

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.1
(118 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
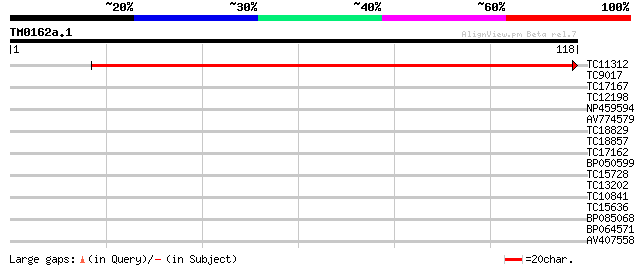
Score E
Sequences producing significant alignments: (bits) Value
TC11312 weakly similar to UP|O04147 (O04147) Cyclic phosphodiest... 211 3e-56
TC9017 weakly similar to GB|BAC20772.1|23617089|AP004664 valine-... 28 0.52
TC17167 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived f... 26 1.5
TC12198 similar to UP|Q9LHU3 (Q9LHU3) ESTs D40069(S1808), partia... 26 1.5
NP459594 Krm protein [Lotus japonicus] 26 1.5
AV774579 26 2.0
TC18829 homologue to UP|Q8L4X4 (Q8L4X4) Amino acid permease-like... 25 2.6
TC18857 homologue to UP|Q9ZS21 (Q9ZS21) Glyoxalase I , complete 24 7.5
TC17162 similar to UP|Q8H5F1 (Q8H5F1) OJ1165_F02.16 protein, par... 24 7.5
BP050599 24 7.5
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 23 9.8
TC13202 23 9.8
TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1,... 23 9.8
TC15636 similar to UP|Q93WR7 (Q93WR7) MAP kinase kinase, partial... 23 9.8
BP085068 23 9.8
BP064571 23 9.8
AV407558 23 9.8
>TC11312 weakly similar to UP|O04147 (O04147) Cyclic phosphodiesterase
(Protein kinase-like protein) , partial (84%)
Length = 621
Score = 211 bits (536), Expect = 3e-56
Identities = 101/101 (100%), Positives = 101/101 (100%)
Frame = +1
Query: 18 AIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQV 77
AIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQV
Sbjct: 16 AIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQV 195
Query: 78 TVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS 118
TVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS
Sbjct: 196 TVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS 318
>TC9017 weakly similar to GB|BAC20772.1|23617089|AP004664 valine--tRNA
ligase-like protein {Oryza sativa (japonica
cultivar-group);} , partial (7%)
Length = 686
Score = 27.7 bits (60), Expect = 0.52
Identities = 13/25 (52%), Positives = 14/25 (56%)
Frame = -3
Query: 87 FFYQCVYLLLHPSPPVVETSAHCSN 111
F YQCV LLLH ET + C N
Sbjct: 78 FSYQCVILLLHLGKAF*ETLSFCRN 4
>TC17167 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived factor
2-like protein precursor (SDF2-like protein), partial
(47%)
Length = 588
Score = 26.2 bits (56), Expect = 1.5
Identities = 11/18 (61%), Positives = 11/18 (61%)
Frame = -1
Query: 86 TFFYQCVYLLLHPSPPVV 103
T F C YLLL PS P V
Sbjct: 252 TLFTNCTYLLLSPSNPTV 199
>TC12198 similar to UP|Q9LHU3 (Q9LHU3) ESTs D40069(S1808), partial (32%)
Length = 588
Score = 26.2 bits (56), Expect = 1.5
Identities = 13/43 (30%), Positives = 21/43 (48%)
Frame = -3
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAH 108
R+AC L+ F T +F+ + L P+PP+ + AH
Sbjct: 211 RAACGSLEVFYETQHPNGHQVYFHTQSFHLEKPTPPLQSSMAH 83
>NP459594 Krm protein [Lotus japonicus]
Length = 632
Score = 26.2 bits (56), Expect = 1.5
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +1
Query: 93 YLLLHPSPPVVETSAHCSNHF 113
+LLL PS P+ HC +HF
Sbjct: 82 FLLLEPSLPLPYPQIHCPSHF 144
>AV774579
Length = 425
Score = 25.8 bits (55), Expect = 2.0
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = -1
Query: 91 CVYLLLHPSPPVVETSAHCSNHFG 114
C YLLLHP + S +C FG
Sbjct: 155 CTYLLLHPPDLDIVLSCYCK*IFG 84
>TC18829 homologue to UP|Q8L4X4 (Q8L4X4) Amino acid permease-like protein,
partial (27%)
Length = 713
Score = 25.4 bits (54), Expect = 2.6
Identities = 12/52 (23%), Positives = 29/52 (55%), Gaps = 5/52 (9%)
Frame = +1
Query: 48 ITVVGAITLTADDALNKLRSACEE-----LKAFQVTVDRVAAGTFFYQCVYL 94
+TV+GA+T A ++K+ CE+ ++ ++ D + +G +Y +++
Sbjct: 286 LTVMGAVTFYAYFLMSKVLEHCEKSGRRHIRFRELAADVLGSGWMYYFVIFI 441
>TC18857 homologue to UP|Q9ZS21 (Q9ZS21) Glyoxalase I , complete
Length = 877
Score = 23.9 bits (50), Expect = 7.5
Identities = 16/46 (34%), Positives = 22/46 (47%)
Frame = +3
Query: 51 VGAITLTADDALNKLRSACEELKAFQVTVDRVAAGTFFYQCVYLLL 96
+GA+T TA D + +ACE L V G ++ Q V L L
Sbjct: 558 IGAVTQTA*DKTFRSTTACEVL---*VNASECRNGCYYIQSVVLSL 686
>TC17162 similar to UP|Q8H5F1 (Q8H5F1) OJ1165_F02.16 protein, partial (40%)
Length = 784
Score = 23.9 bits (50), Expect = 7.5
Identities = 10/32 (31%), Positives = 15/32 (46%)
Frame = +2
Query: 87 FFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS 118
++Y +Y + V HC+ HF Y SS
Sbjct: 689 YYYYNIYCICVQILHVFYYRVHCTCHFAYSSS 784
>BP050599
Length = 511
Score = 23.9 bits (50), Expect = 7.5
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = +2
Query: 99 SPPVVETSAHCSNHFGYKSS 118
SPP ++AHC +HF SS
Sbjct: 443 SPPSHPSAAHCPSHFLPSSS 502
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 23.5 bits (49), Expect = 9.8
Identities = 8/14 (57%), Positives = 12/14 (85%)
Frame = +2
Query: 98 PSPPVVETSAHCSN 111
P+PP +ET+A CS+
Sbjct: 242 PTPPPLETTASCSS 283
>TC13202
Length = 431
Score = 23.5 bits (49), Expect = 9.8
Identities = 15/51 (29%), Positives = 22/51 (42%), Gaps = 23/51 (45%)
Frame = +2
Query: 86 TFFYQCVYLLLH----------------PSPPVVE-------TSAHCSNHF 113
TFF+ ++LLLH P PP + +SAH ++HF
Sbjct: 20 TFFFFSIFLLLHFPLQP*TTHHLLLLAWPPPPKAKPPPQRAFSSAHANHHF 172
>TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(59%)
Length = 991
Score = 23.5 bits (49), Expect = 9.8
Identities = 11/20 (55%), Positives = 12/20 (60%), Gaps = 1/20 (5%)
Frame = -3
Query: 97 HPSPPVV-ETSAHCSNHFGY 115
H PPVV + S CSNH Y
Sbjct: 626 HLHPPVVSQCSWECSNHHNY 567
>TC15636 similar to UP|Q93WR7 (Q93WR7) MAP kinase kinase, partial (46%)
Length = 1184
Score = 23.5 bits (49), Expect = 9.8
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = -2
Query: 46 PHITVVGAITLTADDALNKLRSACEELKAFQVTVDRV 82
P+IT V L + D+L E+++ FQV VD +
Sbjct: 181 PYITFVVVAILASIDSLWLPSMINEQIRRFQVPVDNM 71
>BP085068
Length = 440
Score = 23.5 bits (49), Expect = 9.8
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = -2
Query: 19 IPPDHVRPRIAKLMTDLRS 37
I PDH RPRI + T S
Sbjct: 406 IRPDHFRPRILSITTSRTS 350
>BP064571
Length = 439
Score = 23.5 bits (49), Expect = 9.8
Identities = 16/45 (35%), Positives = 22/45 (48%), Gaps = 3/45 (6%)
Frame = -1
Query: 73 KAFQVTVDRVAAGTFFYQC---VYLLLHPSPPVVETSAHCSNHFG 114
+ FQ + R+ TF + + +LLHPS VE S S H G
Sbjct: 349 RPFQYLILRLLCPTFLRKGRSRLTMLLHPSVSSVEPSLTPSLHIG 215
>AV407558
Length = 310
Score = 23.5 bits (49), Expect = 9.8
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +1
Query: 87 FFYQCVYLLLHPSPPV 102
F YQ +LL HPSP V
Sbjct: 31 FLYQIWFLLEHPSPLV 78
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,351,683
Number of Sequences: 28460
Number of extensions: 28243
Number of successful extensions: 175
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 175
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 175
length of query: 118
length of database: 4,897,600
effective HSP length: 79
effective length of query: 39
effective length of database: 2,649,260
effective search space: 103321140
effective search space used: 103321140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0162a.1


