

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
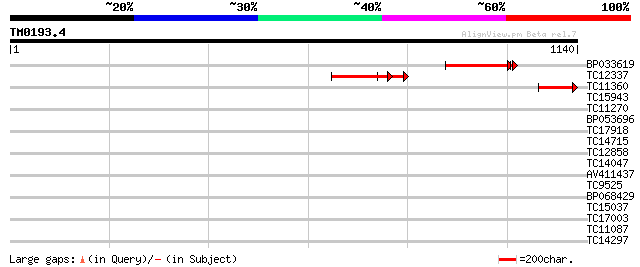
Query= TM0193.4
(1140 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP033619 285 6e-80
TC12337 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (11%) 238 5e-63
TC11360 homologue to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%) 153 1e-37
TC15943 similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific fungal... 32 0.57
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 32 0.57
BP053696 32 0.57
TC17918 weakly similar to UP|Q9FLU4 (Q9FLU4) Genomic DNA, chromo... 32 0.74
TC14715 32 0.74
TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter,... 31 0.97
TC14047 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {... 31 1.3
AV411437 30 1.7
TC9525 similar to UP|Q9AVS2 (Q9AVS2) Trypsin inhibitor precursor... 30 2.2
BP068429 29 3.7
TC15037 similar to UP|O24457 (O24457) Pyruvate dehydrogenase E1 ... 29 3.7
TC17003 homologue to UP|Q40329 (Q40329) Serine proteinase inhibi... 28 6.3
TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protei... 28 6.3
TC14297 similar to UP|Q9STA6 (Q9STA6) RAD23 protein, partial (81%) 28 8.2
>BP033619
Length = 432
Score = 285 bits (730), Expect(2) = 6e-80
Identities = 130/134 (97%), Positives = 132/134 (98%)
Frame = -2
Query: 877 SDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPC 936
SDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPC
Sbjct: 431 SDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPC 252
Query: 937 CAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYT 996
CAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYT
Sbjct: 251 CAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYT 72
Query: 997 SSVDLKGYDSGIIQ 1010
SSVDLKGYDS + +
Sbjct: 71 SSVDLKGYDSALFK 30
Score = 30.8 bits (68), Expect(2) = 6e-80
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = -1
Query: 1008 IIQASSFRTYHTP 1020
IIQASSFRTYHTP
Sbjct: 39 IIQASSFRTYHTP 1
>TC12337 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (11%)
Length = 676
Score = 238 bits (606), Expect = 5e-63
Identities = 122/123 (99%), Positives = 122/123 (99%)
Frame = +3
Query: 648 IPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSF 707
IPFLVLA GVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSF
Sbjct: 18 IPFLVLAGGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSF 197
Query: 708 ISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPD 767
ISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPD
Sbjct: 198 ISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPD 377
Query: 768 KGI 770
KGI
Sbjct: 378 KGI 386
Score = 70.9 bits (172), Expect = 1e-12
Identities = 40/68 (58%), Positives = 47/68 (68%), Gaps = 4/68 (5%)
Frame = +3
Query: 739 IVLDSQRAEDKRVDCFP----CIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWG 794
I++D + K ++ F + H+F GIRQRKPGLLARYMKEVHAPILSIWG
Sbjct: 486 ILVDQKSYLLKNIESFDDFC*SLSFHNF-----SGIRQRKPGLLARYMKEVHAPILSIWG 650
Query: 795 VKIVVIAI 802
VKIVVIAI
Sbjct: 651 VKIVVIAI 674
>TC11360 homologue to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%)
Length = 651
Score = 153 bits (387), Expect = 1e-37
Identities = 78/78 (100%), Positives = 78/78 (100%)
Frame = +1
Query: 1063 MGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
MGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP
Sbjct: 1 MGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 180
Query: 1123 PSRCTIIEQEENRSSTSS 1140
PSRCTIIEQEENRSSTSS
Sbjct: 181 PSRCTIIEQEENRSSTSS 234
>TC15943 similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific fungal
endoglucanase inhibitor protein precursor, partial (15%)
Length = 968
Score = 32.0 bits (71), Expect = 0.57
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 175 NATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNS 213
N SS +P++ + SCS+T GC+ D P C+N+
Sbjct: 284 NHYNSSTYRPVSCDSKSCSNTGAGCTSCDGPLKPGCTNN 400
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 32.0 bits (71), Expect = 0.57
Identities = 27/103 (26%), Positives = 47/103 (45%), Gaps = 9/103 (8%)
Frame = -3
Query: 154 FIGRKAAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTS-LGCS--------CGDC 204
F P+S S + F +++ ++ + S +S S +S CS CG C
Sbjct: 553 FTSTTTTPSSSSSSSSSSFPCSSSTTTSSSSSSSSPFSSSSSSSCACSTRDLPRVGCGVC 374
Query: 205 PSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFILAVLYIIL 247
SSS S+S+S+T + S S+ L + ++ +L +IL
Sbjct: 373 SSSSSSSSSSSSTPWFSFSTSLSFFFLPPPLIIILILILILIL 245
>BP053696
Length = 509
Score = 32.0 bits (71), Expect = 0.57
Identities = 15/36 (41%), Positives = 21/36 (57%)
Frame = -2
Query: 839 NVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQ 874
NV+ L IGP +YF+ + +SES H C +SQ
Sbjct: 109 NVTCCLHIGPTVYFLTEEAYVASESLHDTGTCLLSQ 2
>TC17918 weakly similar to UP|Q9FLU4 (Q9FLU4) Genomic DNA, chromosome 5, TAC
clone:K18P6, partial (35%)
Length = 583
Score = 31.6 bits (70), Expect = 0.74
Identities = 15/42 (35%), Positives = 24/42 (56%)
Frame = +2
Query: 220 KANSCSIKVGSLTVKCVDFILAVLYIILICVFLGWALYHRIR 261
+ S I S+ VK +DFIL L ++++ ++ W LY IR
Sbjct: 221 EVTSLKINTSSMEVKYLDFILVPLGVLVLGIYHVWLLYTIIR 346
>TC14715
Length = 637
Score = 31.6 bits (70), Expect = 0.74
Identities = 25/103 (24%), Positives = 38/103 (36%), Gaps = 7/103 (6%)
Frame = +1
Query: 867 NQLCSISQCNSDSLLNEISKAS--LVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTN 924
N CS++ D +++ ++ L TS +A + W+ P F +
Sbjct: 1 NLFCSVTCSICDGCISQCTEPMD*LWSTTSNLASSPTNTRTTMACWLQPAGFLWDLSWIW 180
Query: 925 GSYCPP-----DDQPPCCAPEDDSCVSGACKDCTTCFRHSDLR 962
G C P CC SC+S CT C +DLR
Sbjct: 181 GQLCKPAVAYVSPTKYCCIRSLSSCIS-----CTACSSSTDLR 294
>TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter, partial
(51%)
Length = 882
Score = 31.2 bits (69), Expect = 0.97
Identities = 15/45 (33%), Positives = 21/45 (46%)
Frame = +2
Query: 907 FLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKD 951
F W SP + CCR + PCC+ SC +GAC++
Sbjct: 326 FGAWCSPSRYFCCRLTLPS*LT----ELPCCSRSSRSC*TGACEN 448
>TC14047 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (67%)
Length = 842
Score = 30.8 bits (68), Expect = 1.3
Identities = 19/43 (44%), Positives = 23/43 (53%), Gaps = 2/43 (4%)
Frame = -2
Query: 45 CCTKAQFDTLQTQVQQ--AIPFLVGCPACLRNFLNLFCELTCS 85
CC++ +F LQ QQ A LVGCP CL F C+L S
Sbjct: 715 CCSRPRFCHLQAVYQQR*ASVGLVGCPLCL-EFWPSHCQLCLS 590
>AV411437
Length = 417
Score = 30.4 bits (67), Expect = 1.7
Identities = 22/64 (34%), Positives = 29/64 (44%)
Frame = -1
Query: 174 PNATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTV 233
P SS P +VS S S++ SSS CSNS ST + ++S S S V
Sbjct: 201 PPTWASSSSPPPSVSNSSSSES--------VSSSSNCSNSPSTPSSSSSSSSSSSSSAIV 46
Query: 234 KCVD 237
C +
Sbjct: 45 SCAE 34
>TC9525 similar to UP|Q9AVS2 (Q9AVS2) Trypsin inhibitor precursor, partial
(85%)
Length = 547
Score = 30.0 bits (66), Expect = 2.2
Identities = 18/62 (29%), Positives = 27/62 (43%), Gaps = 1/62 (1%)
Frame = +3
Query: 892 ETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPC-CAPEDDSCVSGACK 950
+ S+I + DD ++ A CC + YC P C C+ ++C S ACK
Sbjct: 144 QNSFITQVLPKGTDDTNYYVKSTATACC----DNCYCTKSIPPQCHCSDIGETCHS-ACK 308
Query: 951 DC 952
C
Sbjct: 309 TC 314
>BP068429
Length = 412
Score = 29.3 bits (64), Expect = 3.7
Identities = 20/69 (28%), Positives = 35/69 (49%), Gaps = 1/69 (1%)
Frame = -3
Query: 777 LLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEP-GLEQEIVLPRDSYLQG 835
LL M+ + P S W + +++ ++ V +ASI + + P L +L D YL G
Sbjct: 344 LLILQMRALFHPPASFWRMSLLIPSLLVLGLMASIGVRIQNSPKALLMSNLLHSD*YLLG 165
Query: 836 YFNNVSEYL 844
F+ V ++L
Sbjct: 164 AFSFVWKFL 138
>TC15037 similar to UP|O24457 (O24457) Pyruvate dehydrogenase E1 alpha
subunit (At1g01090/T25K16_8) , partial (20%)
Length = 499
Score = 29.3 bits (64), Expect = 3.7
Identities = 20/69 (28%), Positives = 34/69 (48%), Gaps = 1/69 (1%)
Frame = +3
Query: 777 LLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEP-GLEQEIVLPRDSYLQG 835
LL M+ + P S W + +++ + V +ASI + + P L +L D YL G
Sbjct: 111 LLNLQMRALFHPAASFWRMSLLIPKVLVLGLMASIGVRIQNSPKALLMSNLLHSD*YLLG 290
Query: 836 YFNNVSEYL 844
F+ V ++L
Sbjct: 291 AFSFVWKFL 317
>TC17003 homologue to UP|Q40329 (Q40329) Serine proteinase inhibitor,
partial (53%)
Length = 523
Score = 28.5 bits (62), Expect = 6.3
Identities = 20/70 (28%), Positives = 28/70 (39%), Gaps = 1/70 (1%)
Frame = +1
Query: 884 ISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPC-CAPEDD 942
++ A S+IA+ DD ++ A CC N C P C C +
Sbjct: 79 VADARSFDPNSFIAQVLPKGTDDTSYYVKSTATACC----NSCPCTRSIPPQCRCTDIGE 246
Query: 943 SCVSGACKDC 952
+C S ACK C
Sbjct: 247 TCHS-ACKAC 273
>TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protein
(OJ1118_G09.13 protein), partial (6%)
Length = 689
Score = 28.5 bits (62), Expect = 6.3
Identities = 24/72 (33%), Positives = 34/72 (46%)
Frame = +1
Query: 162 NSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKA 221
++PGSP +P AT ++ +VS S + T PSS NSASTT +
Sbjct: 121 STPGSP-----KPPATATTNSPSSSVSTSSGAPTPKPSRTTPSPSS----NSASTTAASS 273
Query: 222 NSCSIKVGSLTV 233
+ S SLT+
Sbjct: 274 SKSSTHPPSLTL 309
>TC14297 similar to UP|Q9STA6 (Q9STA6) RAD23 protein, partial (81%)
Length = 1472
Score = 28.1 bits (61), Expect = 8.2
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 199 CSCGDCPSSSVCSNSASTTINKANSCSIK 227
C C C SSS CS+++ TT SC +K
Sbjct: 667 CKCTTCKSSSSCSSASGTTC----SCYLK 741
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,359,373
Number of Sequences: 28460
Number of extensions: 308266
Number of successful extensions: 1680
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1654
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1677
length of query: 1140
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1040
effective length of database: 2,051,600
effective search space: 2133664000
effective search space used: 2133664000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0193.4


