

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
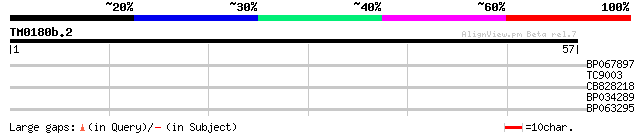
Query= TM0180b.2
(57 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP067897 26 1.8
TC9003 25 2.4
CB828218 25 3.1
BP034289 24 5.3
BP063295 23 9.0
>BP067897
Length = 500
Score = 25.8 bits (55), Expect = 1.8
Identities = 8/25 (32%), Positives = 15/25 (60%)
Frame = +1
Query: 21 WAAALQGHMMWLQRQDSFKHKFRNL 45
WAA++QG +W + H+F+ +
Sbjct: 295 WAASIQGSQIWPHQSMHMTHEFKTV 369
>TC9003
Length = 613
Score = 25.4 bits (54), Expect = 2.4
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = +1
Query: 10 KKFWVASFLIAWAAALQ 26
+KFW+ FL++W +LQ
Sbjct: 154 RKFWMFIFLLSWTPSLQ 204
>CB828218
Length = 502
Score = 25.0 bits (53), Expect = 3.1
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = -1
Query: 17 FLIAWAAALQGHMMWLQRQDSFKHKFRNLDDDDR 50
F + W +G M +R+ + KHKFR + R
Sbjct: 466 FFLCWIILKRGSQMKSERKIT*KHKFRKFQNRAR 365
>BP034289
Length = 492
Score = 24.3 bits (51), Expect = 5.3
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = +3
Query: 13 WVASFLIAWAAALQGHMMWL 32
WV S L +WA+ + M+W+
Sbjct: 312 WVLSLLQSWASTVAVFMIWM 371
>BP063295
Length = 537
Score = 23.5 bits (49), Expect = 9.0
Identities = 7/21 (33%), Positives = 13/21 (61%)
Frame = -1
Query: 11 KFWVASFLIAWAAALQGHMMW 31
K W AS ++W + ++ H +W
Sbjct: 519 KSWHASSALSWXSPMRAH*VW 457
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.133 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,189,570
Number of Sequences: 28460
Number of extensions: 12039
Number of successful extensions: 59
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 59
length of query: 57
length of database: 4,897,600
effective HSP length: 33
effective length of query: 24
effective length of database: 3,958,420
effective search space: 95002080
effective search space used: 95002080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0180b.2


