ ++W + R + + +F+ +F + +SF++FI+ LIY
Sbjct: 270 VMGLQWWGLKR*GRLQGVICFIFRFMFLFTLEILLSFLFFIYALIY 133
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.150 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,702,174
Number of Sequences: 28460
Number of extensions: 42825
Number of successful extensions: 352
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 352
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 352
length of query: 242
length of database: 4,897,600
effective HSP length: 88
effective length of query: 154
effective length of database: 2,393,120
effective search space: 368540480
effective search space used: 368540480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0197.8


