

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0185.17
(469 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
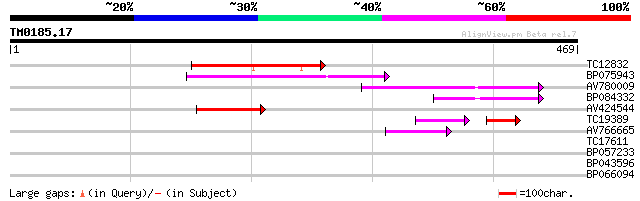
Score E
Sequences producing significant alignments: (bits) Value
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 91 5e-19
BP075943 86 1e-17
AV780009 65 2e-11
BP084332 52 3e-07
AV424544 48 4e-06
TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 40 4e-05
AV766665 42 2e-04
TC17611 27 7.1
BP057233 27 7.1
BP043596 27 9.3
BP066094 27 9.3
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 90.5 bits (223), Expect = 5e-19
Identities = 50/120 (41%), Positives = 76/120 (62%), Gaps = 9/120 (7%)
Frame = +2
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD--GEIIGLSQ 208
++ LYVDDML+ G + + V++ K+ L+ FDMKDLG A+ ILG++I RD I LSQ
Sbjct: 89 IILLLYVDDMLVVGPNKDRVQELKAQLAREFDMKDLGPANKILGMQIHRDRKDRRIWLSQ 268
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPH-------KRCPVAQLEYSKAIRCLMYAM 261
+Y++KVL++F+ DC +STP N KL +R ++++ Y+ A+ LMYAM
Sbjct: 269 KNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSMIPSSEAERMEMSRVPYASAVGSLMYAM 448
>BP075943
Length = 547
Score = 85.9 bits (211), Expect = 1e-17
Identities = 57/170 (33%), Positives = 87/170 (50%), Gaps = 2/170 (1%)
Frame = -3
Query: 147 SGKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGL 206
SG LYVDD+L+ G D++E++ KS L + F +KDLGEA LG++I R I L
Sbjct: 539 SGSFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVL 360
Query: 207 SQSHYIEKVLKKFDHFDCKSVSTPFDQNTK-LQPHKRCPVAQL-EYSKAIRCLMYAMTCT 264
+Q Y +++ HF + N++ L + P+ + Y + + L+Y T T
Sbjct: 359 NQRKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNT-T 183
Query: 265 RPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSVL 314
RPDI + V +LS++ P+ H + LKYL G+ L Y S L
Sbjct: 182 RPDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPASSSTL 33
>AV780009
Length = 529
Score = 65.5 bits (158), Expect = 2e-11
Identities = 44/151 (29%), Positives = 76/151 (50%), Gaps = 1/151 (0%)
Frame = -1
Query: 292 RVLKYLNGTINYSLIYNG-YPSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKK 350
RVL+Y+ G L ++ P L+ Y+ + W C S +G LG +SW +KK
Sbjct: 517 RVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWRTKK 338
Query: 351 QTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQV 410
QT ++ S+ AE+ ALAA E L L + + + V + CD+Q L A++
Sbjct: 337 QTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKL--NVPLPVPLFCDNQSALHIAHNPT 164
Query: 411 YNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
++ +++HI L V+ + G+I ++ + T
Sbjct: 163 FHERTKHIELDCHVVRAKLQAGLIHLLPIST 71
>BP084332
Length = 368
Score = 51.6 bits (122), Expect = 3e-07
Identities = 26/91 (28%), Positives = 54/91 (58%)
Frame = +2
Query: 351 QTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQV 410
Q+ IA ST AE+I+ A CS + + +++ L + I L + I+CD+ +S + + +
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI---LESNIPIYCDNTAAISLSKNPI 202
Query: 411 YNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
+ +++HI +++ ++D + GV+ + FV T
Sbjct: 203 LHSRAKHIEVKYHFIRDYVQKGVLLLKFVDT 295
>AV424544
Length = 276
Score = 47.8 bits (112), Expect = 4e-06
Identities = 23/57 (40%), Positives = 35/57 (61%)
Frame = +3
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHY 211
+YVDD+++ G DLNE++ K+ L F +KDLG LG+++ R + LSQ Y
Sbjct: 105 VYVDDLILAGNDLNEIQCVKNKLDIQFRIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (6%)
Length = 498
Score = 40.0 bits (92), Expect(2) = 4e-05
Identities = 19/45 (42%), Positives = 26/45 (57%)
Frame = +3
Query: 336 IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLL 380
+ GGAV+W S+ Q C+A ST AEFIA E + ++N L
Sbjct: 3 LVTFAGGAVAWPSRLQKCVALSTAEAEFIAATEACHELLWMKNFL 137
Score = 23.5 bits (49), Expect(2) = 4e-05
Identities = 7/28 (25%), Positives = 18/28 (64%)
Frame = +2
Query: 395 IHCDSQCTLSKAYSQVYNGKSRHIVLRH 422
++CD+Q + + ++ +S+HI +R+
Sbjct: 173 LYCDNQGAIYLGQNSTFHSRSKHIDVRY 256
>AV766665
Length = 601
Score = 42.0 bits (97), Expect = 2e-04
Identities = 22/54 (40%), Positives = 31/54 (56%)
Frame = +3
Query: 312 SVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIA 365
S ++G+TYA ++ + D ST G L G V+W SK+Q IA S+ AE A
Sbjct: 435 SSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTWRSKQQNIIARSSGEAELRA 596
Score = 30.0 bits (66), Expect = 0.84
Identities = 16/39 (41%), Positives = 21/39 (53%)
Frame = +1
Query: 154 CLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVI 192
C+YVD M I G D E + + L+ FD+K L VI
Sbjct: 118 CVYVDHMTIVGDDEVEKQILREKLAPQFDLKKLKYISVI 234
>TC17611
Length = 409
Score = 26.9 bits (58), Expect = 7.1
Identities = 10/32 (31%), Positives = 19/32 (59%)
Frame = -1
Query: 212 IEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRC 243
I K +++ +HFD K + +N ++QP + C
Sbjct: 247 ISKKVRRTNHFDSKDLHRIL*KNIQIQPQRTC 152
>BP057233
Length = 473
Score = 26.9 bits (58), Expect = 7.1
Identities = 11/54 (20%), Positives = 28/54 (51%)
Frame = -2
Query: 391 RQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKE 444
R+ + CD+ + A + V + +S+HI + +++D + + + +V T +
Sbjct: 457 RKPILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQ 296
>BP043596
Length = 569
Score = 26.6 bits (57), Expect = 9.3
Identities = 20/81 (24%), Positives = 32/81 (38%)
Frame = +3
Query: 389 LMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKESCRS 448
L+ +V C+L ++ +HI H + I N +S K CRS
Sbjct: 327 LLSKVRYDIGDSCSLDYLIDSIFPLFRKHIEPDLHHAQIRINNSFVSSFREAIYKVKCRS 506
Query: 449 FDERSDKGNGVEDIVGDESQT 469
S +G+ G+E+QT
Sbjct: 507 ---ESSEGHSDSVFQGNETQT 560
>BP066094
Length = 532
Score = 26.6 bits (57), Expect = 9.3
Identities = 11/42 (26%), Positives = 23/42 (54%)
Frame = +2
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIK 196
+YVDD++ + + ++ + F+M+ +GE LGI+
Sbjct: 407 IYVDDIIFGSANPSLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.351 0.154 0.546
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,612,997
Number of Sequences: 28460
Number of extensions: 126128
Number of successful extensions: 1023
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 1012
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1020
length of query: 469
length of database: 4,897,600
effective HSP length: 94
effective length of query: 375
effective length of database: 2,222,360
effective search space: 833385000
effective search space used: 833385000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0185.17


