

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.10
(265 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
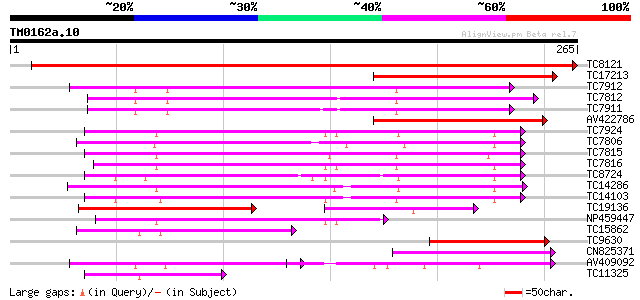
Sequences producing significant alignments: (bits) Value
TC8121 UP|Q9LKJ5 (Q9LKJ5) Multifunctional transport intrinsic me... 374 e-104
TC17213 similar to UP|AAS48064 (AAS48064) NIP2, partial (30%) 128 1e-30
TC7912 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic p... 107 2e-24
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 105 1e-23
TC7911 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic p... 101 1e-22
AV422786 100 3e-22
TC7924 homologue to UP|CAE53883 (CAE53883) Aquaporin, partial (95%) 96 6e-21
TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1,... 96 8e-21
TC7815 similar to UP|Q9AVB5 (Q9AVB5) Plasma membrane intrinsic p... 94 2e-20
TC7816 homologue to UP|Q9ATM4 (Q9ATM4) Plasma membrane integral ... 93 4e-20
TC8724 homologue to UP|Q9AQU5 (Q9AQU5) Plasma membrane integral ... 93 5e-20
TC14286 homologue to UP|O65357 (O65357) Aquaporin 2, complete 92 1e-19
TC14103 similar to UP|Q84XC6 (Q84XC6) Plasma intrinsic protein 2... 89 8e-19
TC19136 similar to UP|AAQ11827 (AAQ11827) Nodulin-like intrinsic... 88 1e-18
NP459447 plasma membrane intrinsic protein homolog [Lotus japoni... 70 5e-13
TC15862 homologue to UP|O22339 (O22339) Aquaporin-like transmemb... 66 7e-12
TC9630 similar to UP|AAS48063 (AAS48063) NIP3, partial (18%) 59 6e-10
CN825371 52 1e-07
AV409092 50 4e-07
TC11325 similar to UP|Q8LNX5 (Q8LNX5) Plasma membrane intrinsic ... 43 5e-05
>TC8121 UP|Q9LKJ5 (Q9LKJ5) Multifunctional transport intrinsic membrane
protein 2, complete
Length = 1213
Score = 374 bits (959), Expect = e-104
Identities = 177/255 (69%), Positives = 219/255 (85%)
Frame = +2
Query: 11 HDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTL 70
H+VV++VNKDASKTIE SDT +V F+QK++AE VGT+F IF GCASIVVNKNNDNVVTL
Sbjct: 131 HEVVVNVNKDASKTIEVSDTNFTVSFLQKVIAELVGTYFFIFAGCASIVVNKNNDNVVTL 310
Query: 71 PGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLAS 130
PGIALVWGL +MVL+YS+GHISGAHFNPA T AFA+TKRFPW QV Y+++Q+LG+ LAS
Sbjct: 311 PGIALVWGLAVMVLVYSLGHISGAHFNPAATIAFASTKRFPWKQVPAYVSAQVLGSTLAS 490
Query: 131 GILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIG 190
G L+++FSG H+QF+G +P+G+NLQAFVIEFI TF L+F++ VATD+RAIGE+AGI +G
Sbjct: 491 GTLRLIFSGKHNQFAGALPTGSNLQAFVIEFIITFFLIFILFGVATDDRAIGEVAGIVVG 670
Query: 191 STLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYT 250
ST+LLN+L +GPITGASMNPAR++G A H++YR I +Y +S GAVAGAWV+NI+RYT
Sbjct: 671 STVLLNVLFAGPITGASMNPARSIGSAFVHNEYRGIWIYLLSPTLGAVAGAWVYNIVRYT 850
Query: 251 DKPLHEITKGSSILK 265
DKPL EITK S LK
Sbjct: 851 DKPLREITKNVSFLK 895
>TC17213 similar to UP|AAS48064 (AAS48064) NIP2, partial (30%)
Length = 578
Score = 128 bits (321), Expect = 1e-30
Identities = 58/86 (67%), Positives = 74/86 (85%)
Frame = +3
Query: 171 ISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYF 230
IS V+TDNRAIGEMAG+AIG+T+LLN++I+GPITGASMNPAR+LGPAI HS+Y+ I +Y
Sbjct: 3 ISGVSTDNRAIGEMAGLAIGATILLNVIIAGPITGASMNPARSLGPAIVHSEYKGIWIYL 182
Query: 231 VSTIFGAVAGAWVFNILRYTDKPLHE 256
VS + GAVAG W +N +RYT+ LH+
Sbjct: 183 VSPVLGAVAGTWSYNFIRYTNYTLHQ 260
>TC7912 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
partial (95%)
Length = 1045
Score = 107 bits (267), Expect = 2e-24
Identities = 70/215 (32%), Positives = 105/215 (48%), Gaps = 7/215 (3%)
Frame = +2
Query: 29 DTYTSVFFVQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMV 83
D S+ ++ +AEF+ T +F G S I K + P IA+ L V
Sbjct: 56 DDSFSLGSIKAYIAEFISTLLFVFAGVGSAIAYGKVTSDAALDPAGLVAIAVCHAFALFV 235
Query: 84 LIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQ 143
+ +ISG H NPAVTF A + + Y +QLLG+++A +L + G
Sbjct: 236 AVSVGANISGGHVNPAVTFGLALGGQITILTGIFYWIAQLLGSIVACFLLHYVTGGLETP 415
Query: 144 FSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISG 201
G + V E I TF L++ + A A D + ++G +A IAIG + NIL +G
Sbjct: 416 VHGLAAGVGAFEGVVTEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAG 595
Query: 202 PITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFG 236
P +G SMNPAR+ GPA+ + I +Y+V + G
Sbjct: 596 PFSGGSMNPARSFGPAVVSGNFHDIWIYWVGPLIG 700
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 105 bits (261), Expect = 1e-23
Identities = 68/218 (31%), Positives = 109/218 (49%), Gaps = 7/218 (3%)
Frame = +3
Query: 37 VQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHI 91
++ +AEF+ TF +F G S I NK D+ P ++ L V + +I
Sbjct: 132 LRSALAEFISTFIFVFAGSGSGIAYNKLTDDGAATPAGLISASIAHAFALFVAVSISANI 311
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSG 151
SG H NPAVTF +++ YI +QLLG+++AS +L + F + G
Sbjct: 312 SGGHVNPAVTFGLFVGGNISFLRGVIYIIAQLLGSIVASLLLAFVTGLAVPAFGLSAGVG 491
Query: 152 TNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITGASMN 209
A V+E + TF L++ + A A D + ++G +A IAIG + NIL+ G GASMN
Sbjct: 492 VG-NALVLEIVMTFGLVYTVYATAVDPKKGSLGTIAPIAIGFIVGANILLGGAFDGASMN 668
Query: 210 PARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
PA + GPA+ + +Y+V + G ++ ++
Sbjct: 669 PAVSFGPAVVSWSWSNHWIYWVGPLVGGGIAGLIYEVV 782
>TC7911 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
complete
Length = 1018
Score = 101 bits (252), Expect = 1e-22
Identities = 67/207 (32%), Positives = 105/207 (50%), Gaps = 7/207 (3%)
Frame = +1
Query: 37 VQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHI 91
++ +AEF+ T +F G S I K + P +A+ L V + +I
Sbjct: 109 IKAYIAEFISTLIFVFAGVGSAIAYGKLTSDAALDPAGLVAVAICHAFALFVAVSVGANI 288
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSG 151
SG H NPAVTF A + + Y +QLLG+++A +LK + G ++ +G
Sbjct: 289 SGGHVNPAVTFGLALGGQITLLTGLFYWIAQLLGSIVACFLLKFVTGGLTLPIH-SVAAG 465
Query: 152 TNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITGASMN 209
+ V E I TF L++ + A A D + ++G +A IAIG + NIL +GP +G SMN
Sbjct: 466 AG-EGVVTEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAGPFSGGSMN 642
Query: 210 PARTLGPAIFHSKYRAIVVYFVSTIFG 236
PAR+ GPA+ + +Y+V + G
Sbjct: 643 PARSFGPAVVSGDFHDNWIYWVGPLIG 723
>AV422786
Length = 427
Score = 100 bits (248), Expect = 3e-22
Identities = 46/81 (56%), Positives = 64/81 (78%)
Frame = -3
Query: 171 ISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYF 230
++AVATD+RA+GE+AGIA+G+T+LLNILISGP +G SMNP RTLGPA+ Y+ I +Y
Sbjct: 425 VTAVATDSRAVGELAGIAVGATVLLNILISGPTSGGSMNPVRTLGPAVAAGNYKHIWIYL 246
Query: 231 VSTIFGAVAGAWVFNILRYTD 251
V+ GA+AGA V+ +++ D
Sbjct: 245 VAPTLGALAGAGVYTLVKLRD 183
>TC7924 homologue to UP|CAE53883 (CAE53883) Aquaporin, partial (95%)
Length = 1078
Score = 95.9 bits (237), Expect = 6e-21
Identities = 63/221 (28%), Positives = 110/221 (49%), Gaps = 15/221 (6%)
Frame = +2
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHIS 92
F + L+AEF+ T ++ A+++ +K V L GIA +G ++ VL+Y IS
Sbjct: 119 FYRALIAEFIATLLFLYVTVATVIGHKKQTGPCDGVGLLGIAWSFGGMIFVLVYCTAGIS 298
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ I+ Y+ +Q LGA+ G++K ++ G ++
Sbjct: 299 GGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGLVKAFMKHPYNSLGGGANSVA 478
Query: 150 SG-TNLQAFVIEFITTFLLMFVISAVATDNRA-----IGEMAGIAIGSTLLLNILISGPI 203
G + A E I TF+L++ + + R+ + +A + IG + + L + PI
Sbjct: 479 HGYSKGSALGAEIIGTFVLVYTVFSATDPKRSARDSHVPVLAPLPIGFAVFMVHLATIPI 658
Query: 204 TGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
TG +NPAR+ G A+ + + +++V GA+A A
Sbjct: 659 TGTGINPARSFGAAVIFNNGKVWDDQWIFWVGPFVGALAAA 781
>TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1, complete
Length = 1413
Score = 95.5 bits (236), Expect = 8e-21
Identities = 69/229 (30%), Positives = 109/229 (47%), Gaps = 19/229 (8%)
Frame = +3
Query: 32 TSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDN---VVTLPGIALVWGLVLMVLIYSV 88
TS F + +AEFV TF ++ +++ D+ V + GIA +G ++ L+Y
Sbjct: 213 TSWSFYRAGIAEFVATFLFLYITVLTVMGVSGADSKCKTVGIQGIAWAFGGMIFALVYCT 392
Query: 89 GHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTI 148
ISG H NPAVTF ++ + YI Q LGA++ +G++K T GT+
Sbjct: 393 AGISGGHINPAVTFGLFLARKLSLTRAIFYIVMQCLGAIVGAGVVKGFEGKTK---YGTL 563
Query: 149 PSGTNLQA--------FVIEFITTFLLMFVISAVATDNRAIGE-----MAGIAIGSTLLL 195
G N A E + TF+L++ + + R+ + +A + IG + L
Sbjct: 564 GGGANFVAPGYTKGDGLGAEIVGTFVLVYTVFSATDAKRSARDSHVPILAPLPIGFAVFL 743
Query: 196 NILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
L + PITG +NPAR+LG AI +K R +++V GA A
Sbjct: 744 VHLATIPITGTGINPARSLGAAIIFNKDRGWDDHWIFWVGPFIGAALAA 890
>TC7815 similar to UP|Q9AVB5 (Q9AVB5) Plasma membrane intrinsic protein
2-1, complete
Length = 1450
Score = 94.0 bits (232), Expect = 2e-20
Identities = 59/223 (26%), Positives = 110/223 (48%), Gaps = 17/223 (7%)
Frame = +1
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV-----VTLPGIALVWGLVLMVLIYSVGH 90
F + L+AEF+ T ++ +++ + ++ + GIA +G ++ +L+Y
Sbjct: 172 FYRALIAEFIATLLFLYVTVLTVIGHSRLNSADPCDGAGILGIAWAFGGMIFILVYCTAG 351
Query: 91 ISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTI-- 148
ISG H NPAVTF ++ I+ YI +Q LGA+ G+ K ++++ G
Sbjct: 352 ISGGHINPAVTFGLFIGRKVSLIRALLYIVAQCLGAISGVGLAKAFQKSFYNRYGGAANF 531
Query: 149 --PSGTNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAGIAIGSTLLLNILISG 201
P +N A E I TF+L++ + + R + +A + IG + + L +
Sbjct: 532 VQPGYSNGTALGAEIIGTFVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATI 711
Query: 202 PITGASMNPARTLGPAIFHSK---YRAIVVYFVSTIFGAVAGA 241
P+TG +NPAR+ G A+ +++ + +++V GA+A A
Sbjct: 712 PVTGTGINPARSFGAAVIYNEDKIWDDHWIFWVGPFIGALAAA 840
>TC7816 homologue to UP|Q9ATM4 (Q9ATM4) Plasma membrane integral protein
ZmPIP2-7, complete
Length = 1523
Score = 93.2 bits (230), Expect = 4e-20
Identities = 64/224 (28%), Positives = 110/224 (48%), Gaps = 22/224 (9%)
Frame = +1
Query: 40 LVAEFVGTFFLIFTGCASIVVNKNNDNV----------VTLPGIALVWGLVLMVLIYSVG 89
++AEFV T ++ +I+ K ++ V + GIA +G ++ VL+Y
Sbjct: 295 VIAEFVATLLFLYITVLTIIGYKRQTDLGIPGNTECDGVGILGIAWAFGGMIFVLVYCTA 474
Query: 90 HISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG--- 146
ISG H NPAVTF ++ ++ YI +Q LGA+ +G+ K ++++ G
Sbjct: 475 GISGGHINPAVTFGLFVGRKVSLVRALLYIIAQCLGAICGAGLAKGFQKSFYNRYGGGAN 654
Query: 147 TIPSG-TNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAGIAIGSTLLLNILIS 200
TI G + A E I TF+L++ + + R + +A + IG + + L +
Sbjct: 655 TISDGYSKGTALGAEIIGTFVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLAT 834
Query: 201 GPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
P+TG +NPAR+ GPA+ ++ + VY+V GA+ A
Sbjct: 835 IPVTGTGINPARSFGPAVIFNEDKVWDDQWVYWVGPFVGALVAA 966
>TC8724 homologue to UP|Q9AQU5 (Q9AQU5) Plasma membrane integral protein
ZmPIP1-4 (Plasma membrane integral protein ZmPIP1-3),
partial (93%)
Length = 1560
Score = 92.8 bits (229), Expect = 5e-20
Identities = 73/224 (32%), Positives = 111/224 (48%), Gaps = 18/224 (8%)
Frame = +1
Query: 36 FVQKLVAEFVGTF-FLIFTGCASIVVNK--NNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
F + +AEFV TF FL T + VN+ N + V + GIA +G ++ L+Y IS
Sbjct: 235 FYRAGIAEFVATFLFLYITVLTVMGVNRSPNKCSSVGIQGIAWAFGGMIFALVYCTAGIS 414
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGT--HDQFSG---- 146
G H NPAVTF ++ + YI Q LGA+ +G++K F G + F G
Sbjct: 415 GGHINPAVTFGLFLARKLSLTRALFYIVMQCLGAICGAGVVKG-FEGNARFEMFKGGANV 591
Query: 147 TIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA------IGEMAGIAIGSTLLLNILIS 200
P T E + TF+L++ + + ATD + + +A + IG + L L +
Sbjct: 592 VNPGYTKGDGLGAEIVGTFVLVYTVFS-ATDAKRNARDSHVPLLAPLPIGFAVFLVHLAT 768
Query: 201 GPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
PITG +NPAR+LG AI +++ A +++V GA A
Sbjct: 769 IPITGTGINPARSLGAAIIYNRDHAWDDHWIFWVGPFIGAALAA 900
Score = 37.0 bits (84), Expect = 0.003
Identities = 28/109 (25%), Positives = 53/109 (47%), Gaps = 11/109 (10%)
Frame = +1
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFV-ISAVATDNRAIGEMAGI-------AIGSTLLLNI 197
G + S + +A + EF+ TFL +++ + V NR+ + + + A G + +
Sbjct: 214 GELKSWSFYRAGIAEFVATFLFLYITVLTVMGVNRSPNKCSSVGIQGIAWAFGGMIFALV 393
Query: 198 LISGPITGASMNPARTLGPAIFHSKYRAI---VVYFVSTIFGAVAGAWV 243
+ I+G +NPA T G +F ++ ++ + Y V GA+ GA V
Sbjct: 394 YCTAGISGGHINPAVTFG--LFLARKLSLTRALFYIVMQCLGAICGAGV 534
>TC14286 homologue to UP|O65357 (O65357) Aquaporin 2, complete
Length = 1278
Score = 91.7 bits (226), Expect = 1e-19
Identities = 62/239 (25%), Positives = 115/239 (47%), Gaps = 24/239 (10%)
Frame = +1
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---------VTLPGIALVWG 78
++ T F + ++AEF+ T ++ +++ K +V V + GIA +G
Sbjct: 190 AEELTKWSFYRAVIAEFIATLLFLYITVLTVIGYKVQSDVKNGGDDCGGVGILGIAWAFG 369
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFS 138
++ +L+Y ISG H NPAVTF ++ I+ Y+ +Q LGA+ G++K
Sbjct: 370 GMIFILVYCTAGISGGHINPAVTFGLFLARKVSLIRAILYMVAQCLGAICGVGLVKAFQK 549
Query: 139 GTHDQFSGTIPS-------GTNLQAFVIEFITTFLLMFVISAVATDNRA-----IGEMAG 186
++++ G S GT L A E I TF+L++ + + R+ + +A
Sbjct: 550 SHYNKYGGGANSLNDGYSTGTGLGA---EIIGTFVLVYTVFSATDPKRSARDSHVPVLAP 720
Query: 187 IAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGAW 242
+ IG + + L + P+TG +NPAR+ G A+ ++ + +++V GA A+
Sbjct: 721 LPIGFAVFMVHLATIPVTGTGINPARSFGAAVIFNQSKPWDDHWIFWVGPFIGAAIAAF 897
>TC14103 similar to UP|Q84XC6 (Q84XC6) Plasma intrinsic protein 2,2, partial
(94%)
Length = 1199
Score = 89.0 bits (219), Expect = 8e-19
Identities = 66/230 (28%), Positives = 108/230 (46%), Gaps = 24/230 (10%)
Frame = +3
Query: 36 FVQKLVAEFVGTF-FLIFTGCASIVVNKNNDNVVT--------LPGIALVWGLVLMVLIY 86
F + L+AEFV T FL T I + D T + GIA +G ++ +L+Y
Sbjct: 198 FYRALIAEFVATLLFLYITVLTVIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVY 377
Query: 87 SVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG 146
ISG H NPAVTF ++ ++ Y+ +Q LGA+ G++K ++++ G
Sbjct: 378 CTAGISGGHINPAVTFGLFLARKVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHG 557
Query: 147 -------TIPSGTNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAGIAIGSTLL 194
GT L A E I TF+L++ + + R + +A + IG +
Sbjct: 558 GANMLADGYSKGTGLGA---EIIGTFVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVF 728
Query: 195 LNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
+ L + PITG +NPAR+ G A+ ++ +A +++V GA A
Sbjct: 729 MVHLATIPITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPFIGATIAA 878
>TC19136 similar to UP|AAQ11827 (AAQ11827) Nodulin-like intrinsic protein
NIP1-2 (Fragment), partial (29%)
Length = 556
Score = 88.2 bits (217), Expect = 1e-18
Identities = 44/83 (53%), Positives = 53/83 (63%)
Frame = +2
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV QK++AEFVGTF LIF A +VN D L G A GL +M +I S+GHIS
Sbjct: 305 SVPLTQKILAEFVGTFILIFAATAGPIVNNKYDGAEGLMGNAATAGLTVMFIILSIGHIS 484
Query: 93 GAHFNPAVTFAFATTKRFPWIQV 115
GAH NP++T AFA + FPW QV
Sbjct: 485 GAHLNPSLTIAFAAFRHFPWSQV 553
Score = 39.3 bits (90), Expect = 7e-04
Identities = 26/77 (33%), Positives = 38/77 (48%), Gaps = 5/77 (6%)
Frame = +2
Query: 148 IPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGI-----AIGSTLLLNILISGP 202
IPS Q + EF+ TF+L+F +A N G+ G T++ IL G
Sbjct: 299 IPSVPLTQKILAEFVGTFILIFAATAGPIVNNKYDGAEGLMGNAATAGLTVMFIILSIGH 478
Query: 203 ITGASMNPARTLGPAIF 219
I+GA +NP+ T+ A F
Sbjct: 479 ISGAHLNPSLTIAFAAF 529
>NP459447 plasma membrane intrinsic protein homolog [Lotus japonicus]
Length = 578
Score = 69.7 bits (169), Expect = 5e-13
Identities = 45/144 (31%), Positives = 72/144 (49%), Gaps = 7/144 (4%)
Frame = +1
Query: 41 VAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHISGAHFN 97
+AEF+ TF ++ +++ K + N+ V + GIA +G ++ L+Y ISG H N
Sbjct: 73 IAEFIATFLFLYITVLTVMGVKRSPNMCASVGIQGIAWAFGGMIFALVYCTAGISGGHIN 252
Query: 98 PAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIPSG-TN 153
PAVTF ++ + YI Q LGA+ +G++K + G TI G T
Sbjct: 253 PAVTFGLFLARKLSLTRAVYYIVMQCLGAICGAGVVKGFQPKQYQALGGGANTIAHGYTK 432
Query: 154 LQAFVIEFITTFLLMFVISAVATD 177
E I TF+L++ + + ATD
Sbjct: 433 GSGLGAEIIGTFVLVYTVFS-ATD 501
Score = 36.2 bits (82), Expect = 0.006
Identities = 37/138 (26%), Positives = 61/138 (43%), Gaps = 20/138 (14%)
Frame = +1
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFV-ISAVATDNRAIGEMAGI-------AIGSTLLLNI 197
G + S + +A + EFI TFL +++ + V R+ A + A G + +
Sbjct: 37 GELASWSFWRAGIAEFIATFLFLYITVLTVMGVKRSPNMCASVGIQGIAWAFGGMIFALV 216
Query: 198 LISGPITGASMNPARTLGPAIFHSKYRAI---VVYFVSTIFGAVAGAWV---FNILRY-- 249
+ I+G +NPA T G +F ++ ++ V Y V GA+ GA V F +Y
Sbjct: 217 YCTAGISGGHINPAVTFG--LFLARKLSLTRAVYYIVMQCLGAICGAGVVKGFQPKQYQA 390
Query: 250 ----TDKPLHEITKGSSI 263
+ H TKGS +
Sbjct: 391 LGGGANTIAHGYTKGSGL 444
>TC15862 homologue to UP|O22339 (O22339) Aquaporin-like transmembrane
channel protein, partial (61%)
Length = 576
Score = 65.9 bits (159), Expect = 7e-12
Identities = 36/106 (33%), Positives = 59/106 (54%), Gaps = 3/106 (2%)
Frame = +3
Query: 32 TSVFFVQKLVAEFVGTFFLIFTGCASIV-VNKNNDNVVT--LPGIALVWGLVLMVLIYSV 88
TS F + +AEFV TF ++ +++ VNK++ T + GIA +G ++ L+Y
Sbjct: 189 TSWSFYRAGIAEFVATFLFLYITILTVMGVNKSDTKCKTVGIQGIAWAFGGMIFALVYCT 368
Query: 89 GHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILK 134
ISG H NPAVTF ++ + Y+ Q+LGA+ +G++K
Sbjct: 369 AGISGGHINPAVTFGLFLARKLSLTRAVFYMVMQVLGAICGAGVVK 506
Score = 36.2 bits (82), Expect = 0.006
Identities = 28/101 (27%), Positives = 51/101 (49%), Gaps = 12/101 (11%)
Frame = +3
Query: 155 QAFVIEFITTFLLMFV-------ISAVATDNRAIGEMAGI--AIGSTLLLNILISGPITG 205
+A + EF+ TFL +++ ++ T + +G + GI A G + + + I+G
Sbjct: 207 RAGIAEFVATFLFLYITILTVMGVNKSDTKCKTVG-IQGIAWAFGGMIFALVYCTAGISG 383
Query: 206 ASMNPARTLGPAIFHSKYRAI---VVYFVSTIFGAVAGAWV 243
+NPA T G +F ++ ++ V Y V + GA+ GA V
Sbjct: 384 GHINPAVTFG--LFLARKLSLTRAVFYMVMQVLGAICGAGV 500
>TC9630 similar to UP|AAS48063 (AAS48063) NIP3, partial (18%)
Length = 971
Score = 59.3 bits (142), Expect = 6e-10
Identities = 28/56 (50%), Positives = 39/56 (69%)
Frame = +3
Query: 197 ILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDK 252
IL P TGASMNP RTLGPAI + Y+AI VY V+ + GA++GA ++ ++ D+
Sbjct: 177 ILYCRPETGASMNPVRTLGPAIAANNYKAIWVYLVAPVLGALSGAGIYTAVKLPDE 344
>CN825371
Length = 483
Score = 51.6 bits (122), Expect = 1e-07
Identities = 26/76 (34%), Positives = 41/76 (53%)
Frame = +1
Query: 180 AIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVA 239
+IG +A +AIG + NIL+ GP GA MNPA GP++ ++ V+++ GA
Sbjct: 13 SIGAIAPLAIGLIVGANILVGGPFDGACMNPALAFGPSLVGWRWHYHWVFWLGPFLGAAL 192
Query: 240 GAWVFNILRYTDKPLH 255
A ++ L +P H
Sbjct: 193 AALIYEYLVIPTEPSH 240
>AV409092
Length = 426
Score = 50.1 bits (118), Expect = 4e-07
Identities = 36/115 (31%), Positives = 52/115 (44%), Gaps = 5/115 (4%)
Frame = +3
Query: 29 DTYTSVFFVQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLP----GIALVWGLVLMV 83
D SV ++ +AEF+ T IF G S I N+ + P +AL L V
Sbjct: 57 DDSFSVTSLKAYLAEFIATLLFIFAGVGSAIAYNELTSDAALDPTGLVAVALAHAFALFV 236
Query: 84 LIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFS 138
+ +ISG H NPAVT A + Y +QLLG+++A +L + S
Sbjct: 237 GVSIAANISGGHLNPAVTLGLAVGGNITLLTGFFYWIAQLLGSIVACLLLNFITS 401
Score = 45.1 bits (105), Expect = 1e-05
Identities = 38/137 (27%), Positives = 66/137 (47%), Gaps = 11/137 (8%)
Frame = +3
Query: 130 SGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMF--VISAVA----TDNRAIGE 183
S ++K+ F D FS T +L+A++ EFI T L +F V SA+A T + A+
Sbjct: 24 SKMVKIAFGRFDDSFSVT-----SLKAYLAEFIATLLFIFAGVGSAIAYNELTSDAALDP 188
Query: 184 MAGIAIGST----LLLNILISGPITGASMNPARTLGPAI-FHSKYRAIVVYFVSTIFGAV 238
+A+ L + + I+ I+G +NPA TLG A+ + Y+++ + G++
Sbjct: 189 TGLVAVALAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITLLTGFFYWIAQLLGSI 368
Query: 239 AGAWVFNILRYTDKPLH 255
+ N + P H
Sbjct: 369 VACLLLNFITSKSIPTH 419
>TC11325 similar to UP|Q8LNX5 (Q8LNX5) Plasma membrane intrinsic protein,
partial (41%)
Length = 427
Score = 43.1 bits (100), Expect = 5e-05
Identities = 23/69 (33%), Positives = 38/69 (54%), Gaps = 3/69 (4%)
Frame = +1
Query: 36 FVQKLVAEFVGTFFLIFTGCASIV---VNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
F + +AEFV TF ++ +++ +K+ + V + GIA +G ++ L+Y IS
Sbjct: 208 FYRAGIAEFVATFLFLYITVLTVMGVAKSKSKCSTVGVQGIAWSFGGMIFALVYCTAGIS 387
Query: 93 GAHFNPAVT 101
G H NPA T
Sbjct: 388 GGHINPAXT 414
Score = 31.6 bits (70), Expect = 0.14
Identities = 22/78 (28%), Positives = 37/78 (47%), Gaps = 8/78 (10%)
Frame = +1
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFV----ISAVATDNRAIGEMA--GIA--IGSTLLLNI 197
G + S + +A + EF+ TFL +++ + VA + GIA G + +
Sbjct: 187 GELSSWSFYRAGIAEFVATFLFLYITVLTVMGVAKSKSKCSTVGVQGIAWSFGGMIFALV 366
Query: 198 LISGPITGASMNPARTLG 215
+ I+G +NPA TLG
Sbjct: 367 YCTAGISGGHINPAXTLG 420
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,834,261
Number of Sequences: 28460
Number of extensions: 44534
Number of successful extensions: 326
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 302
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 311
length of query: 265
length of database: 4,897,600
effective HSP length: 89
effective length of query: 176
effective length of database: 2,364,660
effective search space: 416180160
effective search space used: 416180160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0162a.10


