

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0157a.4
(594 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
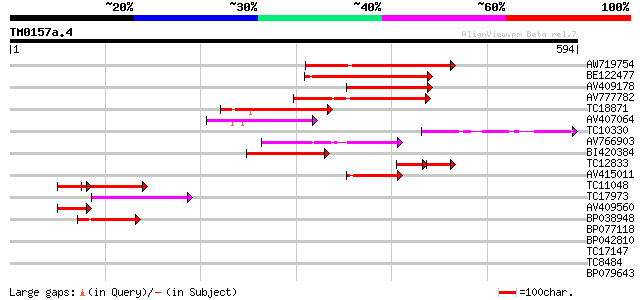
Score E
Sequences producing significant alignments: (bits) Value
AW719754 144 3e-35
BE122477 106 9e-24
AV409178 106 9e-24
AV777782 101 4e-22
TC18871 weakly similar to UP|O04344 (O04344) At2g30510 protein, ... 79 2e-15
AV407064 70 7e-13
TC10330 similar to UP|O04344 (O04344) At2g30510 protein, partial... 70 1e-12
AV766903 69 3e-12
BI420384 67 1e-11
TC12833 similar to UP|Q9LYW0 (Q9LYW0) Photoreceptor-interacting ... 47 6e-11
AV415011 60 1e-09
TC11048 similar to UP|O04343 (O04343) At2g30520 protein, partial... 54 5e-08
TC17973 similar to UP|Q9LIM6 (Q9LIM6) Gb|AAF16571.1, partial (24%) 53 1e-07
AV409560 47 9e-06
BP038948 45 3e-05
BP077118 35 0.026
BP042810 30 1.1
TC17147 similar to UP|Q8LP14 (Q8LP14) Nine-cis-epoxycarotenoid d... 30 1.4
TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), part... 29 1.9
BP079643 29 1.9
>AW719754
Length = 485
Score = 144 bits (364), Expect = 3e-35
Identities = 72/157 (45%), Positives = 108/157 (67%)
Frame = +2
Query: 311 IPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYL 370
+PSYS+ TL+DVD V RI+ +LE ++V D SPS ++ VGKL++ YL
Sbjct: 2 VPSYSYLNETLYDVDCVARILSHFLEGLEAGNTVEGGDAAARSPS---LMLVGKLIDGYL 172
Query: 371 AEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHC 430
+EIA+D NL +F + A +P+Q+ +DG YRA+D+YLKAHP++SE E++ +C ++ C
Sbjct: 173 SEIASDANLKPERFYTFAISLPDQARLFDDGLYRAVDVYLKAHPWVSESEREKICGLLDC 352
Query: 431 QKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETM 467
QKL+ +A HAAQN+RLP++ V+QVL F+Q LR +
Sbjct: 353 QKLTLEACTHAAQNERLPLRAVVQVLFFEQLQLRHAI 463
>BE122477
Length = 420
Score = 106 bits (265), Expect = 9e-24
Identities = 59/136 (43%), Positives = 87/136 (63%), Gaps = 2/136 (1%)
Frame = +3
Query: 310 LIPSYSFPGNTLFDVDTVQRIVVSYLEFE--IGNHSVSSADDEYFSPSQRGIVRVGKLME 367
LIP+YS + L++ D ++ IV ++ E + S SS D SPS +V KL++
Sbjct: 15 LIPTYS-DSDALYNTDCIEHIVQHFIRAESNLTAFSPSSLDQHASSPSSESSKKVAKLID 191
Query: 368 NYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSV 427
+Y+AEIA+D NL K ++AE +PE S DG YRA+DIY KAHP+LS+ EK+ +C++
Sbjct: 192 SYIAEIASDVNLKPGKIRALAEGLPESSRLLHDGLYRALDIYFKAHPWLSDKEKEELCNI 371
Query: 428 MHCQKLSRDARAHAAQ 443
+ +KLS A AHA+Q
Sbjct: 372 IDYRKLSIHACAHASQ 419
>AV409178
Length = 309
Score = 106 bits (265), Expect = 9e-24
Identities = 53/91 (58%), Positives = 67/91 (73%)
Frame = +1
Query: 353 SPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKA 412
SPSQ +V+V KL++NYLAEIA D NL SKF+ IAE +P + DG YRAIDIYLKA
Sbjct: 37 SPSQTALVKVSKLVDNYLAEIAPDANLKLSKFLVIAETLPAHARTVHDGLYRAIDIYLKA 216
Query: 413 HPFLSEMEKKNVCSVMHCQKLSRDARAHAAQ 443
H LS+++KK +C ++ QKLS +A AHAAQ
Sbjct: 217 HQGLSDLDKKKLCKMIDFQKLSHEAGAHAAQ 309
>AV777782
Length = 424
Score = 101 bits (251), Expect = 4e-22
Identities = 55/144 (38%), Positives = 87/144 (60%)
Frame = +1
Query: 298 AMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQR 357
++Q +A + DLL PS S +D + V ++ S+++F S + D+ +F R
Sbjct: 1 SIQFEEATVSDLLYPSTSPLDQNFYDTELVLAVLESFVKFW-KRISPGAVDNRHFL---R 168
Query: 358 GIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLS 417
I +VGKL+++YL +A D N+P KF+++AE +P D YRAI+IYLK HP LS
Sbjct: 169 SIRKVGKLIDSYLQVVARDENMPVQKFVALAETVPAIGRIEHDDLYRAINIYLKVHPDLS 348
Query: 418 EMEKKNVCSVMHCQKLSRDARAHA 441
+++KK +C + CQ+LS + AHA
Sbjct: 349 KVDKKCLCRSLECQRLSPEVCAHA 420
>TC18871 weakly similar to UP|O04344 (O04344) At2g30510 protein, partial
(34%)
Length = 565
Score = 79.3 bits (194), Expect = 2e-15
Identities = 42/121 (34%), Positives = 74/121 (60%), Gaps = 3/121 (2%)
Frame = +2
Query: 221 IMLYAQKSLEGLEIFGKGRKELEPQHEHEK---RVVVETLVSLLPREKNAMSVSSLSMLL 277
++ YA++++ L GKG + + + R ++E++V L P EK + L LL
Sbjct: 122 LITYAERAIRDLT--GKGIRSSDSSETDSRVNQRKLLESIVELFPSEKTVFPMHFLCCLL 295
Query: 278 RAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEF 337
R AI+L + C+ DLEKR+++ L +DDLL+ S+S+ G+ LFD+++V++IV ++E
Sbjct: 296 RCAIHLRASGTCKSDLEKRISLILEHVTVDDLLVLSFSYDGDKLFDLESVRKIVSGFVEK 475
Query: 338 E 338
E
Sbjct: 476 E 478
>AV407064
Length = 377
Score = 70.5 bits (171), Expect = 7e-13
Identities = 43/125 (34%), Positives = 68/125 (54%), Gaps = 9/125 (7%)
Frame = +1
Query: 207 MMGRGFKQFDLGPIIMLYAQKSLEGL---EIFGKGRKEL------EPQHEHEKRVVVETL 257
M RG +Q + + YA+ L GL ++ G+ L P E ++++++E +
Sbjct: 1 MESRGIRQEIIAGSLAFYAKMYLPGLNRRQVSGESSTRLTQMALGSPPCEEDQKILLEEI 180
Query: 258 VSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFP 317
LLP +K + L LLR A+ L +C DLEKR+ MQL QA L+DLL+P++S+
Sbjct: 181 NRLLPLQKGLVQTKFLFGLLRTAMILRVDSSCISDLEKRIGMQLDQANLEDLLMPTFSYS 360
Query: 318 GNTLF 322
TL+
Sbjct: 361 METLY 375
>TC10330 similar to UP|O04344 (O04344) At2g30510 protein, partial (18%)
Length = 607
Score = 69.7 bits (169), Expect = 1e-12
Identities = 57/165 (34%), Positives = 82/165 (49%), Gaps = 2/165 (1%)
Frame = +1
Query: 432 KLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELN 491
KLS +AR HA+QN RLPVQ VL L + Q LR +DGG + + ++L
Sbjct: 4 KLSYEARVHASQNKRLPVQIVLHALYYDQLRLRSGADDGG---EAVAEREQL-------- 150
Query: 492 PVSKVISNLRRENEELKREIVKLKMKLQEIEKPAHESSAPSSPLISAYSPSVNKPPLPRK 551
+ L RENEEL+ E++++K + +++K APSS +
Sbjct: 151 ---QTDVTLVRENEELRLELMRMKTVVSDLQK----GVAPSSS--------------KKI 267
Query: 552 SFMKSVSRKLGRLYPFSRAAADT--VTTPFKDRLKPEKKIRHSIS 594
F SVS+KLG+L PF + DT + D KP ++ R SIS
Sbjct: 268 KFFSSVSKKLGKLNPFRNGSKDTTHLEDGPVDLTKPRRR-RFSIS 399
>AV766903
Length = 548
Score = 68.6 bits (166), Expect = 3e-12
Identities = 48/148 (32%), Positives = 83/148 (55%)
Frame = -3
Query: 264 EKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFD 323
+++++S +L +LR + L + + R LE + L QA LD+LLIPS + + L+D
Sbjct: 504 DQSSISCKTLFGILRVTLSLSISKSSRNKLENMIGSHLDQATLDNLLIPSPNGI-SYLYD 328
Query: 324 VDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADRNLPASK 383
V+ V R + ++L GN P+ + +V +L++ YLAEIA D L ASK
Sbjct: 327 VNLVLRFLKAFLRQ--GNSL----------PTPIQMRKVARLIDLYLAEIAPDPCLKASK 184
Query: 384 FISIAELIPEQSIPTEDGKYRAIDIYLK 411
F+++ +P+ + + D Y A+D+YL+
Sbjct: 183 FLALVTALPDSARDSYDELYHAMDMYLE 100
>BI420384
Length = 319
Score = 66.6 bits (161), Expect = 1e-11
Identities = 38/87 (43%), Positives = 56/87 (63%)
Frame = +3
Query: 249 EKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDD 308
+KR +ETLV +LP EK+A+ + L LLR A + + R +LEKR++ QL QA L +
Sbjct: 54 KKRFFMETLVGVLPPEKDAVPCNFLLRLLRTANMVGVEGSYRQELEKRISWQLDQASLKE 233
Query: 309 LLIPSYSFPGNTLFDVDTVQRIVVSYL 335
L+IPS+S TL DV+ V R+V ++
Sbjct: 234 LVIPSFSHTCGTLLDVELVIRLVKRFV 314
>TC12833 similar to UP|Q9LYW0 (Q9LYW0) Photoreceptor-interacting
protein-like (Non-phototropic hypocotyl-like protein),
partial (15%)
Length = 734
Score = 47.0 bits (110), Expect(2) = 6e-11
Identities = 18/32 (56%), Positives = 27/32 (84%)
Frame = +2
Query: 406 IDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDA 437
IDIYLK HP++++ EK+ +C +M+CQKLS +A
Sbjct: 32 IDIYLKTHPWMTDSEKEQICRLMNCQKLSLEA 127
Score = 37.0 bits (84), Expect(2) = 6e-11
Identities = 16/31 (51%), Positives = 25/31 (80%)
Frame = +1
Query: 437 ARAHAAQNDRLPVQTVLQVLSFQQKHLRETM 467
++ HAAQN+RLP++ V+QVL F+Q LR ++
Sbjct: 124 SQPHAAQNERLPLRVVVQVLFFEQLKLRTSV 216
>AV415011
Length = 238
Score = 59.7 bits (143), Expect = 1e-09
Identities = 30/59 (50%), Positives = 42/59 (70%)
Frame = +2
Query: 353 SPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLK 411
+PS++ ++V KL++ YLAEIA D +LP S F+ +AEL+ T DG YRAID+YLK
Sbjct: 65 NPSKK--IKVVKLVDGYLAEIARDPSLPVSNFVDLAELVSSFPRQTHDGLYRAIDMYLK 235
>TC11048 similar to UP|O04343 (O04343) At2g30520 protein, partial (79%)
Length = 514
Score = 54.3 bits (129), Expect = 5e-08
Identities = 28/71 (39%), Positives = 46/71 (64%), Gaps = 2/71 (2%)
Frame = +2
Query: 76 NASFSLHKM--LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLL 133
N ++H + LRC AE+LEMT++ +L GR++ +L++VA T++GAV+VL LL
Sbjct: 302 NFEITVHNVAVLRCAAEFLEMTDEYCDNSLAGRTEDFLSQVAFFTLTGAVTVLKSCRHLL 481
Query: 134 QVAEKAKLVNR 144
+A+ +V R
Sbjct: 482 PIADDLSIVER 514
Score = 46.2 bits (108), Expect = 1e-05
Identities = 20/35 (57%), Positives = 26/35 (74%)
Frame = +2
Query: 51 SSAMKKTSEWNFPQEIPSDVTVQIDNASFSLHKML 85
S AM++T +W F QEIP+DV VQ+ ASF LHK +
Sbjct: 62 SLAMERTGQWIFSQEIPTDVIVQVGEASFCLHKFM 166
>TC17973 similar to UP|Q9LIM6 (Q9LIM6) Gb|AAF16571.1, partial (24%)
Length = 919
Score = 53.1 bits (126), Expect = 1e-07
Identities = 35/110 (31%), Positives = 54/110 (48%), Gaps = 4/110 (3%)
Frame = +3
Query: 86 RCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRC 145
RC AEYL+MTE+ GNL+ + D + N L ++ L + L +E ++ +RC
Sbjct: 588 RCSAEYLQMTEEVEKGNLIQKLDVFFNSCILRGWKDSIVSLQTTKALHLWSEDLEITSRC 767
Query: 146 IDAIA--YMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVH--WWGED 191
I+AIA ++ S + S+ D V S + R + WWGED
Sbjct: 768 IEAIASKVLSHPTKVSLSHSHSRRVRDDVSSCNETESVRHKSNKGWWGED 917
>AV409560
Length = 375
Score = 47.0 bits (110), Expect = 9e-06
Identities = 21/35 (60%), Positives = 26/35 (74%)
Frame = +3
Query: 51 SSAMKKTSEWNFPQEIPSDVTVQIDNASFSLHKML 85
S AM++T +W F QEIPSDVTV I A +SLHK +
Sbjct: 117 SLAMERTGQWVFSQEIPSDVTVTIGEADYSLHKFM 221
>BP038948
Length = 508
Score = 45.1 bits (105), Expect = 3e-05
Identities = 23/66 (34%), Positives = 42/66 (62%)
Frame = +3
Query: 72 VQIDNASFSLHKMLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPET 131
V+ID +S ++ LRC E+LEMTE+ S NL+ +++ +L+ L ++ +++ L E
Sbjct: 309 VKIDLSSSNVAS-LRCAGEFLEMTEEYSEDNLISKTEQFLSRSVLNSVKDSITALQSCER 485
Query: 132 LLQVAE 137
L+ +AE
Sbjct: 486 LMPLAE 503
Score = 30.0 bits (66), Expect = 1.1
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +3
Query: 58 SEWNFPQEIPSDVTVQIDNASFSLHKM 84
+ W +PSD+ VQ+D+ +F LHK+
Sbjct: 75 NSWFCTTGLPSDIVVQLDDMNFHLHKI 155
>BP077118
Length = 354
Score = 35.4 bits (80), Expect = 0.026
Identities = 16/23 (69%), Positives = 18/23 (77%)
Frame = +3
Query: 62 FPQEIPSDVTVQIDNASFSLHKM 84
F QEIPSDVTV I A +SLHK+
Sbjct: 45 FSQEIPSDVTVTIGEADYSLHKV 113
>BP042810
Length = 503
Score = 30.0 bits (66), Expect = 1.1
Identities = 14/44 (31%), Positives = 24/44 (53%), Gaps = 4/44 (9%)
Frame = -2
Query: 44 LRKKELKSSAMKKTSE----WNFPQEIPSDVTVQIDNASFSLHK 83
+ + K S+ +K S W +PSD+ V++D+ +F LHK
Sbjct: 376 IHNSQTKMSSSEKHSSKGQAWFCTTGLPSDIVVEVDDMTFHLHK 245
>TC17147 similar to UP|Q8LP14 (Q8LP14) Nine-cis-epoxycarotenoid
dioxygenase4, partial (22%)
Length = 663
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = -1
Query: 522 EKPAHESSAPSSPLISAYSPSVNKPPLP 549
E P+ ++S P SP + SPS + PPLP
Sbjct: 276 EHPSPKTSTPPSPHHARNSPSTHHPPLP 193
>TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), partial (76%)
Length = 1121
Score = 29.3 bits (64), Expect = 1.9
Identities = 16/55 (29%), Positives = 25/55 (45%)
Frame = +1
Query: 524 PAHESSAPSSPLISAYSPSVNKPPLPRKSFMKSVSRKLGRLYPFSRAAADTVTTP 578
P + +PSS ++ +PS + PP P S + L YP S A + +P
Sbjct: 175 PPNSPPSPSSSTVATPNPSDSHPPPPSPPPSPSATNSLNPPYPTSTTPARSKPSP 339
>BP079643
Length = 451
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/51 (27%), Positives = 20/51 (38%)
Frame = +2
Query: 499 NLRRENEELKREIVKLKMKLQEIEKPAHESSAPSSPLISAYSPSVNKPPLP 549
N E + + + P H S PS P +S + S + PPLP
Sbjct: 182 NFFHRTEPFSTHFLSILLPSLSSSSPLHPPSLPSPPSLSPFFRSPSSPPLP 334
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,094,962
Number of Sequences: 28460
Number of extensions: 114347
Number of successful extensions: 777
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 766
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 773
length of query: 594
length of database: 4,897,600
effective HSP length: 95
effective length of query: 499
effective length of database: 2,193,900
effective search space: 1094756100
effective search space used: 1094756100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0157a.4


