

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
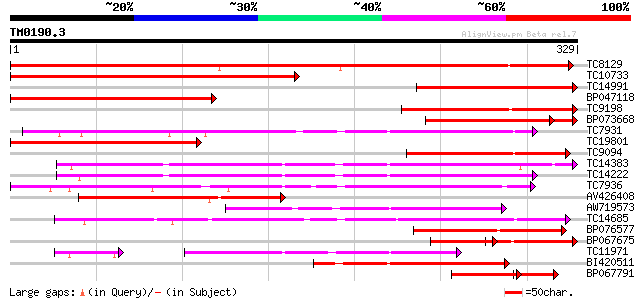
Score E
Sequences producing significant alignments: (bits) Value
TC8129 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (90%) 409 e-115
TC10733 similar to UP|Q9SQ81 (Q9SQ81) Gibberellin 2 beta-hydroxy... 343 3e-95
TC14991 similar to UP|O04162 (O04162) Dioxygenase, partial (28%) 187 2e-48
BP047118 181 2e-46
TC9198 similar to UP|Q9SQ80 (Q9SQ80) Gibberellin 2 beta-hydroxyl... 171 2e-43
BP073668 137 7e-37
TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial... 145 9e-36
TC19801 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (34%) 141 1e-34
TC9094 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (28%) 137 2e-33
TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-ca... 134 2e-32
TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carb... 132 8e-32
TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, pa... 124 2e-29
AV426408 118 1e-27
AW719573 116 6e-27
TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 102 9e-23
BP076577 93 5e-20
BP067675 60 6e-20
TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partia... 82 2e-19
BI420511 91 3e-19
BP067791 68 1e-17
>TC8129 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (90%)
Length = 1278
Score = 409 bits (1052), Expect = e-115
Identities = 207/335 (61%), Positives = 257/335 (75%), Gaps = 8/335 (2%)
Frame = +2
Query: 1 MVLLSKQ-ATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVS 59
MV+ S+Q A + +K C T +P VDLS P+AK++IV AC+EFGFFK+INH +
Sbjct: 116 MVVHSQQGALNELFLIKTCKPTNLFTEVPEVDLSHPEAKTLIVNACKEFGFFKIINHDIP 295
Query: 60 MEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNV 119
ME IS LE+EAL+FF P +EKEK L +PFGYG+KRIG NGDVGWVEYLL TN +V
Sbjct: 296 MELISNLENEALSFFMQPQSEKEKASLDDPFGYGSKRIGKNGDVGWVEYLLFNTNPDVIS 475
Query: 120 P----VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV 175
P + QN F ++DY AV+N+ CEIL+LMA+GL I+ +NV S+L+ D++SD +
Sbjct: 476 PNPLLLLEQNQNVFSSAVNDYTLAVKNLSCEILELMADGLGIEQRNVFSRLIRDEKSDCI 655
Query: 176 FRVNHYPACPEMGLD---GENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPD 232
FR+NHYPAC E+ L G NLIGFGEHTDPQ+IS+LRSNNTSGLQI LRDG+W+S+PPD
Sbjct: 656 FRLNHYPACGELELQAIGGRNLIGFGEHTDPQMISVLRSNNTSGLQICLRDGTWVSIPPD 835
Query: 233 YSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGG 292
+SFFINVGDSLQVMTNG FRSVKHRVL++ SR+SMIYFGG PL+EK+APLPSL+
Sbjct: 836 QTSFFINVGDSLQVMTNGSFRSVKHRVLSDTAMSRVSMIYFGGAPLTEKLAPLPSLV-SK 1012
Query: 293 KEESLYKEFTWFEYKNSSYGSRLADNRLGHFERIA 327
+E+SLYKEFTW EYK ++Y RLADNRL FE+ A
Sbjct: 1013EEDSLYKEFTWGEYKKAAYKLRLADNRLTPFEKSA 1117
>TC10733 similar to UP|Q9SQ81 (Q9SQ81) Gibberellin 2 beta-hydroxylase,
partial (52%)
Length = 616
Score = 343 bits (879), Expect = 3e-95
Identities = 168/168 (100%), Positives = 168/168 (100%)
Frame = +1
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM
Sbjct: 112 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 291
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP
Sbjct: 292 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 471
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLM 168
VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLM
Sbjct: 472 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLM 615
>TC14991 similar to UP|O04162 (O04162) Dioxygenase, partial (28%)
Length = 507
Score = 187 bits (476), Expect = 2e-48
Identities = 91/93 (97%), Positives = 93/93 (99%)
Frame = +2
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSL+KGGKEES
Sbjct: 2 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLMKGGKEES 181
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
LYKEFTWFEY+NSSYGSRLADNRLGHFERIAAS
Sbjct: 182 LYKEFTWFEYQNSSYGSRLADNRLGHFERIAAS 280
>BP047118
Length = 476
Score = 181 bits (458), Expect = 2e-46
Identities = 89/120 (74%), Positives = 95/120 (79%)
Frame = -2
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LSK TEQYSY+ NC T S TIPVVDLSKPDAK++IVKACEEFGFFKVINHGV M
Sbjct: 376 MVVLSKPTTEQYSYILNCKPTTFSNTIPVVDLSKPDAKTLIVKACEEFGFFKVINHGVPM 197
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E IS LESEA FFSMP+ +KEK G NPFGYG K IG NGDVGWVEYLLL NQE P
Sbjct: 196 ETISMLESEASKFFSMPLNQKEKAGPPNPFGYGTKNIGQNGDVGWVEYLLLTNNQEYIPP 17
>TC9198 similar to UP|Q9SQ80 (Q9SQ80) Gibberellin 2 beta-hydroxylase,
partial (30%)
Length = 555
Score = 171 bits (432), Expect = 2e-43
Identities = 84/102 (82%), Positives = 92/102 (89%)
Frame = +3
Query: 228 SVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPS 287
SVP D+ SFFI VGDSLQVMTNGRFRSV+HRVLANGF+SRLSMIYFGGPPLS+KIAPLP
Sbjct: 3 SVPSDHDSFFIYVGDSLQVMTNGRFRSVRHRVLANGFKSRLSMIYFGGPPLSQKIAPLPC 182
Query: 288 LLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
L+K G E SLYKEFTW+EYK S+Y SRLADNRLG+FERI AS
Sbjct: 183 LMK-GNESSLYKEFTWYEYKKSAYASRLADNRLGNFERITAS 305
>BP073668
Length = 487
Score = 137 bits (345), Expect(2) = 7e-37
Identities = 66/75 (88%), Positives = 71/75 (94%)
Frame = -3
Query: 242 DSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEF 301
DSLQVMTNGRFRSV R+LANGF SRLSMIYFGGPPLSEKIAPLPSL+KGGKEESLYKEF
Sbjct: 485 DSLQVMTNGRFRSVNIRLLANGFPSRLSMIYFGGPPLSEKIAPLPSLMKGGKEESLYKEF 306
Query: 302 TWFEYKNSSYGSRLA 316
TWFE KNSSYGS+++
Sbjct: 305 TWFEXKNSSYGSQIS 261
Score = 33.1 bits (74), Expect(2) = 7e-37
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = -1
Query: 314 RLADNRLGHFERIAAS 329
RLAD RLGHFERIAAS
Sbjct: 268 RLADTRLGHFERIAAS 221
>TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial (29%)
Length = 1477
Score = 145 bits (366), Expect = 9e-36
Identities = 100/315 (31%), Positives = 161/315 (50%), Gaps = 16/315 (5%)
Frame = +1
Query: 8 ATEQYSYVKNCMATPCSFTI------PVVDLSKPDAKSI---IVKACEEFGFFKVINHGV 58
+T SYV+ + P + T+ PVVDL D I KA EE+GFF+VINHGV
Sbjct: 58 STVPLSYVQPPESRPGTVTVASGKAVPVVDLGGHDRAETLKQIFKASEEYGFFQVINHGV 237
Query: 59 SMEAISQLESEALNFFSMPVTEKEKTGLANPFG----YGNKRIGHNGDVGWVEYLLLK-- 112
S E I + F +MP EK +P G Y ++ I + + + + L
Sbjct: 238 SNELIEDTLNIFKEFHAMPAEEKLSESSRDPNGSCMLYTSREINNKDCIQFWKDTLRHPC 417
Query: 113 TNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQS 172
T E + P ++R V++ Y +R +G +IL+++ EGL + P+ + L
Sbjct: 418 TPSEEFMEFWPLKPARYREVVEKYTKELRELGSQILEMLCEGLGLDPRYCCAGL----SE 585
Query: 173 DSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPP 231
+ + NHYP CPE L +G +H DP +++ LL+ + + LQ+ +DG WIS+ P
Sbjct: 586 NPLLLSNHYPPCPEPSLT----LGLSKHRDPTLVTILLQDTDVNALQV-FKDGEWISIEP 750
Query: 232 DYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKG 291
+F +N+G LQ+++NGR V+HRV+ N SR ++ YF P E + P +L++
Sbjct: 751 IPYAFVVNIGLLLQIISNGRLVGVEHRVVTNSSISRTTVAYFIRPKKEEIVEPAKALVRT 930
Query: 292 GKEESLYKEFTWFEY 306
G +Y+ + E+
Sbjct: 931 G-AHPIYRSIAFEEF 972
>TC19801 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (34%)
Length = 508
Score = 141 bits (356), Expect = 1e-34
Identities = 67/111 (60%), Positives = 86/111 (77%)
Frame = +2
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LS+ Q+ +VK C T IPVVDLS P+AKS+IV+AC+EFGFFK+INHGV +
Sbjct: 173 MVVLSQPPLNQFFHVKPCKPTTFFTGIPVVDLSDPEAKSLIVQACKEFGFFKLINHGVPL 352
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLL 111
+ +S LE+EA+ FF P TEK++ G +PFGYG+K+IG NGDVGWVEYLLL
Sbjct: 353 DLMSNLENEAVRFFRKPQTEKDRAGPPDPFGYGSKKIGSNGDVGWVEYLLL 505
>TC9094 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (28%)
Length = 677
Score = 137 bits (345), Expect = 2e-33
Identities = 68/95 (71%), Positives = 81/95 (84%)
Frame = +1
Query: 231 PDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLK 290
PD++SFFI VGD+LQVMTNGRF+SVKHRV+A+ +SRLSMIYFGG LSEKIAPL SL+
Sbjct: 1 PDHTSFFIYVGDTLQVMTNGRFKSVKHRVMADTTKSRLSMIYFGGASLSEKIAPLESLMS 180
Query: 291 GGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFER 325
G E+SLY+EFTW EYK ++Y SRL DNRLG FE+
Sbjct: 181 KG-EQSLYQEFTWSEYKKAAYSSRLGDNRLGPFEK 282
>TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1249
Score = 134 bits (337), Expect = 2e-32
Identities = 96/313 (30%), Positives = 159/313 (50%), Gaps = 11/313 (3%)
Frame = +3
Query: 28 PVVDLSK------PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
PV+ L K D K I ACE +GFF+++NHG+ + + +E + + + ++
Sbjct: 51 PVISLEKLNGEERKDIKLQIKDACENWGFFELVNHGIPHDVMDTVERLTKEHYRICMEQR 230
Query: 82 EKTGLANPFGYGNKRIGHN-GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAV 140
K +A+ G + + D+ W L+ E N+ +++R V+ D+ +
Sbjct: 231 FKELMASK---GLEAVQTEVKDMDWESTFHLRHLPESNISEIPDLTDEYRKVMKDFALRI 401
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGE 199
+ ++LDL+ E L ++ K L K + + +V +YP CP+ L + G
Sbjct: 402 EKLSEDLLDLLCENLGLE-KGYLKKAFHGSRGPTFGTKVANYPPCPKPNL----VKGLRA 566
Query: 200 HTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHR 258
HTD II L + + SGLQ+ L+DG W+ VPP S +N+GD L+V+TNG+++SV+HR
Sbjct: 567 HTDAGGIILLFQDDKVSGLQL-LKDGEWVDVPPMRHSIVVNLGDQLEVITNGKYKSVEHR 743
Query: 259 VLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEE--SLYKEFTWFEYKNSSYGSRLA 316
V+A +R+S+ F P I P P LL+ EE LY +F + +Y G +
Sbjct: 744 VIAQTDGTRMSIASFYNPGSDAVIYPAPELLEKEAEEKNQLYPKFVFEDYMKLYAGLKF- 920
Query: 317 DNRLGHFERIAAS 329
D + FE + AS
Sbjct: 921 DAKEPRFEALKAS 959
>TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carboxylate
oxidase homolog (ACC oxidase) , partial (97%)
Length = 1597
Score = 132 bits (332), Expect = 8e-32
Identities = 88/288 (30%), Positives = 149/288 (51%), Gaps = 9/288 (3%)
Frame = +1
Query: 28 PVVDLSKPDAKS------IIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
PVVD+SK + + +I ACE +GFF+++NH +S E + +E + + ++
Sbjct: 58 PVVDMSKLNTEERVATMELIKDACENWGFFELVNHEISTEFMDTVERLTKEHYKRFMEQR 237
Query: 82 EKTGLANPFGYGNKRIGHN-GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAV 140
K +A G + + D+ W L+ N+ E +R V+ ++ +
Sbjct: 238 FKEMVATK---GLETVQSEIDDLDWESTFFLRHLPSSNISEVPDLDEDYRKVMKEFAVKL 408
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGE 199
+ +LDL+ E L ++ K L K+ + + +V++YP CP+ L + G
Sbjct: 409 EKLAENLLDLLCENLGLE-KGYLKKVFYGSKGPNFGTKVSNYPPCPKPDL----IKGLRA 573
Query: 200 HTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHR 258
HTD II L + + SGLQ+ L+D WI VPP S +N+GD L+V+TNG+++SV HR
Sbjct: 574 HTDAGGIILLFQDDKVSGLQL-LKDDQWIDVPPMRHSIVVNLGDQLEVITNGKYKSVMHR 750
Query: 259 VLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
V+A +R+S+ F P I+P PSL+K + +Y +F + +Y
Sbjct: 751 VIAQTDGARMSLASFYNPGDDAVISPAPSLVKEDETSQVYPKFVFEDY 894
>TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(94%)
Length = 1492
Score = 124 bits (312), Expect = 2e-29
Identities = 90/330 (27%), Positives = 165/330 (49%), Gaps = 25/330 (7%)
Frame = +3
Query: 1 MVLLSKQATEQYSYVKNCMATP------CSFTIPVVDLS--------KPDAKSIIVKACE 46
+ L++Q T + S+V++ P S IPV+ L+ + + + IV+ACE
Sbjct: 165 LTTLAQQNTLESSFVRDEDERPKVAYNNFSNEIPVISLAGIDEVDDRRSEICNKIVEACE 344
Query: 47 EFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK---EKTGLANPFGYGNKRIGHNGDV 103
+G F+V++HGV E +S + + A FF++P EK + TG + +
Sbjct: 345 NWGIFQVVDHGVDTELVSHMTTLAKEFFALPPEEKLRFDMTGGKKGGFIVSSHLQGESVQ 524
Query: 104 GWVEYLLLKTNQEVNVPVHSQN-------PEKFRCVLDDYLCAVRNMGCEILDLMAEGLK 156
W E + + P+ +++ P ++ V ++Y + + C++L++++E +
Sbjct: 525 DWREIVTY-----FSYPIRNRDYSRWPDTPAGWKAVTEEYSEKLMGLACKLLEVLSEAMG 689
Query: 157 IQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSG 216
++ K L+K +D V VN+YP CP+ L +G HTDP I+LL + G
Sbjct: 690 LE-KEALTKACVDMDQKVV--VNYYPKCPQPDLT----LGLKRHTDPGTITLLLQDQVGG 848
Query: 217 LQISLRDG-SWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGG 275
LQ + +G +WI+V P +F +N+GD ++NGRF++ H+ + N SRLS+ F
Sbjct: 849 LQATRDNGKTWITVQPVEGAFVVNLGDHGHYLSNGRFKNADHQAVVNSNSSRLSIATFQN 1028
Query: 276 PPLSEKIAPLPSLLKGGKEESLYKEFTWFE 305
P + PL ++ G++ + + T+ E
Sbjct: 1029PAPDATVYPLK--VREGEKSVMEEPITFAE 1112
>AV426408
Length = 385
Score = 118 bits (296), Expect = 1e-27
Identities = 58/124 (46%), Positives = 82/124 (65%), Gaps = 4/124 (3%)
Frame = +3
Query: 41 IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHN 100
+V+ACEE+GFFKV+NH V E +S+LE + FFS +EK + G A PFGYG + IG N
Sbjct: 9 VVRACEEYGFFKVVNHNVPREVVSRLEEKGTEFFSKSTSEKTQAGPATPFGYGCRNIGLN 188
Query: 101 GDVGWVEYLLLKTN----QEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLK 156
GD+G +EYLLL TN E + + + +P KF C + DY+ + + C+IL+L+AEGL
Sbjct: 189 GDMGDLEYLLLHTNPLSFTETSKTI-ANDPTKFSCAVSDYIEEAKELACKILELVAEGLW 365
Query: 157 IQPK 160
+ K
Sbjct: 366 VPDK 377
>AW719573
Length = 521
Score = 116 bits (290), Expect = 6e-27
Identities = 62/164 (37%), Positives = 96/164 (57%), Gaps = 1/164 (0%)
Frame = +2
Query: 126 PEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP 185
P+ FR L+ Y + N+ +I++LMA LKI PK + + + R+N+YP CP
Sbjct: 53 PQPFRENLEIYCAELNNLSIKIMELMANALKIDPKEITE---IYNEGSQTIRMNYYPPCP 223
Query: 186 EMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSL 244
+ E +IG HTD ++ LL++N+ GLQI +DG W+ V P ++F IN+GD L
Sbjct: 224 QP----ERVIGLKSHTDGAGLTILLQANDIEGLQIR-KDGHWVPVQPLPNAFIINIGDML 388
Query: 245 QVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSL 288
++ TNG + S++HR N + R+S+ F GP + +AP PSL
Sbjct: 389 EITTNGIYASIEHRATVNSEKERISLATFYGPNMQATLAPAPSL 520
>TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (38%)
Length = 1323
Score = 102 bits (254), Expect = 9e-23
Identities = 84/311 (27%), Positives = 142/311 (45%), Gaps = 12/311 (3%)
Frame = +1
Query: 27 IPVVDLSKPDAKSIIV-----KACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
IP +DL+ + V +A GFF+V+NHGV ++ + + + F P E+
Sbjct: 298 IPTIDLAAVNESRAAVVDQIRRAASSVGFFQVVNHGVPLDLLRRTVAAIKAFHEQPEEER 477
Query: 82 EKTGLANPFGYG-----NKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDY 136
+ G G N + + W + L L+ S PE R + ++
Sbjct: 478 ARV-YTREMGTGVSYISNVDLFQSKAASWRDTLQLRMGP--TAVDQSMIPEVCRQEVMEW 648
Query: 137 LCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIG 196
V +G + L++EGL + + L+ + V ++YP CP+ L +G
Sbjct: 649 DKEVVRVGEVLYGLLSEGLGLDAGRLSEMGLVQGR---VMVGHYYPFCPQPDLT----VG 807
Query: 197 FGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVK 256
H DP ++++ ++ GLQ+ +G W+ V P + IN+GD LQ+++N ++S
Sbjct: 808 LNSHADPGALTVVLQDHIGGLQVRTTEG-WVDVKPLDGALVINIGDLLQIISNEEYKSAD 984
Query: 257 HRVLANGF-QSRLSMIYFGGPPLSEKI-APLPSLLKGGKEESLYKEFTWFEYKNSSYGSR 314
HRVLAN + R+S+ F P EK+ PLP L K +LY+ FT E+ +
Sbjct: 985 HRVLANSANEPRVSVAVFLNPGNREKLFGPLPELTSAEK-PALYRNFTLDEFLTRFFKKE 1161
Query: 315 LADNRLGHFER 325
L L ++ R
Sbjct: 1162LDGKSLTNYFR 1194
>BP076577
Length = 392
Score = 93.2 bits (230), Expect = 5e-20
Identities = 45/89 (50%), Positives = 61/89 (67%)
Frame = -2
Query: 235 SFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKE 294
+F +NVGD L+VMTNGRF SV+HR + N F+SR+SM YFG PPL I P +L +
Sbjct: 391 AFCVNVGDVLEVMTNGRFVSVRHRAMTNSFKSRMSMAYFGAPPLHACIV-APPVLVTPER 215
Query: 295 ESLYKEFTWFEYKNSSYGSRLADNRLGHF 323
SL++ FTW +YK ++Y RL D+R+ F
Sbjct: 214 PSLFRPFTWADYKKATYSLRLGDSRMELF 128
>BP067675
Length = 386
Score = 60.1 bits (144), Expect(2) = 6e-20
Identities = 31/53 (58%), Positives = 38/53 (71%)
Frame = -1
Query: 277 PLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
P +EK+AP PSL+ +E+SLYKEFTW EYK ++Y RLADNRL F I S
Sbjct: 275 PRTEKLAPSPSLVSK-EEDSLYKEFTWGEYKKAAYTLRLADNRLTPFREICWS 120
Score = 53.5 bits (127), Expect(2) = 6e-20
Identities = 26/39 (66%), Positives = 31/39 (78%)
Frame = -3
Query: 245 QVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIA 283
QVMTNG FRSVKHR+L++ SR+SMIYFGG P KI+
Sbjct: 372 QVMTNGSFRSVKHRLLSDTAMSRVSMIYFGGAPSH*KIS 256
>TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partial (69%)
Length = 730
Score = 81.6 bits (200), Expect(2) = 2e-19
Identities = 51/164 (31%), Positives = 88/164 (53%), Gaps = 3/164 (1%)
Frame = +3
Query: 102 DVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKN 161
D+ W + + N+ S E+ R +D Y+ + + ++ LM+E L ++ K+
Sbjct: 243 DIDWESAFFIWHRPKSNIKQISNLSEELRQTMDQYIEKLIQLAEDLSGLMSENLGLE-KD 419
Query: 162 VLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGEHTDPQ-IISLLRSNNTSGLQI 219
+ K ++ +V YP CP + E + G EHTD II LL+ + GL+
Sbjct: 420 YIKKAFSGSDGPAMGTKVAKYPQCP----NPELVRGLREHTDAGGIILLLQDDQVPGLEF 587
Query: 220 SLRDGSWISVPPDYSS-FFINVGDSLQVMTNGRFRSVKHRVLAN 262
S +DG W+ +PP ++ FIN GD ++V++NG ++SV HRV+A+
Sbjct: 588 S-KDGKWVEIPPSKNNAIFINTGDQIEVLSNGLYKSVVHRVMAD 716
Score = 30.4 bits (67), Expect(2) = 2e-19
Identities = 15/49 (30%), Positives = 28/49 (56%), Gaps = 9/49 (18%)
Frame = +2
Query: 27 IPVVDLS------KPDAKSIIVKACEEFGFFKVINHGVS---MEAISQL 66
IP++D S + + +++ +AC+ +G F + NHG+ ME + QL
Sbjct: 8 IPIIDFSTLNGDKRGETMALLHEACKNWGCFLIENHGIDKSLMEKVKQL 154
>BI420511
Length = 384
Score = 90.5 bits (223), Expect = 3e-19
Identities = 51/115 (44%), Positives = 73/115 (63%), Gaps = 1/115 (0%)
Frame = +3
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSS 235
+V +YPACP+ L + G HTD II LL+ + SGLQ+ L+DG W+ VPP S
Sbjct: 24 KVANYPACPKPEL----VKGLRAHTDAGGIILLLQDDKVSGLQL-LKDGQWVDVPPMRHS 188
Query: 236 FFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLK 290
IN+GD L+V+TNG+++SV+HRV+A +R+S+ F P I P P+LL+
Sbjct: 189 IVINLGDQLEVITNGKYKSVEHRVIAKTDGTRMSIASFYNPANDAVIYPAPALLE 353
>BP067791
Length = 468
Score = 67.8 bits (164), Expect(2) = 1e-17
Identities = 32/41 (78%), Positives = 34/41 (82%)
Frame = -2
Query: 257 HRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESL 297
HR+LANGF SRLSMIYFGGPPLSEKIAP PSL+K K L
Sbjct: 446 HRLLANGFPSRLSMIYFGGPPLSEKIAPFPSLMKRRKRREL 324
Score = 37.7 bits (86), Expect(2) = 1e-17
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = -1
Query: 293 KEESLYKEFTWFEYKNSSYGSRLADN 318
K+ + + WFEY+NSSYGSRLAD+
Sbjct: 336 KKRACIRSLHWFEYQNSSYGSRLADH 259
Score = 33.5 bits (75), Expect = 0.050
Identities = 29/77 (37%), Positives = 32/77 (40%), Gaps = 1/77 (1%)
Frame = -3
Query: 250 GRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNS 309
GRFRSV G PP K P GGKEESLYKEFT
Sbjct: 466 GRFRSVDTGFWPMGSHRGYP*FTLVAPP*VRK*HPSLLS*NGGKEESLYKEFTLV*VPEF 287
Query: 310 SYGSRLA-DNRLGHFER 325
+++ RLG FER
Sbjct: 286 KLRFQIS*PPRLGPFER 236
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,631,906
Number of Sequences: 28460
Number of extensions: 74666
Number of successful extensions: 450
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 421
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 422
length of query: 329
length of database: 4,897,600
effective HSP length: 91
effective length of query: 238
effective length of database: 2,307,740
effective search space: 549242120
effective search space used: 549242120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0190.3


