

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.7
(1813 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
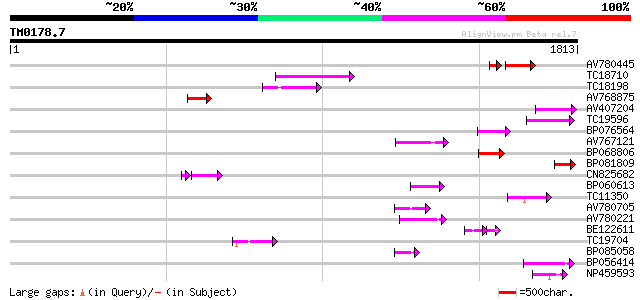
Score E
Sequences producing significant alignments: (bits) Value
AV780445 117 4e-39
TC18710 154 2e-37
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 122 5e-28
AV768875 110 3e-24
AV407204 100 3e-21
TC19596 99 5e-21
BP076564 88 1e-17
AV767121 84 2e-16
BP068806 84 3e-16
BP081809 80 3e-15
CN825682 77 7e-15
BP060613 76 5e-14
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 69 7e-12
AV780705 67 3e-11
AV780221 66 6e-11
BE122611 49 9e-11
TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA po... 65 1e-10
BP085058 52 8e-07
BP056414 52 1e-06
NP459593 Pge1 protein [Lotus japonicus] 49 9e-06
>AV780445
Length = 504
Score = 117 bits (292), Expect(2) = 4e-39
Identities = 51/98 (52%), Positives = 68/98 (69%)
Frame = +1
Query: 1584 NNIRFLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINL 1643
N+++ LAG+W VWKWRNN+V + PW++D AW+KL H+HDE + FS GD D +L +
Sbjct: 211 NSVKCLAGVW*VWKWRNNMVFEDSPWSVDEAWRKLGHEHDEFITFSDGDAGDPGSWLSSR 390
Query: 1644 WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWL 1681
W+PP VKLN DGS ++ MG GGLIRD +G WL
Sbjct: 391 WQPPRQGTVKLNVDGSGREQDRCMGAGGLIRDDQGKWL 504
Score = 63.5 bits (153), Expect(2) = 4e-39
Identities = 24/38 (63%), Positives = 29/38 (76%)
Frame = +2
Query: 1534 RCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFRLPD 1571
RC +PEED HCLRDCPHSQE+W +LGAW W+ + D
Sbjct: 65 RCPAPEEDI*HCLRDCPHSQEIWLRLGAWAWTNLGVGD 178
>TC18710
Length = 843
Score = 154 bits (388), Expect = 2e-37
Identities = 85/254 (33%), Positives = 136/254 (53%), Gaps = 1/254 (0%)
Frame = +2
Query: 851 YFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGF 910
+F L+ + P N+ + + +P + +++++ + + G DG
Sbjct: 23 FFPELYTSTPPNSSESA*MLFLP*SLKI*IKSLNLR*LQKKLRLLFFLLETSQPRGMDGM 202
Query: 911 HPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCN 970
+ F++K ++V D + + E I + + ETL+ LIPKV P ++ +FRPIS C
Sbjct: 203 NGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLVTLIPKVPHPESINQFRPISGCT 382
Query: 971 VVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMA-KKGTL 1029
+YK+I+K+ V R++ + +I PMQ+ F+ G I DN + QE H + A +
Sbjct: 383 FLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDNLLIVQEAFHAINRPGALGRNHS 562
Query: 1030 AFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGR 1089
K+D+ KAYD V W FLE +L+ FGF V++IM VSG + + NG P P R
Sbjct: 563 IIKLDMNKAYDRVEWKFLESSLLAFGFSTNWVKMIMILVSGVSYNYKINGVVGPKLLPQR 742
Query: 1090 GLRKGDPLSPYLFV 1103
GLR+GDP SPYLF+
Sbjct: 743 GLRQGDPFSPYLFL 784
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 122 bits (306), Expect = 5e-28
Identities = 66/194 (34%), Positives = 114/194 (58%), Gaps = 6/194 (3%)
Frame = -3
Query: 808 DRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNM 867
D NT +FHA TI RR N+I ++K G W + ++ + E NYF ++++ P +
Sbjct: 566 DHNTRFFHASTIQRRDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSSP------L 405
Query: 868 QHVM-----MPRL-TEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWEL 921
Q++ +P+L T+ HK A+V +E+ +A+ M A G DG + F+++ E+
Sbjct: 404 QYLRECLAPIPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNREI 225
Query: 922 VGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVV 981
V +D+ + D F+ G + L ETL+ LIPK+ +++FRPIS C+ +YK+I+K++V
Sbjct: 224 VKNDVNNAILDFFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIV 45
Query: 982 NRIRPLLNRIIGPM 995
R++P + +I PM
Sbjct: 44 ARLKPDMVNLISPM 3
>AV768875
Length = 232
Score = 110 bits (274), Expect = 3e-24
Identities = 45/77 (58%), Positives = 63/77 (81%)
Frame = +1
Query: 569 KREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRASRFAAILEGCGMVDL 628
+R+ LW HL IR+++ +PW+++GDMNEIL P++VRGG F P RA RFA +L+ C ++DL
Sbjct: 1 QRDILWQHLSTIRASISIPWLLIGDMNEILFPSQVRGGEFHPNRAQRFATVLDDCNLIDL 180
Query: 629 GMSGGRYTWFRKQNNRL 645
GM GG++TWFRK+NNRL
Sbjct: 181 GMVGGKFTWFRKRNNRL 231
>AV407204
Length = 432
Score = 99.8 bits (247), Expect = 3e-21
Identities = 59/133 (44%), Positives = 70/133 (52%), Gaps = 2/133 (1%)
Frame = -2
Query: 1680 WLGGFMSH--TSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRL 1737
W+ GF+S + FLAE LALRDGL LAW G R+++ DC L D + R
Sbjct: 428 WVNGFISQYLEAASCSFLAEALALRDGLRLAWDNGTRKLVCNSDCKELIDAIADPSRASF 249
Query: 1738 HDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLARRGGFLSTIGVVELMVAPLELEH 1797
H H VL+EI LM R WR + W RD N AD LA+RG L G L ELE
Sbjct: 248 HAHGWVLREIHALMTRNWRVELFWCCRDVNMVADCLAKRGSLLPVAGYSALATPSPELEV 69
Query: 1798 LLLKDRLAIR*SV 1810
LLLKD + * V
Sbjct: 68 LLLKDIIGFS*FV 30
>TC19596
Length = 499
Score = 99.4 bits (246), Expect = 5e-21
Identities = 58/154 (37%), Positives = 89/154 (57%)
Frame = +1
Query: 1653 KLNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQG 1712
++++DG+F+ MG GG++RDA GAW+ GF + + G+ AE+ AL+ GL L W
Sbjct: 4 RMDSDGAFKHDDDRMGMGGIVRDAHGAWISGFYAGSLGGDALRAEIAALKHGLTLLWNAH 183
Query: 1713 HRRIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADF 1772
RR EVDC + + L DR + H A L +IR L++R W ++ + R++NA+AD
Sbjct: 184 VRRATCEVDCLDIVEALE-NDRYQFHALASELLDIRLLLDRDWTVTLAYVPREANAAADC 360
Query: 1773 LARRGGFLSTIGVVELMVAPLELEHLLLKDRLAI 1806
LA G L + L P EL+ +L +D LA+
Sbjct: 361 LAGLGASL-LCPLTCLESPPQELQPILARDLLAL 459
>BP076564
Length = 371
Score = 87.8 bits (216), Expect = 1e-17
Identities = 41/105 (39%), Positives = 62/105 (59%)
Frame = +2
Query: 1497 LHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW 1556
L EK + L LH ALP NA R+ C +A + GC RCS+ +D +HCL+DCPHS+E+W
Sbjct: 56 LKVVEKCKFMVGLGLHQALPINARRHACSMAATGGCGRCSAAVKDILHCLQDCPHSREVW 235
Query: 1557 GKLGAWGWSTFRLPDLKA*VKTQLQGSNNIRFLAGLWGVWKWRNN 1601
+LG + F D + + + ++ N + + LW +W+ RNN
Sbjct: 236 NRLGCSSFGGFLSSDGWSWLLSIMKQPNVEKLIYILWWIWR*RNN 370
>AV767121
Length = 525
Score = 84.3 bits (207), Expect = 2e-16
Identities = 53/176 (30%), Positives = 81/176 (45%), Gaps = 6/176 (3%)
Frame = +2
Query: 1233 LSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRG 1292
LS WK ++ G++ L +SV++A+P + + F LP + + MR F+W + +
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 1293 WHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGESI 1352
V W + +PKE GGL IKD+ FN +LLGK W LW +V+ + H
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQPDH---- 349
Query: 1353 FSVHRRANASPIWQGILK-----ARDQLQEGFQFRMGDG-SSSVWYHDWTGKGKLA 1402
+ S W IL A G + +G+G + W DW G G L+
Sbjct: 350 -----YSCGSSWWNDILSLCPEDADGWFSSGLKKLVGEGDQTKFWSEDWLGSGLLS 502
>BP068806
Length = 418
Score = 83.6 bits (205), Expect = 3e-16
Identities = 37/83 (44%), Positives = 54/83 (64%)
Frame = +3
Query: 1500 PEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKL 1559
PEK+ V++W ++ A+ N+ R+RCKLA SP RC ED HCLRDC ++++W +L
Sbjct: 153 PEKIRVWVWFIMQGAIQVNSFRFRCKLATSPAGPRCQHQFEDIWHCLRDCRIARDVWVRL 332
Query: 1560 GAWGWSTFRLPDLKA*VKTQLQG 1582
A GW F + + +A VK+Q QG
Sbjct: 333 NALGWPGFFVSNAQAWVKSQAQG 401
>BP081809
Length = 299
Score = 80.1 bits (196), Expect = 3e-15
Identities = 38/66 (57%), Positives = 48/66 (72%)
Frame = -1
Query: 1743 VLQEIRELMERPWRCSVHWTNRDSNASADFLARRGGFLSTIGVVELMVAPLELEHLLLKD 1802
VL EI EL+ R WRC ++W +RD N D+LARRGGF S+ GV EL P +L+HLLLKD
Sbjct: 296 VLGEIHELLNRSWRCIMNWISRDGNGPTDYLARRGGFSSSSGVRELDAPPPKLDHLLLKD 117
Query: 1803 RLAIR* 1808
+L +R*
Sbjct: 116 KLTLR* 99
>CN825682
Length = 626
Score = 77.4 bits (189), Expect(2) = 7e-15
Identities = 41/102 (40%), Positives = 54/102 (52%), Gaps = 1/102 (0%)
Frame = -3
Query: 581 RSNLVLPWMIVGDMNEILHPNEVRGG*FL-PTRASRFAAILEGCGMVDLGMSGGRYTWFR 639
+SN W GD NEILH E G PTR F L+G ++DL + G R+TW
Sbjct: 321 KSNTNTFWTCYGDFNEILHHQEKEGARDQDPTRIQVFRDFLDGANLMDLELKGCRFTWLS 142
Query: 640 KQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCP 681
N L+ +RLDR + + WR+ +P A+ AL I SDH P
Sbjct: 141 NLRNGLVTKERLDRVLVNWTWRLDYPNATAMALPIISSDHSP 16
Score = 21.6 bits (44), Expect(2) = 7e-15
Identities = 10/29 (34%), Positives = 15/29 (51%)
Frame = -1
Query: 549 VWRENLSWVCSAIYASPIPVKREELWSHL 577
VW + + IYA+PI +R LW +
Sbjct: 416 VWPNKEIVLGTFIYANPIFRQRRNLWPQI 330
>BP060613
Length = 378
Score = 75.9 bits (185), Expect = 5e-14
Identities = 36/106 (33%), Positives = 53/106 (49%)
Frame = +1
Query: 1283 IWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVL 1342
+W + G RG H W LT+ K+ GGL KD N +LL K W + +N LWVQ+L
Sbjct: 43 VWWSSKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQIL 222
Query: 1343 AHKYLHGESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS 1388
Y +R AS +W +L R+ L + Q+++G G +
Sbjct: 223 KALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSGET 360
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 68.9 bits (167), Expect = 7e-12
Identities = 49/149 (32%), Positives = 72/149 (47%), Gaps = 10/149 (6%)
Frame = -3
Query: 1592 LWGVWKWRNNVVL---DPLPWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINL----- 1643
LW +WK RN V+ +P P + ++ D + L+ S + IN+
Sbjct: 978 LWAIWKNRNRVIFQLEEPNPVNTIFQIRHMQQDAN-LMSTSQSNKQTTRERNINVTSARD 802
Query: 1644 --WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVLAL 1701
W+PPP KLN+D S++ + S G +IRD G + G S +P ++E LAL
Sbjct: 801 RSWRPPPLGVYKLNSDASWKAPSPSCTVGVVIRDNLGLLISGTASRPLAPSPIVSEALAL 622
Query: 1702 RDGLCLAWVQGHRRIIREVDCAGLSDILR 1730
R+GL LA G RI+ E DC L + R
Sbjct: 621 REGLILARSLGLDRILSESDCQPLIEACR 535
>AV780705
Length = 524
Score = 67.0 bits (162), Expect = 3e-11
Identities = 37/119 (31%), Positives = 58/119 (48%), Gaps = 2/119 (1%)
Frame = -2
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
K+KL WK LLN G+ L K++I AIP+Y M + P++ + + + F W K+ G
Sbjct: 415 KEKLEGWKETLLNQAGKEVLIKAIIQAIPSYAMTIVHFPKTFCNSLYALVADF-WWKSQG 239
Query: 1290 QRGWHRVNWKTLTQP--KEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKY 1346
+ P KE GG+ +D+ N + L + W + N LWV+V+ Y
Sbjct: 238 EGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQNLAFLARQAWRVLTNPEALWVRVMKSLY 62
>AV780221
Length = 461
Score = 65.9 bits (159), Expect = 6e-11
Identities = 45/155 (29%), Positives = 74/155 (47%), Gaps = 4/155 (2%)
Frame = +3
Query: 1247 VCLAKSVISAIPTYTMQVFWLPRS-INHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPK 1305
+CL +SV+ A+P + + F LP +N F + G + V W T+ +PK
Sbjct: 3 LCLIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFFGREVEGGKKIAWVKWSTVCRPK 182
Query: 1306 EHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG--ESIFSVHRRANASP 1363
+ GGL ++++G FN +LLGK W + LW +VL KY + +S S +++
Sbjct: 183 DEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKYKNAIPQSASSWWNDLHSTC 362
Query: 1364 IWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTG 1397
G +Q G R+G+G+ W +W G
Sbjct: 363 FEDG---GGGWMQRGLCRRIGEGTEVKFWNENWLG 458
>BE122611
Length = 357
Score = 48.5 bits (114), Expect(2) = 9e-11
Identities = 29/75 (38%), Positives = 38/75 (50%)
Frame = -3
Query: 1454 GIDRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHD 1513
G DR W + S+G S Y L E G W + K +APEK+ F+WLV +
Sbjct: 337 GTDRILWTEESNGCYSAKSTYNLLIED---GGNESRFWSLVWKANAPEKVRFFLWLVGRN 167
Query: 1514 ALPTNANRYRCKLAV 1528
ALP+NA R+ L V
Sbjct: 166 ALPSNARRHHITLLV 122
Score = 36.6 bits (83), Expect(2) = 9e-11
Identities = 18/42 (42%), Positives = 20/42 (46%)
Frame = -2
Query: 1526 LAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTF 1567
LA S C RC +P ED H R CP S AW W+ F
Sbjct: 131 LASSAACIRCGAPSEDANHVFRWCPGS--------AWMWTQF 30
>TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA
polymerase-like protein, partial (3%)
Length = 530
Score = 65.1 bits (157), Expect = 1e-10
Identities = 37/153 (24%), Positives = 78/153 (50%), Gaps = 7/153 (4%)
Frame = -3
Query: 711 KEVVQSAWTGETET-------VAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQR 763
KE ++ AW +ET ++GK + +++ + + + ++ +L +Q
Sbjct: 522 KETIKKAWDNTSETASSHCSNLSGKTTNTKLALTEWHHQNFSRADKEIAELKNQLEHLQ- 346
Query: 764 ELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRK 823
NY + + + + + + L RQEEL W Q++R + D+NT +FHA T+ RR+
Sbjct: 345 --NYSNSGEDWQEIKQRKMRLKDLWRQEELFWSQRSRIKWLKSGDKNTKFFHASTVQRRE 172
Query: 824 RNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLF 856
RN+I R+K G+W + + ++ +++ ++
Sbjct: 171 RNRIERIKDMQGTWVRSQLAINRDVPDFYPDIY 73
>BP085058
Length = 452
Score = 52.0 bits (123), Expect = 8e-07
Identities = 31/79 (39%), Positives = 43/79 (54%)
Frame = +2
Query: 1232 KLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQR 1291
+L+ K LLN ++ LAK V+SAI Y MQ+ WLP+SI +I +R F+W
Sbjct: 212 RLAC*KAYLLNKPARLTLAKPVLSAISMYVMQLNWLPQSICDQICTVVRHFVW---ENVN 382
Query: 1292 GWHRVNWKTLTQPKEHGGL 1310
G V W+T+ K GL
Sbjct: 383 GIPLVAWETIQSRKSMVGL 439
>BP056414
Length = 606
Score = 51.6 bits (122), Expect = 1e-06
Identities = 43/167 (25%), Positives = 76/167 (44%), Gaps = 4/167 (2%)
Frame = -3
Query: 1644 WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTSI---GNPFLAEVLA 1700
W PP + K+N DG+ + GG++RDA G+W GF + F E++A
Sbjct: 562 WTRPPEGWFKVNVDGALSGDLQA-SCGGVVRDAAGSWSKGFARSLGVLRWHXAFFVELMA 386
Query: 1701 LRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVH 1760
++ + ++I E D + ++L+ + H + + +I++ + +
Sbjct: 385 VQTAVDFIMSWDIPQVIIESDSQQVVELLQNFNASSPHQYGEIAWDIQQKISVHGSIVIT 206
Query: 1761 WTNRDSNASADFLARRGGFLSTIGVVELMVAPL-ELEHLLLKDRLAI 1806
T R++N AD+LA+ G LS AP E LL KD ++I
Sbjct: 205 STPREANFLADYLAKVG--LSLPFGTHSFDAPFGECHSLLQKDLMSI 71
>NP459593 Pge1 protein [Lotus japonicus]
Length = 633
Score = 48.5 bits (114), Expect = 9e-06
Identities = 34/123 (27%), Positives = 57/123 (45%), Gaps = 10/123 (8%)
Frame = +1
Query: 1672 LIRDAKGAWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCA-------- 1723
++RD++GAWL + + G+ FL E+LA++ G+ A GH + DC+
Sbjct: 16 VVRDSRGAWLSATLVCYNYGSAFLTEILAVKLGIRHALDIGHIYVFCLSDCSQALQALHA 195
Query: 1724 --GLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLARRGGFLS 1781
++ E CR+HD LQ++ ++ +R N +AD L R G L
Sbjct: 196 DTNITSYWTREKICRVHDIIATLQDV----------NLQHIDRVKNNTADKLVREAGRLE 345
Query: 1782 TIG 1784
+G
Sbjct: 346 YMG 354
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.144 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,001,774
Number of Sequences: 28460
Number of extensions: 626579
Number of successful extensions: 3748
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 3672
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3739
length of query: 1813
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1710
effective length of database: 1,966,220
effective search space: 3362236200
effective search space used: 3362236200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0178.7


