

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
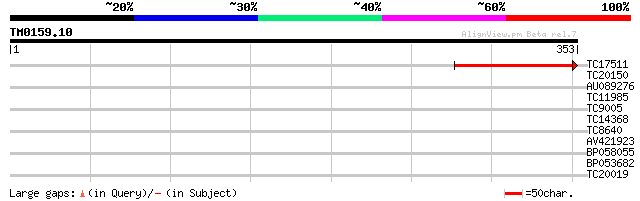
Query= TM0159.10
(353 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17511 similar to UP|AAR92260 (AAR92260) At3g46180, partial (14%) 156 4e-39
TC20150 homologue to UP|CHSY_PUELO (P23569) Chalcone synthase (... 29 1.0
AU089276 29 1.4
TC11985 similar to GB|AAF40452.1|7211981|F13M7 ESTs gb|N65605, g... 28 2.3
TC9005 similar to UP|Q84L76 (Q84L76) Expansin precursor, partial... 27 5.2
TC14368 similar to UP|Q9LLC2 (Q9LLC2) Xyloglucan endotransglycos... 27 5.2
TC8640 27 6.8
AV421923 27 6.8
BP058055 26 8.8
BP053682 26 8.8
TC20019 high affinity ammonium transporter [Lotus corniculatus v... 26 8.8
>TC17511 similar to UP|AAR92260 (AAR92260) At3g46180, partial (14%)
Length = 606
Score = 156 bits (395), Expect = 4e-39
Identities = 76/76 (100%), Positives = 76/76 (100%)
Frame = +3
Query: 278 ALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDS 337
ALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDS
Sbjct: 3 ALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDS 182
Query: 338 IPLVQSGDSNNLKDNP 353
IPLVQSGDSNNLKDNP
Sbjct: 183 IPLVQSGDSNNLKDNP 230
>TC20150 homologue to UP|CHSY_PUELO (P23569) Chalcone synthase
(Naringenin-chalcone synthase) , partial (24%)
Length = 419
Score = 29.3 bits (64), Expect = 1.0
Identities = 12/42 (28%), Positives = 23/42 (54%)
Frame = -2
Query: 142 KRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVW 183
K GP + A ++ ++F + G+D++ YS+ R N +W
Sbjct: 199 KMVAGPSAL*ALRISATLTIFEMKFVGKDVNLYSK*RSNVIW 74
>AU089276
Length = 563
Score = 28.9 bits (63), Expect = 1.4
Identities = 15/52 (28%), Positives = 25/52 (47%)
Frame = -2
Query: 208 GYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLS 259
G+ ++ F +T C + TG + HL+P+ + RH CF + L S
Sbjct: 505 GFQFILYCSYFCSTTCRLSHTATG---EMHLVPSTXYFGRHISCFQHVLLAS 359
>TC11985 similar to GB|AAF40452.1|7211981|F13M7 ESTs gb|N65605, gb|N38087,
gb|T20485, gb|T13726, gb|N38339, gb|F15440 and gb|N97201
come from this gene. {Arabidopsis thaliana;}, partial
(88%)
Length = 1334
Score = 28.1 bits (61), Expect = 2.3
Identities = 18/68 (26%), Positives = 36/68 (52%), Gaps = 6/68 (8%)
Frame = -1
Query: 78 SLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYV------SFPVQTLAKCAKMIP 131
SLL SK + ++ CL+ S + T CQ+E+ ++ S P+ + + + ++P
Sbjct: 644 SLLFSKPTI-----VFSLCLLHSSLLPATLCQFESSPFLFLSPFSSLPLLSSCEPSPLVP 480
Query: 132 VMIWGTII 139
V +W +I+
Sbjct: 479 V-VWPSIL 459
>TC9005 similar to UP|Q84L76 (Q84L76) Expansin precursor, partial (92%)
Length = 1277
Score = 26.9 bits (58), Expect = 5.2
Identities = 11/46 (23%), Positives = 24/46 (51%)
Frame = +2
Query: 299 WFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQSG 344
+FS PL W ++ ++ + + +AP +++++ PL Q G
Sbjct: 59 FFSAPLHWSPFLSQPVLESPAFTPAALGRAPTPPSTAEATPLEQWG 196
>TC14368 similar to UP|Q9LLC2 (Q9LLC2) Xyloglucan endotransglycosylase XET2
, partial (46%)
Length = 565
Score = 26.9 bits (58), Expect = 5.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +3
Query: 66 CNRITTSAVSAGSLLASKKALDPVAPIYKYCL 97
C R T S ++A + KA P API++ CL
Sbjct: 321 CRRTTWSTITALTRRDFHKASQPNAPIHRICL 416
>TC8640
Length = 722
Score = 26.6 bits (57), Expect = 6.8
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 1/49 (2%)
Frame = +2
Query: 287 TRQLVSIMLSCVWFSHPLSWEQ-WIGAVIVFGSLYAKSFTRKAPQKTTS 334
T L + + C LS+E+ WI ++F Y FTR+A K S
Sbjct: 392 TSTLCKVFVVCGLVYSELSYEKNWIDFFLIFFFCYISHFTRRASGKLYS 538
>AV421923
Length = 463
Score = 26.6 bits (57), Expect = 6.8
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = -2
Query: 208 GYDMEIHNQIFYTTLCSCILSLTGLILQG 236
G+D+E+ + +TL LS+ GL+L+G
Sbjct: 243 GHDIEVPEIMLKSTLLLSFLSMLGLVLEG 157
>BP058055
Length = 483
Score = 26.2 bits (56), Expect = 8.8
Identities = 9/21 (42%), Positives = 16/21 (75%)
Frame = -3
Query: 211 MEIHNQIFYTTLCSCILSLTG 231
M+ + +++T LCSC+L +TG
Sbjct: 214 MKKTSYLYFTLLCSCVLFVTG 152
>BP053682
Length = 483
Score = 26.2 bits (56), Expect = 8.8
Identities = 20/70 (28%), Positives = 32/70 (45%), Gaps = 5/70 (7%)
Frame = +1
Query: 96 CLVSVSNILTTTCQYEA-----LKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYM 150
C ++V+N + C + L +FP TL ++MIP I + MQ+R +
Sbjct: 259 CSLAVANA*VSVCSDKGAPESILLLATFPAITLGSVSRMIPPSIISS--MQRRIRSKFSS 432
Query: 151 LAFLVTLGCS 160
+F TL S
Sbjct: 433 ASFWSTLALS 462
>TC20019 high affinity ammonium transporter [Lotus corniculatus var.
japonicus]
Length = 1521
Score = 26.2 bits (56), Expect = 8.8
Identities = 12/44 (27%), Positives = 20/44 (45%)
Frame = +1
Query: 38 YGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLA 81
+G + K Y Y S +LVF ++ + + AGS+ A
Sbjct: 97 FGAISNKFTDTAYAVDNTYLLFSAYLVFAMQLGFAMLCAGSVRA 228
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,946,158
Number of Sequences: 28460
Number of extensions: 101054
Number of successful extensions: 833
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 820
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 832
length of query: 353
length of database: 4,897,600
effective HSP length: 91
effective length of query: 262
effective length of database: 2,307,740
effective search space: 604627880
effective search space used: 604627880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0159.10


