

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195b.8
(83 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
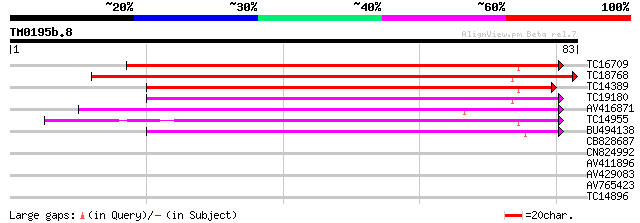
Score E
Sequences producing significant alignments: (bits) Value
TC16709 weakly similar to PIR|T47573|T47573 peptide transport-li... 76 1e-15
TC18768 60 7e-11
TC14389 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter... 57 8e-10
TC19180 similar to PIR|T10255|T10255 nitrite transport protein, ... 52 2e-08
AV416871 49 2e-07
TC14955 UP|O22305 (O22305) Peptide transporter, complete 47 6e-07
BU494138 46 1e-06
CB828687 36 0.001
CN824992 33 0.007
AV411896 31 0.046
AV429083 25 3.3
AV765423 25 3.3
TC14896 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protei... 24 5.6
>TC16709 weakly similar to PIR|T47573|T47573 peptide transport-like protein
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(41%)
Length = 712
Score = 75.9 bits (185), Expect = 1e-15
Identities = 37/66 (56%), Positives = 47/66 (71%), Gaps = 2/66 (3%)
Frame = +1
Query: 18 DYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSFG--SF 75
D +PWRL T TQVEELK ++R+LP+WAT I+F+ VY +STL V QG MN G +F
Sbjct: 277 DSVDPWRLCTETQVEELKSILRMLPVWATGIIFATVYGQMSTLFVLQGQTMNTHVGNSNF 456
Query: 76 NIPPAS 81
IPPA+
Sbjct: 457 KIPPAA 474
>TC18768
Length = 738
Score = 60.1 bits (144), Expect = 7e-11
Identities = 32/72 (44%), Positives = 48/72 (66%), Gaps = 1/72 (1%)
Frame = +2
Query: 13 ESRSGDYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSF 72
E+ S + WRL +V QVE++KIL+ V+PI++ IVF+ + A L T V+QG+ M+
Sbjct: 284 EAGSNTKKSSWRLCSVGQVEQVKILLSVIPIFSCTIVFNTILAQLQTFSVQQGSAMDTHL 463
Query: 73 -GSFNIPPASCE 83
SF+IPPAS +
Sbjct: 464 TKSFHIPPASLQ 499
>TC14389 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter, partial
(72%)
Length = 1990
Score = 56.6 bits (135), Expect = 8e-10
Identities = 28/61 (45%), Positives = 41/61 (66%), Gaps = 1/61 (1%)
Frame = +3
Query: 21 NPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSFG-SFNIPP 79
+PWRL TVTQVEE K + +++PI T ++ + TL ++QGT ++RS G +F IPP
Sbjct: 849 SPWRLCTVTQVEETKQMTKMVPILITTLIPCTMLIQAHTLFIKQGTTLDRSMGPNFEIPP 1028
Query: 80 A 80
A
Sbjct: 1029A 1031
>TC19180 similar to PIR|T10255|T10255 nitrite transport protein,
chloroplast - cucumber {Cucumis sativus;} , partial
(12%)
Length = 408
Score = 52.0 bits (123), Expect = 2e-08
Identities = 29/62 (46%), Positives = 37/62 (58%), Gaps = 1/62 (1%)
Frame = +1
Query: 21 NPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSF-GSFNIPP 79
N WRL TV +VEELK +IR+ PIWA+ I+ YA T ++Q M+R SF IP
Sbjct: 34 NKWRLNTVHRVEELKSIIRMGPIWASGILLITAYAQQGTFSLQQAKTMDRHITKSFQIPA 213
Query: 80 AS 81
S
Sbjct: 214GS 219
>AV416871
Length = 354
Score = 48.5 bits (114), Expect = 2e-07
Identities = 26/72 (36%), Positives = 40/72 (55%), Gaps = 1/72 (1%)
Frame = +3
Query: 11 DSESRSGDYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQG-TMMN 69
D E + + W+L VTQVE KI++ + P++ I+ + A + T V+QG TM
Sbjct: 105 DMELEKPESTSQWKLCRVTQVENAKIILSMGPVFLCTIIMTLCLAQIQTFSVQQGYTMDT 284
Query: 70 RSFGSFNIPPAS 81
+ FN+PPAS
Sbjct: 285 KITKDFNMPPAS 320
>TC14955 UP|O22305 (O22305) Peptide transporter, complete
Length = 2300
Score = 47.0 bits (110), Expect = 6e-07
Identities = 30/77 (38%), Positives = 46/77 (58%), Gaps = 1/77 (1%)
Frame = +1
Query: 6 IDLVSDSESRSGDYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQG 65
+D + ES + D +NP TVTQVE K+++ + IW ++ + +AL ST+ V QG
Sbjct: 919 LDRAAIRESNT-DLSNP--PCTVTQVEGTKLVLGMFQIWLLMLIPTNCWALESTIFVRQG 1089
Query: 66 TMMNRSFG-SFNIPPAS 81
T M+R+ G F +P AS
Sbjct: 1090TTMDRTLGPKFRLPAAS 1140
>BU494138
Length = 518
Score = 46.2 bits (108), Expect = 1e-06
Identities = 26/62 (41%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +2
Query: 21 NPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSFGS-FNIPP 79
N R+ ++ QVEE+K L R+ PIWA I+ A T V Q M+R GS F IP
Sbjct: 2 NSARVVSIQQVEEIKCLARIFPIWAAGILGFTAMAQQGTFTVSQAMKMDRHIGSKFQIPA 181
Query: 80 AS 81
S
Sbjct: 182GS 187
>CB828687
Length = 554
Score = 36.2 bits (82), Expect = 0.001
Identities = 15/29 (51%), Positives = 22/29 (75%)
Frame = +1
Query: 21 NPWRLFTVTQVEELKILIRVLPIWATEIV 49
+PWRL TVTQVEE K + +++PI T ++
Sbjct: 454 SPWRLCTVTQVEETKQMTKMVPILITTLI 540
>CN824992
Length = 708
Score = 33.5 bits (75), Expect = 0.007
Identities = 14/33 (42%), Positives = 23/33 (69%)
Frame = +2
Query: 8 LVSDSESRSGDYANPWRLFTVTQVEELKILIRV 40
L++ ES+ + + W L TVTQVEE+K ++R+
Sbjct: 608 LITSQESKQENECSLWHLSTVTQVEEVKCILRL 706
>AV411896
Length = 323
Score = 30.8 bits (68), Expect = 0.046
Identities = 15/29 (51%), Positives = 19/29 (64%)
Frame = +1
Query: 7 DLVSDSESRSGDYANPWRLFTVTQVEELK 35
D+ SD G ++PWRL T+ QVEELK
Sbjct: 247 DITSD-----GSASDPWRLCTIDQVEELK 318
>AV429083
Length = 316
Score = 24.6 bits (52), Expect = 3.3
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = -3
Query: 45 ATEIVFSAVYALLSTLLVEQGTMMN 69
+ E +FS V+ L T+ ++QGT+ N
Sbjct: 227 SVEYIFSLVHVLS*TIPIQQGTISN 153
>AV765423
Length = 395
Score = 24.6 bits (52), Expect = 3.3
Identities = 9/22 (40%), Positives = 17/22 (76%)
Frame = +1
Query: 23 WRLFTVTQVEELKILIRVLPIW 44
W++F+ +Q+ ELK+ ++ PIW
Sbjct: 259 WQIFSASQLMELKVHMK--PIW 318
>TC14896 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protein S18,
complete
Length = 906
Score = 23.9 bits (50), Expect = 5.6
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 1/65 (1%)
Frame = -3
Query: 14 SRSGDYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTM-MNRSF 72
S S +P LF T ++L +LPI E+ + ++ STL+ M N SF
Sbjct: 253 SNSAALNSPALLFMSTSAFLQQMLANLLPIPLMEVRANMIFCFPSTLVFSTRRMCWNSSF 74
Query: 73 GSFNI 77
+ +I
Sbjct: 73 ATRDI 59
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,359,397
Number of Sequences: 28460
Number of extensions: 12902
Number of successful extensions: 65
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 65
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 65
length of query: 83
length of database: 4,897,600
effective HSP length: 59
effective length of query: 24
effective length of database: 3,218,460
effective search space: 77243040
effective search space used: 77243040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0195b.8


