

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.13
(433 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
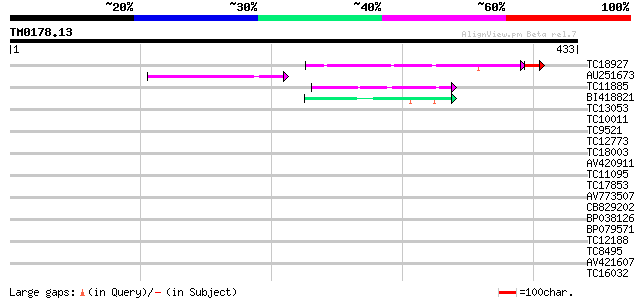
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 139 2e-36
AU251673 48 3e-06
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 46 1e-05
BI418821 42 2e-04
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 39 0.002
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 37 0.008
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 36 0.011
TC12773 33 0.092
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 32 0.27
AV420911 32 0.27
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 31 0.35
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 31 0.35
AV773507 31 0.45
CB829202 30 1.0
BP038126 29 1.3
BP079571 24 1.6
TC12188 similar to GB|AAF48767.2|22832753|AE003506 CG6474-PA {Dr... 28 2.3
TC8495 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-resp... 28 2.3
AV421607 28 2.3
TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%) 28 2.9
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 139 bits (351), Expect(2) = 2e-36
Identities = 74/178 (41%), Positives = 98/178 (54%), Gaps = 10/178 (5%)
Frame = -2
Query: 227 KTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSF 286
K K IEAIDN R + +PNQG+GG + GR D+ + +QKKP+ RPQ +GT+SG
Sbjct: 560 KAKSIEAIDNLRSR--PAFRPNQGSGGPNRSAPGRFDRNKSFQKKPFQRPQNRGTSSGYS 387
Query: 287 YPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKV 346
+ G PRP + C +C++KGH+A+ CP+ L+CWNC K GH +DC PKV
Sbjct: 386 HSFGN--FVPRPTQSDTSEIVCHRCSKKGHFANRCPD--LVCWNCQKTGHSGKDCTNPKV 219
Query: 347 EDAANVAGAR----------RPTAGGRVYFISGTEVDEDEGLIHGNREIPGNSLIVLF 394
E A N AR RP A RVY +SG E +GLI + L +LF
Sbjct: 218 EAATNAIAARRPAPAANKGKRPVASARVYTVSGAESHRADGLIRSVGSVNCKPLTILF 45
Score = 30.0 bits (66), Expect(2) = 2e-36
Identities = 11/15 (73%), Positives = 12/15 (79%)
Frame = -1
Query: 394 FKLWCYTFCYRLSLC 408
F LWCY F +RLSLC
Sbjct: 48 F*LWCYAFIHRLSLC 4
>AU251673
Length = 413
Score = 48.1 bits (113), Expect = 3e-06
Identities = 32/109 (29%), Positives = 51/109 (46%), Gaps = 1/109 (0%)
Frame = +2
Query: 106 VDYATYLLTGEAEYWWRGARAMMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLR 165
V+ A++ L G A W+ TW F F++++ P S R F RL
Sbjct: 23 VELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRDFERLE 202
Query: 166 QG-GMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAV 213
Q GMTV+EY++ L+++ Y + E +RF+ GL L ++V
Sbjct: 203 QAEGMTVSEYSAHFTHLSRYVPY---PLLEEERVKRFVRGLKEYLFKSV 340
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 46.2 bits (108), Expect = 1e-05
Identities = 33/113 (29%), Positives = 48/113 (42%), Gaps = 2/113 (1%)
Frame = +2
Query: 231 IEAIDNARGKFQSQNKP-NQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPT 289
+E + K S ++ ++ S M + R+D+ Y+ PY R +G + +
Sbjct: 176 VEV*KEEKRKMSSDSRSRSRSRSRSPMDRKIRSDRFS-YRDAPYRRDSRRGFSRDNLCK- 349
Query: 290 GGNAIAPRPLAVSLDGVT-CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
N P A V C C GH AS C L CWNC +PGH+A C
Sbjct: 350 --NCKRPGHFARECPNVAICHNCGLPGHIASECTTKSL-CWNCKEPGHMASSC 499
>BI418821
Length = 614
Score = 42.0 bits (97), Expect = 2e-04
Identities = 33/129 (25%), Positives = 41/129 (31%), Gaps = 13/129 (10%)
Frame = +2
Query: 226 EKTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGS 285
+KTK ++ Q K + G R + G G G G
Sbjct: 248 DKTKAVDVTGPNGAALQPTRKDSAPRGFGGWRGGERRNGG------------GGGGGGGG 391
Query: 286 FYPTGGNAIAPRPLAVSLD------GVTCFKCNQKGHYASHCPE-------GPLMCWNCN 332
Y G R S + G C+ C GH A C G C+NC
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGGGAGCYNCG 571
Query: 333 KPGHLARDC 341
GHLARDC
Sbjct: 572 DTGHLARDC 598
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/37 (51%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +3
Query: 306 VTCFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
V C C Q GH + C GPLM C NC GHLA +C
Sbjct: 3 VVCRNCQQLGHMSRDCM-GPLMICHNCGGRGHLAYEC 110
Score = 27.7 bits (60), Expect = 3.8
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +3
Query: 326 LMCWNCNKPGHLARDCRIP 344
++C NC + GH++RDC P
Sbjct: 3 VVCRNCQQLGHMSRDCMGP 59
Score = 26.6 bits (57), Expect = 8.6
Identities = 9/20 (45%), Positives = 10/20 (50%)
Frame = +3
Query: 308 CFKCNQKGHYASHCPEGPLM 327
C C +GH A CP G M
Sbjct: 69 CHNCGGRGHLAYECPSGRFM 128
Score = 26.6 bits (57), Expect = 8.6
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = +3
Query: 135 ITWECFRGAFLDKYFPNSAR 154
+ +EC G F+D+Y PN+ R
Sbjct: 96 LAYECPSGRFMDRYAPNTRR 155
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 36.6 bits (83), Expect = 0.008
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 2/37 (5%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPL--MCWNCNKPGHLARDCR 342
CF C GH+A C G C+ C GH+ R+C+
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCK 481
Score = 30.0 bits (66), Expect = 0.77
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +2
Query: 322 PEGPLMCWNCNKPGHLARDCR 342
P G C+NC GH ARDC+
Sbjct: 353 PPGSGRCFNCGLDGHWARDCK 415
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 36.2 bits (82), Expect = 0.011
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPL--MCWNCNKPGHLARDCR 342
CF C GH+A C G C+ C + GH+ ++C+
Sbjct: 391 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 501
Score = 29.6 bits (65), Expect = 1.0
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +1
Query: 322 PEGPLMCWNCNKPGHLARDCR 342
P G C+NC GH ARDC+
Sbjct: 373 PPGSGRCFNCGIDGHWARDCK 435
>TC12773
Length = 420
Score = 33.1 bits (74), Expect = 0.092
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = +2
Query: 328 CWNCNKPGHLARDCRI 343
CW C +PGH+A DC +
Sbjct: 347 CWKCQRPGHMAEDCLV 394
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 31.6 bits (70), Expect = 0.27
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 326 LMCWNCNKPGHLARDCR 342
L C+ C +PGH AR+CR
Sbjct: 160 LKCYECGEPGHFARECR 210
>AV420911
Length = 418
Score = 31.6 bits (70), Expect = 0.27
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +2
Query: 326 LMCWNCNKPGHLARDCR 342
L C+ C +PGH AR+CR
Sbjct: 368 LKCYECGEPGHFARECR 418
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 31.2 bits (69), Expect = 0.35
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = +1
Query: 326 LMCWNCNKPGHLARDCRI 343
+ C+ C +PGH AR+CR+
Sbjct: 373 MKCYECGEPGHFARECRM 426
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 31.2 bits (69), Expect = 0.35
Identities = 22/83 (26%), Positives = 31/83 (36%)
Frame = +2
Query: 240 KFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPL 299
K Q Q + G +GR+ R + R +G G + GS
Sbjct: 281 KAQPQGSSRDRDDGDRYRERGRDRDDRGDRD----RSRGYGGSRGS-------------- 406
Query: 300 AVSLDGVTCFKCNQKGHYASHCP 322
+G CFKC + GH+A CP
Sbjct: 407 ----NGGECFKCGKPGHFARECP 463
Score = 29.6 bits (65), Expect = 1.0
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = +2
Query: 328 CWNCNKPGHLARDC 341
C+ C KPGH AR+C
Sbjct: 419 CFKCGKPGHFAREC 460
>AV773507
Length = 496
Score = 30.8 bits (68), Expect = 0.45
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +2
Query: 314 KGHYASHCPEGPLMCWNCNKPGHLARDC 341
+G Y + G C+ C +PGH ARDC
Sbjct: 308 RGSYGAGDRVGQDDCFECGRPGHRARDC 391
Score = 26.6 bits (57), Expect = 8.6
Identities = 14/39 (35%), Positives = 17/39 (42%)
Frame = +2
Query: 284 GSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCP 322
G Y +GG V D CF+C + GH A CP
Sbjct: 284 GGGYSSGGRGSYGAGDRVGQDD--CFECGRPGHRARDCP 394
>CB829202
Length = 238
Score = 29.6 bits (65), Expect = 1.0
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = -2
Query: 389 SLIVLFKLWCYTFCYRLSLC 408
S ++LF L C+ FCY +SLC
Sbjct: 108 SPLMLFPLLCFRFCYLISLC 49
>BP038126
Length = 548
Score = 29.3 bits (64), Expect = 1.3
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +2
Query: 303 LDGVTCFKCNQKGHYASHCP 322
+ G TCFKC+Q+GH++ P
Sbjct: 272 MQGDTCFKCHQQGHWSWFWP 331
>BP079571
Length = 414
Score = 24.3 bits (51), Expect(2) = 1.6
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = -1
Query: 305 GVTCFKCNQKGHYASHCPEGPLMCW 329
G CF+ + GH A C G + W
Sbjct: 366 GGGCFRFGEVGHLARDCDGGVAVTW 292
Score = 23.1 bits (48), Expect(2) = 1.6
Identities = 7/13 (53%), Positives = 9/13 (68%)
Frame = -3
Query: 324 GPLMCWNCNKPGH 336
G C++C KPGH
Sbjct: 274 GETHCFHCGKPGH 236
>TC12188 similar to GB|AAF48767.2|22832753|AE003506 CG6474-PA {Drosophila
melanogaster;} , partial (6%)
Length = 297
Score = 28.5 bits (62), Expect = 2.3
Identities = 13/42 (30%), Positives = 22/42 (51%), Gaps = 1/42 (2%)
Frame = -2
Query: 244 QNKPNQGNGGSAMTNQGRNDKGRH-YQKKPYVRPQGQGTASG 284
+++P +GG +G + GRH Y ++ RP G+G G
Sbjct: 164 RSRPRDADGGLHYRERGASGNGRHRYHRRHRHRPPGRGCRRG 39
>TC8495 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-response
regulator 3), partial (9%)
Length = 783
Score = 28.5 bits (62), Expect = 2.3
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = -2
Query: 386 PGNSLIVLFKLWCYTFCYRLSLCDVVKTRC 415
P L+ LF LW T+ R + C +++ RC
Sbjct: 257 PSFRLVYLFLLWYQTYTVRRTNCPLIRGRC 168
>AV421607
Length = 245
Score = 28.5 bits (62), Expect = 2.3
Identities = 8/14 (57%), Positives = 11/14 (78%)
Frame = +3
Query: 328 CWNCNKPGHLARDC 341
C+ C +PGH +RDC
Sbjct: 54 CYKCGRPGHWSRDC 95
Score = 27.7 bits (60), Expect = 3.8
Identities = 14/45 (31%), Positives = 19/45 (42%)
Frame = +3
Query: 278 GQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCP 322
G G+ SGS TG C+KC + GH++ CP
Sbjct: 15 GPGSGSGSGTATG-----------------CYKCGRPGHWSRDCP 98
>TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%)
Length = 631
Score = 28.1 bits (61), Expect = 2.9
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +3
Query: 260 GRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRP 298
GR GR + Y +G+G + G +YP G N A +P
Sbjct: 216 GRGGGGR----RGYSNGRGRGGSRGGYYPNGRNQYADQP 320
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,321,689
Number of Sequences: 28460
Number of extensions: 100831
Number of successful extensions: 600
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 561
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 588
length of query: 433
length of database: 4,897,600
effective HSP length: 93
effective length of query: 340
effective length of database: 2,250,820
effective search space: 765278800
effective search space used: 765278800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0178.13


