

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
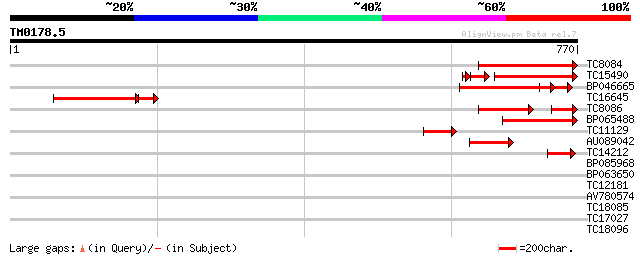
Score E
Sequences producing significant alignments: (bits) Value
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 184 4e-47
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 150 3e-42
BP046665 166 2e-41
TC16645 143 5e-39
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 90 1e-28
BP065488 101 4e-22
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 65 5e-11
AU089042 60 1e-09
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 45 6e-05
BP085968 34 0.077
BP063650 31 0.15
TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%) 30 1.5
AV780574 29 2.5
TC18085 28 7.2
TC17027 similar to UP|Q93XA5 (Q93XA5) Homeodomain leucine zipper... 27 9.4
TC18096 similar to UP|Q9SY59 (Q9SY59) F14N23.5, partial (5%) 27 9.4
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 184 bits (468), Expect = 4e-47
Identities = 87/134 (64%), Positives = 110/134 (81%)
Frame = +1
Query: 637 IDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQF 696
I + LCN TR+IV L Y+I AT++ G +G+ FIPR+ + PS+S PFKF+RR F
Sbjct: 43 IQKFENLCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 222
Query: 697 PVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVS 756
P++LCFAMTINKSQGQSLSHV LYL RPVFTHGQLYVALSRV+SRKGLK+L++D+E V+
Sbjct: 223 PISLCFAMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVT 402
Query: 757 NTTRNVMYQEVFDN 770
NTT+NV+Y+EVF+N
Sbjct: 403 NTTKNVVYREVFEN 444
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 150 bits (380), Expect(3) = 3e-42
Identities = 70/112 (62%), Positives = 92/112 (81%)
Frame = +2
Query: 659 IVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVG 718
I T++ G +G+ IPRM + PS+S PFKF+RRQ P++LCFAMTINKSQG+SLSHVG
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVG 316
Query: 719 LYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
LYL RPV THG LYVAL RV+SRK LK+L++D+E ++NTT+NV+Y+E+F+N
Sbjct: 317 LYLSRPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFEN 472
Score = 38.1 bits (87), Expect(3) = 3e-42
Identities = 18/26 (69%), Positives = 21/26 (80%)
Frame = +1
Query: 626 KVGVPIMLIRNIDQAAGLCNDTRMIV 651
K G IML+RNI QA+GLCN TR+IV
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIV 120
Score = 21.6 bits (44), Expect(3) = 3e-42
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = +3
Query: 615 CSEIPNHAIKLK 626
CS+IPNH I K
Sbjct: 9 CSDIPNHKITEK 44
>BP046665
Length = 524
Score = 166 bits (419), Expect = 2e-41
Identities = 83/131 (63%), Positives = 98/131 (74%), Gaps = 1/131 (0%)
Frame = -1
Query: 612 DFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGE 671
D C IPNH I LK G PIML+RNI QA G CN TR+IV L +I AT++ +G+
Sbjct: 524 DITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGD 345
Query: 672 TTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQL 731
FIPRM + PS+S PFKF+RRQFP++LC AMTINKSQGQSLSHVGLYL R VFTHGQL
Sbjct: 344 DIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQL 165
Query: 732 YVAL-SRVKSR 741
YVAL +++K R
Sbjct: 164 YVALHAKIKKR 132
Score = 45.1 bits (105), Expect = 4e-05
Identities = 20/45 (44%), Positives = 33/45 (72%)
Frame = -2
Query: 720 YLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMY 764
Y+ +++H LS ++SRKGLK+L++D+E V+NTT+NV+Y
Sbjct: 202 YIFLAMYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC16645
Length = 596
Score = 143 bits (360), Expect(2) = 5e-39
Identities = 78/118 (66%), Positives = 85/118 (71%)
Frame = +3
Query: 60 MDGCQQEVPRGEKSHICRIFFQVCVQQKINNMTS*EKRHFSWSIAIYSSWNGRKLLHACF 119
+D QQEVP GEKSHICRI F V + N M S* K+ FSWS+AIY S NGR+LL+ACF
Sbjct: 3 LDERQQEVPGGEKSHICRISF*VSLPPTHNQMGS*AKKIFSWSVAIYCSRNGRELLYACF 182
Query: 120 TN*AKRL*QF*KHKNCKGCCLSHILRCMRSDGINGG*PRVC*WNFNCK*TWFRISAKE 177
N AKRL*QF*KHKN GC LSH CMRS+G GG* RVC*WNFN K* W S KE
Sbjct: 183 INHAKRL*QF*KHKNG*GCGLSHFP*CMRSNGTVGG*SRVC*WNFNFK*IWIWNSTKE 356
Score = 35.8 bits (81), Expect(2) = 5e-39
Identities = 17/27 (62%), Positives = 22/27 (80%)
Frame = +1
Query: 176 KEIIYQNADNKLDFQATINLEKVLDSF 202
+++ Y NAD+KLDF AT +LEKVL SF
Sbjct: 352 RKLFY*NADDKLDFPATRSLEKVLVSF 432
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 90.1 bits (222), Expect(2) = 1e-28
Identities = 41/75 (54%), Positives = 55/75 (72%)
Frame = +1
Query: 637 IDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQF 696
I + LC+ TR+IV L Y+I AT++ G +G+ FIPR+ + PS+S PFKF+RR F
Sbjct: 151 IQKFENLCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 330
Query: 697 PVTLCFAMTINKSQG 711
P++LCFAMTINKSQG
Sbjct: 331 PISLCFAMTINKSQG 375
Score = 54.3 bits (129), Expect(2) = 1e-28
Identities = 23/35 (65%), Positives = 33/35 (93%)
Frame = +3
Query: 736 SRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
SRV+SRKGLK+L++D+E V+NTT+NV+Y+EVF+N
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFEN 473
>BP065488
Length = 439
Score = 101 bits (252), Expect = 4e-22
Identities = 55/103 (53%), Positives = 72/103 (69%), Gaps = 2/103 (1%)
Frame = -1
Query: 670 GETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQ-SLSHVGLYLPRPVFTH 728
G+ FIPRM++ PS S P KF+R QFP++LCFAMTINKSQ V L ++H
Sbjct: 379 GDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQXSVHYPTVALISFLACYSH 200
Query: 729 -GQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
+LYVALS V+SRKGLK+L++ +E V+N T+N +YQEVF N
Sbjct: 199 MDKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKN 71
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 64.7 bits (156), Expect = 5e-11
Identities = 28/46 (60%), Positives = 37/46 (79%)
Frame = +2
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTS 607
PTLESVE++N FML ++PG+ TEYLS DT CK DED+ + + WFT+
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WFTT 595
>AU089042
Length = 191
Score = 60.1 bits (144), Expect = 1e-09
Identities = 29/60 (48%), Positives = 41/60 (68%)
Frame = +3
Query: 625 LKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSN 684
LK GVP+ML+ N+ + GLCN TR+IV+ L +I ATIL+G +G +I M+L PS+
Sbjct: 9 LKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPSD 188
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 44.7 bits (104), Expect = 6e-05
Identities = 21/38 (55%), Positives = 28/38 (73%)
Frame = +2
Query: 731 LYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVF 768
LYVA+SRVKS+ GLK+LI D T+N++Y+EVF
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVF 115
>BP085968
Length = 341
Score = 34.3 bits (77), Expect = 0.077
Identities = 20/43 (46%), Positives = 26/43 (59%), Gaps = 4/43 (9%)
Frame = +1
Query: 352 YCYLEVE----LLIQDFLFLFQ*MRYQPAIFVTVPQKPNY*KK 390
YC+L V LLIQ FLF++Q M YQP I ++ NY K+
Sbjct: 184 YCFLVVAWRKGLLIQGFLFIYQLMIYQPVISSKDLKRLNYFKR 312
Score = 33.5 bits (75), Expect = 0.13
Identities = 30/87 (34%), Positives = 33/87 (37%)
Frame = +3
Query: 314 FYMVLEELDKHLFGIHYLLLCVQGALSS*MSHLAELHRYCYLEVELLIQDFLFLFQ*MRY 373
FYM LE +KHL GIH L + * H L C LE F
Sbjct: 81 FYMDLEVREKHLCGIHGLQV*DHKDSWF*TWHQVVLLLGCCLEERTAHSRFSIHISINDI 260
Query: 374 QPAIFVTVPQKPNY*KKASLIIWDETP 400
QK +KAS IIWDE P
Sbjct: 261 STCNIKQGSQKAELLQKASSIIWDEAP 341
>BP063650
Length = 497
Score = 31.2 bits (69), Expect(2) = 0.15
Identities = 12/31 (38%), Positives = 21/31 (67%)
Frame = +1
Query: 443 QILPVISKGSRSEIVGSAINSSYLWKHCKVM 473
+ L V+ KG +I+ + +N+SYL HC+V+
Sbjct: 58 KFLTVVYKGRMQDIIHAIVNASYL*DHCQVL 150
Score = 20.8 bits (42), Expect(2) = 0.15
Identities = 9/10 (90%), Positives = 9/10 (90%)
Frame = +3
Query: 435 VVLGGDFRQI 444
VVL GDFRQI
Sbjct: 33 VVLQGDFRQI 62
>TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%)
Length = 682
Score = 30.0 bits (66), Expect = 1.5
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = +2
Query: 451 GSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEF 496
G ++ G A LW+ C+++K+ V +QN S EIK +
Sbjct: 314 GRNKKLQGLAAAIIKLWERCEIVKIAVKRGVQNTNSEIMAEEIKVY 451
>AV780574
Length = 566
Score = 29.3 bits (64), Expect = 2.5
Identities = 16/56 (28%), Positives = 26/56 (45%)
Frame = -2
Query: 298 IRMFWMLFCLIMVDSFFYMVLEELDKHLFGIHYLLLCVQGALSS*MSHLAELHRYC 353
I FW LFCL M+ +F Y ++ L + + CV + + +L + R C
Sbjct: 355 ISNFWSLFCLFMLQTFLYEII------LPRLETFIDCVYASSPQPILNLGTIFRCC 206
>TC18085
Length = 551
Score = 27.7 bits (60), Expect = 7.2
Identities = 20/67 (29%), Positives = 28/67 (40%), Gaps = 4/67 (5%)
Frame = -1
Query: 84 VQQKINNMTS*EKRHFSWSIAIYSSWN----GRKLLHACFTN*AKRL*QF*KHKNCKGCC 139
+ +N MT EK ++W I + N R LL *+ +F K + GCC
Sbjct: 473 IDADLNGMT--EKDSYNWMIHWHRKQNHNL*DRWLLGMMIPG*SNSPCRFRKKRRLHGCC 300
Query: 140 LSHILRC 146
H L C
Sbjct: 299 CCHGLNC 279
>TC17027 similar to UP|Q93XA5 (Q93XA5) Homeodomain leucine zipper protein
HDZ1 (Fragment), partial (34%)
Length = 854
Score = 27.3 bits (59), Expect = 9.4
Identities = 15/37 (40%), Positives = 21/37 (56%)
Frame = +1
Query: 578 IPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFK 614
IPG +++ LSYD KS +D GV+ S F + K
Sbjct: 4 IPGSDSKELSYDCCFKSSDD-GVDGTTTASLFAENLK 111
>TC18096 similar to UP|Q9SY59 (Q9SY59) F14N23.5, partial (5%)
Length = 870
Score = 27.3 bits (59), Expect = 9.4
Identities = 13/32 (40%), Positives = 19/32 (58%), Gaps = 3/32 (9%)
Frame = -2
Query: 386 NY*KKASLIIWD---ETPMLNKHCFEALDRSL 414
NY ASL+ W +TP L +HCF ++ + L
Sbjct: 563 NYLHVASLLHWSHLHQTPTLRQHCFLSVSQKL 468
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,285,448
Number of Sequences: 28460
Number of extensions: 192485
Number of successful extensions: 1752
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1739
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1749
length of query: 770
length of database: 4,897,600
effective HSP length: 97
effective length of query: 673
effective length of database: 2,136,980
effective search space: 1438187540
effective search space used: 1438187540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0178.5


