

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0191.15
(398 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
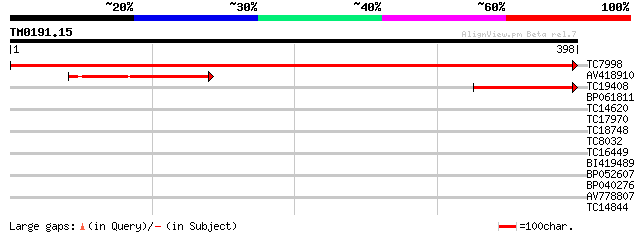
Sequences producing significant alignments: (bits) Value
TC7998 homologue to UP|HEM6_SOYBN (P35055) Coproporphyrinogen II... 830 0.0
AV418910 164 3e-41
TC19408 homologue to PIR|S39523|S39523 coproporphyrinogen oxidas... 157 3e-39
BP061811 37 0.004
TC14620 31 0.32
TC17970 27 4.6
TC18748 similar to GB|AAN28890.1|23506061|AY143951 At4g02590/T10... 27 4.6
TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-... 27 6.0
TC16449 similar to UP|Q940R4 (Q940R4) AT4g16560/dl4305c, partial... 27 6.0
BI419489 27 6.0
BP052607 27 6.0
BP040276 27 7.8
AV778807 27 7.8
TC14844 similar to UP|Q8L553 (Q8L553) SCARECROW transcriptional ... 27 7.8
>TC7998 homologue to UP|HEM6_SOYBN (P35055) Coproporphyrinogen III oxidase,
chloroplast precursor (Coproporphyrinogenase) (Coprogen
oxidase) , partial (88%)
Length = 1573
Score = 830 bits (2144), Expect = 0.0
Identities = 398/398 (100%), Positives = 398/398 (100%)
Frame = +2
Query: 1 MPSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVS 60
MPSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVS
Sbjct: 62 MPSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVS 241
Query: 61 IEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKE 120
IEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKE
Sbjct: 242 IEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKE 421
Query: 121 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPF 180
DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPF
Sbjct: 422 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPF 601
Query: 181 FAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVK 240
FAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVK
Sbjct: 602 FAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVK 781
Query: 241 HFHSIQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA 300
HFHSIQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA
Sbjct: 782 HFHSIQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA 961
Query: 301 TECANSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE 360
TECANSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE
Sbjct: 962 TECANSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE 1141
Query: 361 SILVSLPLTSRWEYDHKPEEGSEEWKLLDACINPKEWI 398
SILVSLPLTSRWEYDHKPEEGSEEWKLLDACINPKEWI
Sbjct: 1142SILVSLPLTSRWEYDHKPEEGSEEWKLLDACINPKEWI 1255
>AV418910
Length = 298
Score = 164 bits (414), Expect = 3e-41
Identities = 86/102 (84%), Positives = 88/102 (85%)
Frame = +2
Query: 42 QTKTMTSLGSKGAVRAGVSIEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREA 101
+TKTM S K VRA VSIEKETPEAERPETFLRGGD A A + S SVRARFEKMIREA
Sbjct: 2 ETKTMRS--KKSVVRATVSIEKETPEAERPETFLRGGDAATA-AESLSVRARFEKMIREA 172
Query: 102 QDSVCDAIEAADGGAKFKEDVWSRPGGGGGISRVLQDGAVWE 143
QDSVCDAIEAADGG KFKEDVWSRPGGGGGISRVLQDG VWE
Sbjct: 173 QDSVCDAIEAADGGGKFKEDVWSRPGGGGGISRVLQDGVVWE 298
>TC19408 homologue to PIR|S39523|S39523 coproporphyrinogen oxidase
precursor - soybean {Glycine max;} , partial (19%)
Length = 593
Score = 157 bits (397), Expect = 3e-39
Identities = 71/73 (97%), Positives = 73/73 (99%)
Frame = +2
Query: 326 HQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILVSLPLTSRWEYDHKPEEGSEEW 385
HQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILVSLPLT+RWEYDHKPEEGSEEW
Sbjct: 2 HQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILVSLPLTARWEYDHKPEEGSEEW 181
Query: 386 KLLDACINPKEWI 398
KLLDAC+NPKEWI
Sbjct: 182 KLLDACMNPKEWI 220
>BP061811
Length = 492
Score = 37.4 bits (85), Expect = 0.004
Identities = 15/17 (88%), Positives = 15/17 (88%)
Frame = -3
Query: 320 DTPFTDHQKAWQQLRRG 336
DTPFTDHQKAWQ LR G
Sbjct: 490 DTPFTDHQKAWQLLRMG 440
>TC14620
Length = 1260
Score = 31.2 bits (69), Expect = 0.32
Identities = 22/82 (26%), Positives = 35/82 (41%)
Frame = -2
Query: 79 DTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKEDVWSRPGGGGGISRVLQD 138
+ A + ++ SV AR + R + + DA+ G A GGGG+ + +
Sbjct: 455 EEAHSAAAKGSVLARLRPVTRRVEPAFEDAVGEVAGVASVAVSFAEVRAGGGGVVVLARV 276
Query: 139 GAVWEKAGVNVSVVYGVMPPDA 160
A AGV V + V+ P A
Sbjct: 275 SATGFLAGVGVGAICPVVWPAA 210
>TC17970
Length = 542
Score = 27.3 bits (59), Expect = 4.6
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +3
Query: 1 MPSCATIVSPPSYAIPFFTSTSSTSSPTAKLLS 33
+P + PP A PF ST+S SSPT L+
Sbjct: 255 VPELTSPSLPPPTASPFKPSTASRSSPTLPFLT 353
>TC18748 similar to GB|AAN28890.1|23506061|AY143951 At4g02590/T10P11_13
{Arabidopsis thaliana;}, partial (36%)
Length = 803
Score = 27.3 bits (59), Expect = 4.6
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = -3
Query: 120 EDVWSRPGGGGGISRVLQDGAVWEK 144
ED+ GGGG+ RV++ +VW +
Sbjct: 153 EDLLEEVVGGGGVGRVIRHDSVWTR 79
>TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-like
protein, partial (97%)
Length = 2230
Score = 26.9 bits (58), Expect = 6.0
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = +1
Query: 9 SPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLI 41
+P IP FT+TS SSP+ K + + NPL+
Sbjct: 694 TPWRILIPTFTATSPPSSPSWKPANLQTRNPLL 792
>TC16449 similar to UP|Q940R4 (Q940R4) AT4g16560/dl4305c, partial (18%)
Length = 613
Score = 26.9 bits (58), Expect = 6.0
Identities = 25/70 (35%), Positives = 29/70 (40%), Gaps = 11/70 (15%)
Frame = -2
Query: 117 KFKEDVWSRPGGGGGISRVLQDGAVWEKAG--------VNVSVVYGVMPPD---AYRAAK 165
K E V GGGGG+ VL +G W G +SV+ GV P D RA
Sbjct: 276 KVAERVGVVGGGGGGVGSVLLNGLRWF*GGGGYC*IGIWKLSVLRGVGPFDMSVKCRA*S 97
Query: 166 GAAASPDEKP 175
A EKP
Sbjct: 96 RAETGEKEKP 67
>BI419489
Length = 479
Score = 26.9 bits (58), Expect = 6.0
Identities = 21/88 (23%), Positives = 37/88 (41%)
Frame = +2
Query: 2 PSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSI 61
P+ SPP A P +S+++PTAK R T S+ + + ++ S
Sbjct: 107 PTAYLTASPPPTATPAAL*KTSSTTPTAKCTVPSRTTTAATAPTGASVAAAASGQSSSSS 286
Query: 62 EKETPEAERPETFLRGGDTAEALSSSSS 89
+P P R ++ AL++ +S
Sbjct: 287 SPRSP----PPLSARHSTSSTALTARNS 358
>BP052607
Length = 466
Score = 26.9 bits (58), Expect = 6.0
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -1
Query: 179 PFFAAGISSVLHPKNPFAP 197
P FAAG SSV+ P+ P AP
Sbjct: 334 PEFAAGSSSVVRPQGPGAP 278
>BP040276
Length = 556
Score = 26.6 bits (57), Expect = 7.8
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +1
Query: 163 AAKGAAASPDEKPGPVPFFAAGISSVLHPKNP 194
A GAAASP+ P P P ++ +S+ P +P
Sbjct: 112 AMSGAAASPNHSPSPSP-WSQIVSAAAPPSSP 204
>AV778807
Length = 590
Score = 26.6 bits (57), Expect = 7.8
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +2
Query: 13 YAIPFFTSTSSTSSPTAKLLSKRRWN 38
Y P+F + +ST PT+K+ + WN
Sbjct: 161 YLSPWFAAINSTQLPTSKVHRSKFWN 238
>TC14844 similar to UP|Q8L553 (Q8L553) SCARECROW transcriptional
regulator-like, partial (49%)
Length = 1051
Score = 26.6 bits (57), Expect = 7.8
Identities = 16/50 (32%), Positives = 23/50 (46%)
Frame = -3
Query: 11 PSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVS 60
P +AIP FTS PT LL L + + + + + VRA +S
Sbjct: 536 PLHAIPPFTSHRVPHLPTQSLLDLHTLQALTRHRGLNRVKQRAVVRARLS 387
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,762,133
Number of Sequences: 28460
Number of extensions: 114944
Number of successful extensions: 714
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 706
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 713
length of query: 398
length of database: 4,897,600
effective HSP length: 92
effective length of query: 306
effective length of database: 2,279,280
effective search space: 697459680
effective search space used: 697459680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0191.15


