

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.7
(848 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
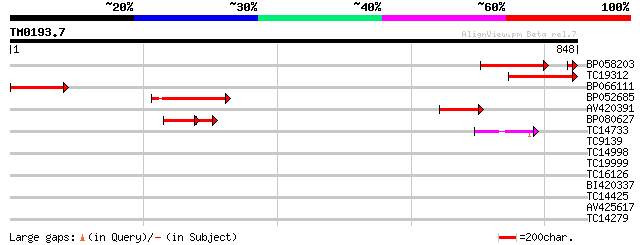
Score E
Sequences producing significant alignments: (bits) Value
BP058203 207 4e-57
TC19312 similar to UP|Q9XIS4 (Q9XIS4) Branching enzyme 3 , part... 198 3e-51
BP066111 169 2e-42
BP052685 157 5e-39
AV420391 84 1e-16
BP080627 66 4e-16
TC14733 similar to UP|Q41058 (Q41058) Starch branching enzyme I ... 64 1e-10
TC9139 similar to UP|Q9LNA8 (Q9LNA8) F5O11.10, partial (12%) 29 3.6
TC14998 similar to UP|Q9ZU81 (Q9ZU81) At2g48080 protein, partial... 28 4.7
TC19999 weakly similar to UP|AAQ89667 (AAQ89667) At5g25170, part... 28 6.1
TC16126 28 6.1
BI420337 28 7.9
TC14425 similar to PIR|T50796|T50796 chorismate mutase CM2 - Ara... 28 7.9
AV425617 28 7.9
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 28 7.9
>BP058203
Length = 581
Score = 207 bits (527), Expect(2) = 4e-57
Identities = 101/102 (99%), Positives = 101/102 (99%)
Frame = -1
Query: 704 GKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVY 763
GKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRP SFKILSPPRTCVVY
Sbjct: 581 GKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPXSFKILSPPRTCVVY 402
Query: 764 YRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKERE 805
YRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKERE
Sbjct: 401 YRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKERE 276
Score = 32.3 bits (72), Expect(2) = 4e-57
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = -3
Query: 835 ENEVFQDEVEDADE 848
ENEVFQDEVEDADE
Sbjct: 276 ENEVFQDEVEDADE 235
>TC19312 similar to UP|Q9XIS4 (Q9XIS4) Branching enzyme 3 , partial (4%)
Length = 523
Score = 198 bits (504), Expect = 3e-51
Identities = 100/102 (98%), Positives = 101/102 (98%)
Frame = +2
Query: 747 RPNSFKILSPPRTCVVYYRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKEREV 806
RP SF+ILSPPRTCVVYYRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKEREV
Sbjct: 2 RPXSFQILSPPRTCVVYYRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKEREV 181
Query: 807 SNFNWTMETLAAANADVAKIPDELVPAAENEVFQDEVEDADE 848
SNFNWTMETLAAANADVAKIPDELVPAAENEVFQDEVEDADE
Sbjct: 182 SNFNWTMETLAAANADVAKIPDELVPAAENEVFQDEVEDADE 307
>BP066111
Length = 266
Score = 169 bits (428), Expect = 2e-42
Identities = 86/87 (98%), Positives = 86/87 (98%)
Frame = +3
Query: 1 MITSFSLQSFNIASTAHNSRNKQDLAKQNSVELVLGYRNPKGCNRFSFGSRRSIHERVST 60
MITSFSLQSFNIASTAHNSRNKQDLAKQNSVELVLGYRNPKGCNRFSFGSRRSIHERVST
Sbjct: 6 MITSFSLQSFNIASTAHNSRNKQDLAKQNSVELVLGYRNPKGCNRFSFGSRRSIHERVST 185
Query: 61 GFKGVAVITDNKSAMSATEEDLENIGI 87
FKGVAVITDNKSAMSATEEDLENIGI
Sbjct: 186CFKGVAVITDNKSAMSATEEDLENIGI 266
>BP052685
Length = 419
Score = 157 bits (398), Expect = 5e-39
Identities = 72/118 (61%), Positives = 92/118 (77%)
Frame = +2
Query: 213 WIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINS 272
WIK++ P + PY+G+Y+DPP E+Y F++P+ +PK+ RIYE+H+GMSS EP+IN+
Sbjct: 71 WIKFSVQAPGEI--PYNGIYYDPPEEEKYVFRHPQTKRPKSLRIYESHIGMSSPEPKINT 244
Query: 273 YKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLI 330
Y F DD+LPRI+ YN VQ+MA+ EHSYYASFG HVTNFFA SSR GTPEDLK LI
Sbjct: 245 YANFRDDVLPRIKKLGYNAVQIMAIQEHSYYASFG*HVTNFFAPSSRLGTPEDLKSLI 418
>AV420391
Length = 205
Score = 83.6 bits (205), Expect = 1e-16
Identities = 37/66 (56%), Positives = 49/66 (74%)
Frame = +1
Query: 643 FDKAMNLLDDKFSFLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDL 702
FD+AM L+++F F+ S Q +S NE DKVIVFERG+LVFVFNFH +Y Y++GC
Sbjct: 7 FDRAMQHLEERFGFMTSEHQYISRKNEGDKVIVFERGNLVFVFNFHWNNSYYDYRIGCLH 186
Query: 703 PGKYRV 708
PGKY++
Sbjct: 187 PGKYKI 204
>BP080627
Length = 391
Score = 66.2 bits (160), Expect(2) = 4e-16
Identities = 26/53 (49%), Positives = 39/53 (73%)
Frame = -2
Query: 231 VYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPR 283
++W+PP + Y++KY P PK+ RIYEAHVG+S SEP+I+S+ +F D + R
Sbjct: 381 IHWEPPPEQAYKWKYTSPKAPKSLRIYEAHVGISGSEPKISSFSDFTDQVFFR 223
Score = 35.8 bits (81), Expect(2) = 4e-16
Identities = 14/32 (43%), Positives = 21/32 (64%)
Frame = -3
Query: 280 ILPRIRANNYNTVQLMAVMEHSYYASFGYHVT 311
+LP I+ YN +QL+ V+EH Y + GY V+
Sbjct: 116 VLPHIKEAGYNAIQLIGVVEHKDYFTVGYRVS 21
>TC14733 similar to UP|Q41058 (Q41058) Starch branching enzyme I precursor ,
partial (8%)
Length = 1188
Score = 63.5 bits (153), Expect = 1e-10
Identities = 34/99 (34%), Positives = 52/99 (52%), Gaps = 3/99 (3%)
Frame = +2
Query: 696 YKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILS 755
Y++GC PG Y++ALDSD FGG R+ H ++FT + +++RP SF + +
Sbjct: 5 YRIGCLHPGXYKIALDSDDPSFGGFNRLNHAAEYFTT--------DGWYDDRPRSFLVYA 160
Query: 756 PPRTCVVYYRVDESQEENSIS---NLVGVQETSTAADIV 791
P RT VVY D + E + + + ET+ D V
Sbjct: 161 PSRTAVVYALADGIELEPEVEP**EVCRINETNHGIDAV 277
>TC9139 similar to UP|Q9LNA8 (Q9LNA8) F5O11.10, partial (12%)
Length = 1084
Score = 28.9 bits (63), Expect = 3.6
Identities = 19/58 (32%), Positives = 30/58 (50%), Gaps = 13/58 (22%)
Frame = -3
Query: 792 ANIPDGSSAS-------KEREVSNFNWT------METLAAANADVAKIPDELVPAAEN 836
+N PD + AS K++++SN +T E + AA + KIP ++VPA N
Sbjct: 917 SNCPDITKASCXVNIPYKQKQISNATFTCQRVSVFERIIAAFTERDKIPSKIVPAENN 744
>TC14998 similar to UP|Q9ZU81 (Q9ZU81) At2g48080 protein, partial (6%)
Length = 586
Score = 28.5 bits (62), Expect = 4.7
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 5/57 (8%)
Frame = +2
Query: 219 VDPTKFAAPYDGVY--WDPPLSERYQFKYPRPPKPKA---PRIYEAHVGMSSSEPRI 270
++P K A GV+ W+ P + + PR K + P E H+G SSSEP I
Sbjct: 62 LNPRKLAGGGTGVFLPWNVPSRKPAKHLPPRAQKGRLLALPSPAEPHMGESSSEPSI 232
>TC19999 weakly similar to UP|AAQ89667 (AAQ89667) At5g25170, partial (37%)
Length = 563
Score = 28.1 bits (61), Expect = 6.1
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -1
Query: 478 PGLGRPISEVGIGFDYRLAMAIPDKWIDY 506
P L R ++++GIGF L A KW+ +
Sbjct: 191 PILARRLTQLGIGFSVSLTQAALQKWLQF 105
>TC16126
Length = 464
Score = 28.1 bits (61), Expect = 6.1
Identities = 13/30 (43%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Frame = -2
Query: 403 RWWLEEF-KFDGFRFDGVTSMLYHHHGVNI 431
RWWLEE+ + GFR V +L H G +
Sbjct: 160 RWWLEEWQRRKGFRVGSVKWVLEHERGKRV 71
>BI420337
Length = 397
Score = 27.7 bits (60), Expect = 7.9
Identities = 19/64 (29%), Positives = 24/64 (36%), Gaps = 5/64 (7%)
Frame = +1
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP-----KAPRIYEAHVGMSSSEPRI 270
Y + P + P Y PP + +KYP PP P P A+ S P
Sbjct: 130 YHSPPPPMHSPPPPYHYSSPPPPPKKPYKYPSPPPPVYKYKSPPPRTPAYYYKSPPPPPK 309
Query: 271 NSYK 274
N YK
Sbjct: 310 NPYK 321
>TC14425 similar to PIR|T50796|T50796 chorismate mutase CM2 - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (58%)
Length = 1144
Score = 27.7 bits (60), Expect = 7.9
Identities = 29/116 (25%), Positives = 48/116 (41%), Gaps = 3/116 (2%)
Frame = +2
Query: 230 GVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRA--N 287
G Y +P + + P P P P H G +S + +K + D++LP + A +
Sbjct: 338 GRYQNPEENAFFPEHLPSPIVPSYPFTQILHPGAASININKSIWKNYFDELLPILVASGD 517
Query: 288 NYNTVQLMAV-MEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLM 342
+ N Q A + S H F A + +P+D + LI + GL L+
Sbjct: 518 DGNYAQTAASDLSLLQAISRRIHYGKFVAEAKFRESPQDYEPLIRAKDTEGLMKLL 685
>AV425617
Length = 405
Score = 27.7 bits (60), Expect = 7.9
Identities = 19/49 (38%), Positives = 26/49 (52%), Gaps = 3/49 (6%)
Frame = -3
Query: 749 NSFKILSPPRTCVVYYRVDE---SQEENSISNLVGVQETSTAADIVANI 794
NSFK LS TC + + E SQ + +N +G+ ETS D +NI
Sbjct: 244 NSFKELS---TCSILHNYSEMGWSQNQLLEANNIGMTETSMVDDFPSNI 107
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 27.7 bits (60), Expect = 7.9
Identities = 21/66 (31%), Positives = 26/66 (38%), Gaps = 2/66 (3%)
Frame = +1
Query: 211 PAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAP--RIYEAHVGMSSSEP 268
P W K + P PY Y+ P Y +K P PP P P Y++ S S P
Sbjct: 4 PYWYK-SPPPPPPVPKPY---YYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPP 171
Query: 269 RINSYK 274
YK
Sbjct: 172 PPYYYK 189
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,984,199
Number of Sequences: 28460
Number of extensions: 222516
Number of successful extensions: 1041
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1014
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1038
length of query: 848
length of database: 4,897,600
effective HSP length: 98
effective length of query: 750
effective length of database: 2,108,520
effective search space: 1581390000
effective search space used: 1581390000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0193.7


