

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0176a.2
(342 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
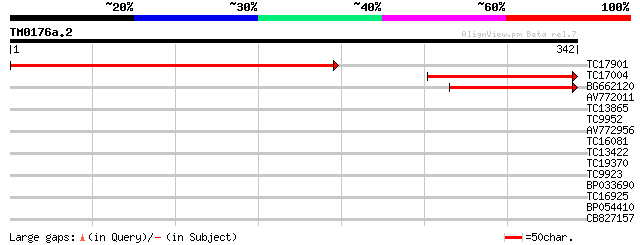
TC17901 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_150... 434 e-123
TC17004 similar to GB|AAM70808.1|21645108|AE003790 CG3271-PA {Dr... 192 5e-50
BG662120 124 2e-29
AV772011 39 0.002
TC13865 similar to UP|V714_ARATH (Q9FMR5) Vesicle-associated mem... 31 0.26
TC9952 29 1.0
AV772956 29 1.3
TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), par... 28 2.9
TC13422 27 3.8
TC19370 homologue to UP|CC21_MEDSA (P24923) Cell division contro... 27 5.0
TC9923 homologue to UP|Q7XHJ0 (Q7XHJ0) Formate dehydrogenase, pa... 27 6.5
BP033690 27 6.5
TC16925 weakly similar to UP|O22208 (O22208) bZIP family transcr... 26 8.5
BP054410 26 8.5
CB827157 26 8.5
>TC17901 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_150, partial
(52%)
Length = 763
Score = 434 bits (1117), Expect = e-123
Identities = 198/198 (100%), Positives = 198/198 (100%)
Frame = +2
Query: 1 MIGGYVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEH 60
MIGGYVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEH
Sbjct: 170 MIGGYVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEH 349
Query: 61 FDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQE 120
FDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQE
Sbjct: 350 FDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQE 529
Query: 121 AASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSA 180
AASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSA
Sbjct: 530 AASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSA 709
Query: 181 VFHSRDVDLTETLDYSSA 198
VFHSRDVDLTETLDYSSA
Sbjct: 710 VFHSRDVDLTETLDYSSA 763
>TC17004 similar to GB|AAM70808.1|21645108|AE003790 CG3271-PA {Drosophila
melanogaster;} , partial (9%)
Length = 543
Score = 192 bits (489), Expect = 5e-50
Identities = 90/90 (100%), Positives = 90/90 (100%)
Frame = +3
Query: 253 VCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWH 312
VCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWH
Sbjct: 3 VCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWH 182
Query: 313 ATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
ATTIPLTYIWWSFIRDDAEFRTSIRVKKAK
Sbjct: 183 ATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 272
>BG662120
Length = 458
Score = 124 bits (311), Expect = 2e-29
Identities = 51/77 (66%), Positives = 64/77 (82%)
Frame = +1
Query: 266 AVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWHATTIPLTYIWWSF 325
A+WAG+S HP+RWKLW VV+ LAM++E YDFPPY G +DAHA+W+A IPLT++WWS+
Sbjct: 16 AIWAGVSSHPARWKLWTVVVGQALAMVMETYDFPPYMGYVDAHAVWNACCIPLTFLWWSY 195
Query: 326 IRDDAEFRTSIRVKKAK 342
IRDDAEFRTS +KK K
Sbjct: 196 IRDDAEFRTSALLKKVK 246
>AV772011
Length = 447
Score = 38.5 bits (88), Expect = 0.002
Identities = 18/26 (69%), Positives = 22/26 (84%)
Frame = -3
Query: 186 DVDLTETLDYSSAVILLGYSLILAIL 211
DVD+ E LDYSSAV+ +GYSL L+IL
Sbjct: 295 DVDVAEKLDYSSAVVPVGYSLTLSIL 218
>TC13865 similar to UP|V714_ARATH (Q9FMR5) Vesicle-associated membrane
protein 714 (AtVAMP714), partial (34%)
Length = 543
Score = 31.2 bits (69), Expect = 0.26
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +2
Query: 136 HGWVSFFILLYYKLPLKDGKEAYYEY 161
H +++ F +LYYK L+ GK +Y+Y
Sbjct: 377 HVFIALFFILYYKSTLEPGKVLFYDY 454
>TC9952
Length = 630
Score = 29.3 bits (64), Expect = 1.0
Identities = 10/31 (32%), Positives = 20/31 (64%)
Frame = +1
Query: 230 IAFVITHVMYINFYKLDYGWNMIVCVVMAVV 260
+ F+ITH +I F+ L + W+ CV++ ++
Sbjct: 457 LEFIITHSSFITFFSLFFCWSDHGCVIIPIL 549
>AV772956
Length = 315
Score = 28.9 bits (63), Expect = 1.3
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = -2
Query: 232 FVITHVMYINFYKLDYGWNMIVCV 255
FVI +VMYI F+ L WN VC+
Sbjct: 158 FVIVYVMYILFFLLFLDWNGGVCL 87
>TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), partial (37%)
Length = 1189
Score = 27.7 bits (60), Expect = 2.9
Identities = 17/49 (34%), Positives = 22/49 (44%), Gaps = 14/49 (28%)
Frame = +3
Query: 176 WFWSAVFHSRDV--------------DLTETLDYSSAVILLGYSLILAI 210
W+W A F S V DL SAV+ LGYSL++A+
Sbjct: 501 WWWKAFFASGSVALYVFLYSVNYLVFDLQSLSGPVSAVLYLGYSLLMAV 647
>TC13422
Length = 490
Score = 27.3 bits (59), Expect = 3.8
Identities = 8/20 (40%), Positives = 14/20 (70%)
Frame = +2
Query: 164 LWHIYGLLSLNSWFWSAVFH 183
LWH +G + +N WF A+++
Sbjct: 248 LWHTFGSIFVNEWFDRAIYN 307
>TC19370 homologue to UP|CC21_MEDSA (P24923) Cell division control protein 2
homolog 1 (Fragment) , partial (34%)
Length = 537
Score = 26.9 bits (58), Expect = 5.0
Identities = 17/47 (36%), Positives = 22/47 (46%), Gaps = 3/47 (6%)
Frame = +3
Query: 272 SGHPSRWKLWLVVIAGGLAMLLEIYDF---PPYEGLLDAHALWHATT 315
SGHP W LW ++ L ++ + F PP E L ALW T
Sbjct: 144 SGHPRTWHLWFQILI-QLVLIFFLVCFAWIPPKESL--PGALWSMNT 275
>TC9923 homologue to UP|Q7XHJ0 (Q7XHJ0) Formate dehydrogenase, partial
(56%)
Length = 755
Score = 26.6 bits (57), Expect = 6.5
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 3/36 (8%)
Frame = +2
Query: 48 CFPHCKFPSDGEHFDRPWYMQ---EPLYLQWKKWDC 80
CF C FPS + +P++ EP L WK+ DC
Sbjct: 59 CFLCCSFPSHCSN-TKPFFFHFL*EPSCLWWKEEDC 163
>BP033690
Length = 546
Score = 26.6 bits (57), Expect = 6.5
Identities = 11/36 (30%), Positives = 17/36 (46%)
Frame = +1
Query: 19 VIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKF 54
V+ ++ V+ R RD + QC +PHC F
Sbjct: 337 VV*GASWSVEYRSRDAVFQCGSQSSQFSYAYPHCSF 444
>TC16925 weakly similar to UP|O22208 (O22208) bZIP family transcription
factor, partial (19%)
Length = 642
Score = 26.2 bits (56), Expect = 8.5
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -3
Query: 54 FPSDGEHFDRPWYMQEPL 71
FPS E F+ PW+ Q PL
Sbjct: 352 FPSWPEQFEMPWFSQWPL 299
>BP054410
Length = 572
Score = 26.2 bits (56), Expect = 8.5
Identities = 12/36 (33%), Positives = 24/36 (66%)
Frame = +3
Query: 173 LNSWFWSAVFHSRDVDLTETLDYSSAVILLGYSLIL 208
++S+F+ +FH LTET + ++ +L+ Y+L+L
Sbjct: 267 VSSYFYMELFHGCINHLTETSEGATDKLLVQYNLLL 374
>CB827157
Length = 536
Score = 26.2 bits (56), Expect = 8.5
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +1
Query: 242 FYKLDYGWNMIVCVVMAVVQLVIWAVWAGL 271
FY L+ W I+ V + +L W +W GL
Sbjct: 151 FYLLNQQWRQILRVPRRLGKLRGWVIWLGL 240
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,310,619
Number of Sequences: 28460
Number of extensions: 148779
Number of successful extensions: 976
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 968
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 976
length of query: 342
length of database: 4,897,600
effective HSP length: 91
effective length of query: 251
effective length of database: 2,307,740
effective search space: 579242740
effective search space used: 579242740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0176a.2


