

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
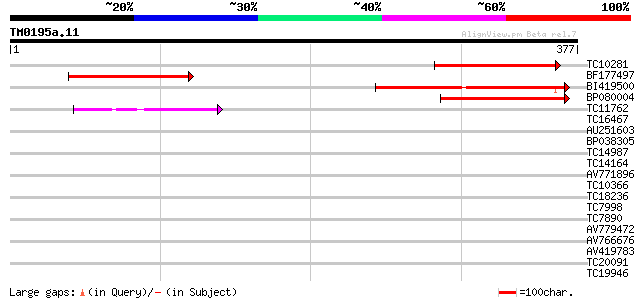
Score E
Sequences producing significant alignments: (bits) Value
TC10281 110 3e-25
BF177497 109 9e-25
BI419500 108 2e-24
BP080004 78 2e-15
TC11762 59 1e-09
TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase ch... 33 0.078
AU251603 30 0.50
BP038305 28 2.5
TC14987 28 2.5
TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, ... 28 2.5
AV771896 28 3.3
TC10366 28 3.3
TC18236 weakly similar to UP|Q9LH79 (Q9LH79) MtN3-like protein (... 28 3.3
TC7998 homologue to UP|HEM6_SOYBN (P35055) Coproporphyrinogen II... 27 4.3
TC7890 27 4.3
AV779472 27 5.6
AV766676 27 5.6
AV419783 27 7.3
TC20091 27 7.3
TC19946 27 7.3
>TC10281
Length = 510
Score = 110 bits (276), Expect = 3e-25
Identities = 52/84 (61%), Positives = 67/84 (78%)
Frame = +2
Query: 283 LSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL 342
LS FLFMSLV RP LF++SMK+FNVAWWAYSFP+T LA+AS +YA +VKG +H +MLVL
Sbjct: 2 LSFFLFMSLVSRPLLFKKSMKKFNVAWWAYSFPLTALALASAEYAHQVKGLTAHAVMLVL 181
Query: 343 LALSVLVSLALTVFTFINSKMLLP 366
+SVLVS+ L + T +N+ + LP
Sbjct: 182 SLISVLVSVVLMIVTALNTSIPLP 253
>BF177497
Length = 332
Score = 109 bits (272), Expect = 9e-25
Identities = 51/83 (61%), Positives = 63/83 (75%)
Frame = +1
Query: 40 NSVLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFT 99
+S+L+++HAGYFRISL+L QALLWK LI P D LR + +PS+AF LW LALFT
Sbjct: 82 SSILSQIHAGYFRISLALSSQALLWKVLIEPIEDAHALRKIFSFIPSTAFTFLWYLALFT 261
Query: 100 LALLSLLYLLRCLFHFNMVKAEF 122
L LS LY+LRC+FHF+MVK EF
Sbjct: 262 LVTLSFLYVLRCVFHFDMVKDEF 330
>BI419500
Length = 546
Score = 108 bits (269), Expect = 2e-24
Identities = 58/132 (43%), Positives = 84/132 (62%), Gaps = 3/132 (2%)
Frame = +1
Query: 244 LPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMK 303
LP L PVFFLF AAP VAS+AW I G FD S++ +F++LFL+ SL R FR
Sbjct: 1 LPKELHPVFFLFVAAPSVASMAWAKIQGSFDYGSRIAYFIALFLYFSLAVRINFFRGF-- 174
Query: 304 RFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINS-- 361
+F++AWWAY+FP+T A+A+ Y+ +V ++ L +VL +S L+ AL V T +++
Sbjct: 175 KFSLAWWAYTFPMTGAAIATIRYSNQVPNVVTKTLCVVLALISTLIVTALLVSTILHAFV 354
Query: 362 -KMLLPDDDPIA 372
+ L P+D IA
Sbjct: 355 FRDLFPNDIAIA 390
>BP080004
Length = 367
Score = 78.2 bits (191), Expect = 2e-15
Identities = 34/86 (39%), Positives = 57/86 (65%)
Frame = -3
Query: 287 LFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALS 346
LF+ CRP F++SM++ V WW SFP+T L +A +Y++EV+G+++ LMLV+ +S
Sbjct: 365 LFLMQACRPNFFKKSMRKLTVTWWVCSFPLTFLGLACAEYSQEVEGSMASALMLVICVVS 186
Query: 347 VLVSLALTVFTFINSKMLLPDDDPIA 372
VLV ++L + + + + LL PI+
Sbjct: 185 VLVFISLMLISVLKIERLLLKKAPIS 108
>TC11762
Length = 601
Score = 58.9 bits (141), Expect = 1e-09
Identities = 31/99 (31%), Positives = 51/99 (51%)
Frame = +2
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F I L + QA+LWKTL T + H+ V L+LW +++ +
Sbjct: 314 LLRFPVSSFGICLGVSSQAILWKTLA--TSPSTEFLHISLKVN----LILWIISVALITT 475
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLL 141
+ +Y L+ + +F V+ E+ H + VN+ FAPWI+ L
Sbjct: 476 VFAIYTLKIILYFEAVRREYYHPIRVNFFFAPWIALLFL 592
>TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase chain H,
chloroplast (NAD(P)H dehydrogenase, chain H)
(NADH-plastoquinone oxidoreductase 49 kDa subunit) ,
complete
Length = 2407
Score = 33.1 bits (74), Expect = 0.078
Identities = 30/90 (33%), Positives = 45/90 (49%), Gaps = 1/90 (1%)
Frame = +3
Query: 230 LFVTLYQRLSGGN-RLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
LFVT+ L G N +P + FF GV +G+ +G F TL+K LFLF
Sbjct: 2061 LFVTVLY-LGGSNISIPYISLFEFFEINKEYGV----FGTTIGIFITLAKTY----LFLF 2213
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+S++ R TL R M + W + P+++
Sbjct: 2214 VSIITRWTLPRLRMDQLLNLGWKFLLPISL 2303
>AU251603
Length = 350
Score = 30.4 bits (67), Expect = 0.50
Identities = 15/57 (26%), Positives = 30/57 (52%), Gaps = 8/57 (14%)
Frame = +2
Query: 92 LWSLALFTLALLSL-LYLLRCLFHFNMVKAEFLHHV-------GVNYLFAPWISWFL 140
+W+ A AL+++ +++ +FH + A L V G+NYL+ +++W L
Sbjct: 167 IWTYAALEFALVAITVHIFYSMFHLTAIYAILLSSVLGFGISMGINYLYIQYVTWRL 337
>BP038305
Length = 491
Score = 28.1 bits (61), Expect = 2.5
Identities = 20/64 (31%), Positives = 29/64 (45%)
Frame = +3
Query: 280 LFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLM 339
LF + L +F+ L+ RRS + +AWW F LA S G+ HV +
Sbjct: 261 LFLMILLVFIVLLSS----RRSWRLSFIAWWLRGFEFWRLAPRSA-------GSFEHVAL 407
Query: 340 LVLL 343
+V L
Sbjct: 408 IVCL 419
>TC14987
Length = 1038
Score = 28.1 bits (61), Expect = 2.5
Identities = 25/100 (25%), Positives = 44/100 (44%), Gaps = 9/100 (9%)
Frame = +3
Query: 36 LASLNSVLTKLHAGYFRISLSLGGQALLWKT---------LIGPTHDKSTLRHVVHMVPS 86
LA L++ L ++ G + S+G LL KT ++GP D ++
Sbjct: 66 LAILSTSLQQISIGSLQKKYSIGSFELLSKTAPIQAVSLLVLGPFIDYYLSAKLITNYKM 245
Query: 87 SAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHV 126
S+ +L+ L TLA+ + C+ F+ V + L H+
Sbjct: 246 SSGAILFILLSCTLAVFCNVSQYLCIGRFSAVSFQVLGHM 365
>TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, partial
(44%)
Length = 1606
Score = 28.1 bits (61), Expect = 2.5
Identities = 18/70 (25%), Positives = 32/70 (45%)
Frame = +2
Query: 158 LWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAV 217
L W F LD+ T +++S A NP+S L+ +VGAQA + +
Sbjct: 263 LHWTFDQATGKLDIAFRHTGITSTDKWVSWAINPSSNLN--SAMVGAQALVAIPQSSGSP 436
Query: 218 CLFSLGMVHY 227
+++ + +Y
Sbjct: 437 KVYTTSIANY 466
>AV771896
Length = 459
Score = 27.7 bits (60), Expect = 3.3
Identities = 19/68 (27%), Positives = 37/68 (53%)
Frame = -1
Query: 293 CRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLA 352
C PT+ ++ ++ +SF VTV ++ + +V G +L+L++ L +LV L
Sbjct: 231 CHPTV---AVPSPGFIFYVFSFVVTVTVLSLGHFPSDV*GL---MLLLLMNHLVILVCLN 70
Query: 353 LTVFTFIN 360
L V ++I+
Sbjct: 69 LLVLSYIS 46
>TC10366
Length = 421
Score = 27.7 bits (60), Expect = 3.3
Identities = 9/21 (42%), Positives = 16/21 (75%)
Frame = +1
Query: 98 FTLALLSLLYLLRCLFHFNMV 118
F+++L ++LYLL C+F F +
Sbjct: 274 FSMSLFTILYLLNCIFFFQYI 336
>TC18236 weakly similar to UP|Q9LH79 (Q9LH79) MtN3-like protein
(AT3g14770/T21E2_2), partial (42%)
Length = 1039
Score = 27.7 bits (60), Expect = 3.3
Identities = 31/122 (25%), Positives = 49/122 (39%), Gaps = 6/122 (4%)
Frame = +1
Query: 174 YGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVT 233
YG F L + N T I ++ A K CLF+ ++ + V+
Sbjct: 268 YGLPFVSPDNILVSTVNGTGAAIEIVYVLIFITLAPKKEKAKIFCLFTFVLLVFSVVIFV 447
Query: 234 LYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGS------IVGGFDTLSKMLFFLSLFL 287
L G +R F FAA +++ +GS +V ++ M FFLSLF+
Sbjct: 448 SLCALHGNSRK-------LFCGFAAAIFSAIMYGSPLSIMRLVIKTKSVEFMPFFLSLFV 606
Query: 288 FM 289
F+
Sbjct: 607 FL 612
>TC7998 homologue to UP|HEM6_SOYBN (P35055) Coproporphyrinogen III oxidase,
chloroplast precursor (Coproporphyrinogenase) (Coprogen
oxidase) , partial (88%)
Length = 1573
Score = 27.3 bits (59), Expect = 4.3
Identities = 21/71 (29%), Positives = 36/71 (50%), Gaps = 2/71 (2%)
Frame = +3
Query: 84 VPSSAFLVLWSLALF--TLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLL 141
V ++A + + LF L L+SLL+++ CL ++ LH + + + P L
Sbjct: 441 VVAAALAGFFKMVLFGRRLGLMSLLFMVSCLLMRIVLPKVLLHLLMRSLVLFP----SLQ 608
Query: 142 LQSAPFVAPKT 152
L+SA F P+T
Sbjct: 609 LESAQFCIPRT 641
>TC7890
Length = 232
Score = 27.3 bits (59), Expect = 4.3
Identities = 10/15 (66%), Positives = 14/15 (92%)
Frame = +3
Query: 279 MLFFLSLFLFMSLVC 293
++FF+SLFLF+SL C
Sbjct: 54 LIFFVSLFLFVSLAC 98
>AV779472
Length = 509
Score = 26.9 bits (58), Expect = 5.6
Identities = 20/77 (25%), Positives = 44/77 (56%), Gaps = 2/77 (2%)
Frame = -2
Query: 99 TLALLSLLYLLRCLFHFNMVKAEFLHHVGVN-YLFAPWISWFLLLQ-SAPFVAPKTATYL 156
T+ +++L L+ L+ N+V A+F+ H+ + YL+ + + + ++ FV Y
Sbjct: 409 TVIIINLFRLM--LYVNNLVLAKFVIHICYDIYLYMLSLFQYSVFS*TSSFVPLIM*IY* 236
Query: 157 VLWWVFAVPVVVLDVKI 173
++W +F VPV+ + V++
Sbjct: 235 IIWSLFNVPVIGMTVEL 185
>AV766676
Length = 544
Score = 26.9 bits (58), Expect = 5.6
Identities = 9/26 (34%), Positives = 16/26 (60%)
Frame = -2
Query: 153 ATYLVLWWVFAVPVVVLDVKIYGQWF 178
A + WWV +P+V+L K+Y ++
Sbjct: 135 ALIAMFWWVLKIPLVLLQ*KLYQVYY 58
>AV419783
Length = 422
Score = 26.6 bits (57), Expect = 7.3
Identities = 22/69 (31%), Positives = 28/69 (39%)
Frame = +3
Query: 90 LVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVA 149
++LW L F LL Y+ R L + K E HH + L W LL QS
Sbjct: 213 VLLWQLGSFLSHLLLFFYMGRSLMSSLLFKLEGSHH---HTLLQQW----LLQQSLALRE 371
Query: 150 PKTATYLVL 158
+LVL
Sbjct: 372 ASQIPFLVL 398
>TC20091
Length = 507
Score = 26.6 bits (57), Expect = 7.3
Identities = 18/70 (25%), Positives = 32/70 (45%), Gaps = 15/70 (21%)
Frame = +3
Query: 87 SAFLVLWSLALFTLALLSLLYLLRC-LFHFNMV------KAEFLH--------HVGVNYL 131
S+F++ W L++F + ++L+ L F+ K +F H H V YL
Sbjct: 15 SSFILSWPLSVFCFFVFIAIFLVHFPLLFFSATSLSPFFKIDFCHPFAFPSLCHTHVIYL 194
Query: 132 FAPWISWFLL 141
F W++ +L
Sbjct: 195 FKSWVNLIVL 224
>TC19946
Length = 485
Score = 26.6 bits (57), Expect = 7.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = -3
Query: 281 FFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVL 319
FFLSL+LF SL C T +S+ ++ + + V++L
Sbjct: 165 FFLSLYLFSSLTCS*T*CLQSINILSILYTKLVYYVSML 49
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,593,366
Number of Sequences: 28460
Number of extensions: 125745
Number of successful extensions: 962
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 954
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 960
length of query: 377
length of database: 4,897,600
effective HSP length: 92
effective length of query: 285
effective length of database: 2,279,280
effective search space: 649594800
effective search space used: 649594800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0195a.11


