

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
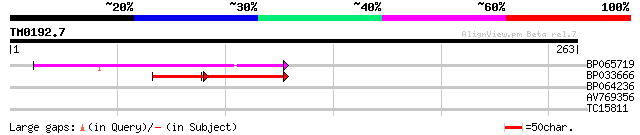
Query= TM0192.7
(263 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065719 52 1e-07
BP033666 37 2e-07
BP064236 30 0.42
AV769356 28 1.2
TC15811 similar to UP|RL13_ARATH (P41127) 60S ribosomal protein ... 27 2.7
>BP065719
Length = 567
Score = 51.6 bits (122), Expect = 1e-07
Identities = 32/120 (26%), Positives = 58/120 (47%), Gaps = 2/120 (1%)
Frame = +3
Query: 12 FSEDVESVAILDNMKALVLDSYSGDSDPK--DHLLYFNTKMVIIAASDAVKCRMFPSTFK 69
F E+V + K +SGDS +H+ + + +A ++ +K + FPS+
Sbjct: 135 FIEEVLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLT 314
Query: 70 STAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARY 129
A WFTTL S+ + F QF + + V+ DL S++++ ES+ +Y+ R+
Sbjct: 315 KNAFTWFTTLAPRSVHTWAQLERIFHEQFFRGECK-VSXKDLASVKRKPAESIDDYLNRF 491
>BP033666
Length = 342
Score = 36.6 bits (83), Expect(2) = 2e-07
Identities = 18/40 (45%), Positives = 25/40 (62%)
Frame = -1
Query: 90 FSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARY 129
F + FL FSAN+ Q T DL++I QQ GE ++ Y + Y
Sbjct: 273 FFN*FLTXFSANQTQKATSADLFNICQQVGERVRPYFSLY 154
Score = 33.9 bits (76), Expect(2) = 2e-07
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = -2
Query: 67 TFKSTAMAWFTTLPRGSISNFRDFSS 92
+FK AM WF P SISNF DFS+
Sbjct: 341 SFKGVAMTWFIKQPPYSISNFTDFST 264
>BP064236
Length = 578
Score = 30.0 bits (66), Expect = 0.42
Identities = 26/91 (28%), Positives = 41/91 (44%), Gaps = 8/91 (8%)
Frame = -3
Query: 13 SEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTA 72
S+ + S AI+D +KAL S P D ++ TKMV IA + + R+ S+
Sbjct: 513 SQKLNSEAIIDFVKALCKVSMEELRSPSDPRVFSLTKMVEIAHYNMNRIRLVWSSIWHVL 334
Query: 73 MAWFTTLPRG--------SISNFRDFSSKFL 95
+F T+ ++ + R S KFL
Sbjct: 333 SDFFVTIGCSANLSIAIFAMDSLRQLSMKFL 241
>AV769356
Length = 574
Score = 28.5 bits (62), Expect = 1.2
Identities = 19/59 (32%), Positives = 26/59 (43%), Gaps = 1/59 (1%)
Frame = -2
Query: 39 PKDHLLYFNTKMV-IIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLV 96
PK + F T ++ + A + P TFK + W T L S SN RD K L+
Sbjct: 546 PKGDVYSFGTVLLEFVTGERATQVAKAPETFKGNLVEWITQLT--SHSNLRDAIDKSLL 376
>TC15811 similar to UP|RL13_ARATH (P41127) 60S ribosomal protein L13 (BBC1
protein homolog), complete
Length = 981
Score = 27.3 bits (59), Expect = 2.7
Identities = 17/46 (36%), Positives = 27/46 (57%), Gaps = 3/46 (6%)
Frame = +2
Query: 163 RKPARSMEEMRA---RASTYILDEEDDAFKRKRAKLEKGDTSPEQR 205
R+ RS+E ++A R TY + F R+ K++ GD+SPE+R
Sbjct: 332 RRRNRSLESLQANVQRLKTY--KAKLVVFPRRARKVKAGDSSPEER 463
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.134 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,968,661
Number of Sequences: 28460
Number of extensions: 30027
Number of successful extensions: 153
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 153
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 153
length of query: 263
length of database: 4,897,600
effective HSP length: 88
effective length of query: 175
effective length of database: 2,393,120
effective search space: 418796000
effective search space used: 418796000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0192.7


