

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
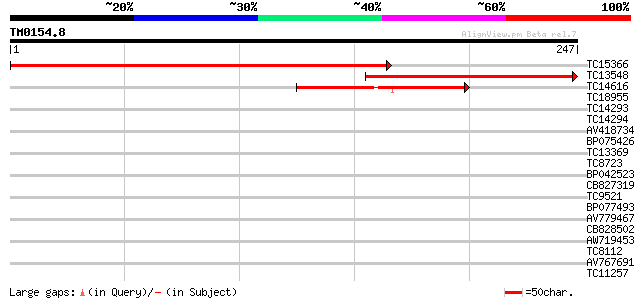
Score E
Sequences producing significant alignments: (bits) Value
TC15366 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Ar... 343 1e-95
TC13548 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Ar... 183 3e-47
TC14616 78 1e-15
TC18955 similar to UP|Q9LU67 (Q9LU67) Phosphatidylserine decarbo... 28 1.9
TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 28 1.9
TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 28 1.9
AV418734 27 2.5
BP075426 27 3.3
TC13369 similar to UP|Q9FPK0 (Q9FPK0) AT4g38460, partial (35%) 27 3.3
TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain,... 27 3.3
BP042523 27 3.3
CB827319 27 4.3
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 27 4.3
BP077493 26 5.6
AV779467 26 7.3
CB828502 26 7.3
AW719453 26 7.3
TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1... 26 7.3
AV767691 26 7.3
TC11257 UP|Q7ZPS7 (Q7ZPS7) Tat protein (Fragment), partial (18%) 25 9.5
>TC15366 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Arabidopsis
thaliana;}, partial (23%)
Length = 568
Score = 343 bits (881), Expect = 1e-95
Identities = 166/166 (100%), Positives = 166/166 (100%)
Frame = +2
Query: 1 RCGVELLPQLKAMATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFT 60
RCGVELLPQLKAMATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFT
Sbjct: 71 RCGVELLPQLKAMATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFT 250
Query: 61 TATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGD 120
TATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGD
Sbjct: 251 TATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGD 430
Query: 121 SVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTIC 166
SVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTIC
Sbjct: 431 SVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTIC 568
>TC13548 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Arabidopsis
thaliana;}, partial (22%)
Length = 515
Score = 183 bits (464), Expect = 3e-47
Identities = 92/92 (100%), Positives = 92/92 (100%)
Frame = +2
Query: 156 PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQS 215
PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQS
Sbjct: 2 PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQS 181
Query: 216 EQGVAATLQSTDITSSQKKWMKRLQGNRENKE 247
EQGVAATLQSTDITSSQKKWMKRLQGNRENKE
Sbjct: 182 EQGVAATLQSTDITSSQKKWMKRLQGNRENKE 277
>TC14616
Length = 1353
Score = 78.2 bits (191), Expect = 1e-15
Identities = 42/78 (53%), Positives = 55/78 (69%), Gaps = 3/78 (3%)
Frame = +3
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV G+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 465 VVAAGLGKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPVFD 641
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 642 TLDVSPLLAFAVLGTLGS 695
>TC18955 similar to UP|Q9LU67 (Q9LU67) Phosphatidylserine decarboxylase,
partial (6%)
Length = 523
Score = 27.7 bits (60), Expect = 1.9
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 4/77 (5%)
Frame = +2
Query: 30 TQTPFALPILKPPISNLTSFLSPPE--LHKSFTTATDNCFRFLHSLASQNPFL--NKILS 85
T P ALP PP++N + + PE SFT + L +L S+N + S
Sbjct: 92 TVVPKALPTPLPPLANYSPMTTSPESLSSLSFTRRCSSRINGLRALLSENRLSVPRPLTS 271
Query: 86 LRSEFDSLCFQIRCSNY 102
F CF + S Y
Sbjct: 272 SLHSFIHSCFLLPISFY 322
>TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (18%)
Length = 540
Score = 27.7 bits (60), Expect = 1.9
Identities = 12/29 (41%), Positives = 20/29 (68%)
Frame = -3
Query: 105 RRWLSSHNFAAVLPGDSVAGLVVGNGIQN 133
RRW+++ AAV G + GL+VG G+++
Sbjct: 418 RRWVTATTAAAVGMGFVLGGLLVGRGVRS 332
>TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (65%)
Length = 1192
Score = 27.7 bits (60), Expect = 1.9
Identities = 12/29 (41%), Positives = 20/29 (68%)
Frame = -2
Query: 105 RRWLSSHNFAAVLPGDSVAGLVVGNGIQN 133
RRW+++ AAV G + GL+VG G+++
Sbjct: 405 RRWVTATTAAAVGMGFVLGGLLVGRGVRS 319
>AV418734
Length = 287
Score = 27.3 bits (59), Expect = 2.5
Identities = 18/55 (32%), Positives = 25/55 (44%), Gaps = 4/55 (7%)
Frame = +3
Query: 52 PPELHKSFTTATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIR----CSNY 102
P L S +C R H ++S N N S S S+C+QIR C+N+
Sbjct: 27 PCSLVSSSWKHVQHCLRNFHGMSSCNWNCNLSQSCPSFMVSICYQIRHHDNCNNF 191
>BP075426
Length = 342
Score = 26.9 bits (58), Expect = 3.3
Identities = 29/91 (31%), Positives = 40/91 (43%), Gaps = 10/91 (10%)
Frame = +3
Query: 54 ELH-KSFTTATDNCFRFLHS----LASQNPFLNKILSLRSEFDSLCFQIRCSNY-RSRRW 107
ELH +S T T +C L + QNPFL+K+ L + L + CS S +
Sbjct: 51 ELHVQSTNTQTRDCACSLEFDL*LNSEQNPFLSKMQQLWMDSHQLDYHCHCSGISSSSKC 230
Query: 108 LSSHNFAAVLPGD----SVAGLVVGNGIQNF 134
+ S + A L VA +VG GI F
Sbjct: 231 MKSESVADFLRIKL*IVIVAISIVGRGISKF 323
>TC13369 similar to UP|Q9FPK0 (Q9FPK0) AT4g38460, partial (35%)
Length = 559
Score = 26.9 bits (58), Expect = 3.3
Identities = 10/38 (26%), Positives = 21/38 (54%)
Frame = -2
Query: 65 NCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNY 102
+CF +L + N F+ ++ L + +C QI+ S++
Sbjct: 318 HCFIYLRRKLN*NSFIQLLIDLHHQGSPICLQIKMSSF 205
>TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain, complete
Length = 1689
Score = 26.9 bits (58), Expect = 3.3
Identities = 24/96 (25%), Positives = 36/96 (37%)
Frame = -3
Query: 8 PQLKAMATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCF 67
P+ K + E + + NT T K I N+T+ PPELH + T +
Sbjct: 1564 PREKGIQVEGEKDNIWQKSHHNTHTD------KNTIQNITTKNPPPELHPHLHSHTPHHL 1403
Query: 68 RFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNYR 103
+ A SL S + +RC+N R
Sbjct: 1402 QLHPGTAGTPRQGRSCCSLPQ*TPSHPYPLRCTNAR 1295
>BP042523
Length = 479
Score = 26.9 bits (58), Expect = 3.3
Identities = 16/50 (32%), Positives = 20/50 (40%)
Frame = +2
Query: 17 TDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNC 66
T P +SH WR+T PP S +S P L + AT C
Sbjct: 53 TPPSATTSHGWRSTD---------PPRSPTSSATKTPSLASKSSLATATC 175
>CB827319
Length = 532
Score = 26.6 bits (57), Expect = 4.3
Identities = 19/67 (28%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Frame = -3
Query: 46 LTSFLSPPELHKSFTTATDNCF--RFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNYR 103
+T F S L S NCF F H+L+ + L+ + + C ++C R
Sbjct: 458 MTLFFSLILLITSLVLCVPNCFSKNFFHNLSVTGLRSHPTLAFSTCIN--CSSVQCGKIR 285
Query: 104 SRRWLSS 110
S RW S+
Sbjct: 284 SSRW*ST 264
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 26.6 bits (57), Expect = 4.3
Identities = 12/34 (35%), Positives = 16/34 (46%)
Frame = -2
Query: 103 RSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLN 136
RSRRW ++H F +P + GN F N
Sbjct: 384 RSRRWTTTHIFPVAVPAPAPFTRTTGNPFCKFNN 283
>BP077493
Length = 375
Score = 26.2 bits (56), Expect = 5.6
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +3
Query: 43 ISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQ 76
I L SP +LH++ T NC F H++ SQ
Sbjct: 255 IPYLLLLTSPCKLHENLCIPTSNCSIFHHAVYSQ 356
>AV779467
Length = 456
Score = 25.8 bits (55), Expect = 7.3
Identities = 12/34 (35%), Positives = 22/34 (64%), Gaps = 1/34 (2%)
Frame = -3
Query: 85 SLRSEFDSLCFQIRC-SNYRSRRWLSSHNFAAVL 117
+L ++F + +C SNY S +W+S H+++ VL
Sbjct: 259 TLIAQFGWAYWAYQCDSNYWSLQWMSDHHYSPVL 158
>CB828502
Length = 536
Score = 25.8 bits (55), Expect = 7.3
Identities = 25/85 (29%), Positives = 37/85 (43%)
Frame = -2
Query: 17 TDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQ 76
T P V+ R+ T F P + P + LT P + +FT+ NC RFL L +
Sbjct: 250 TYPVTVTLLALRSISTEFT-PFI-PSTAFLTFLSHPSQSIFTFTSTVCNCLRFLCELPLE 77
Query: 77 NPFLNKILSLRSEFDSLCFQIRCSN 101
+++ S F +L I SN
Sbjct: 76 QSKRSEM*SRAPIFVNLSSIISSSN 2
>AW719453
Length = 546
Score = 25.8 bits (55), Expect = 7.3
Identities = 26/98 (26%), Positives = 41/98 (41%), Gaps = 4/98 (4%)
Frame = +1
Query: 135 LNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLF----RGLIPPLGGTLDLSPIL 190
++I T L+V L P PP + PL P NL + +PPL +L
Sbjct: 271 MSIVITQLIVSYHLLPPPPRPP*PLPPLENPRPPLPNLLPPPPKPPLPPLDWSLSCKRG- 447
Query: 191 AFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDI 228
N+FTS+ LPA + + + +A T + +
Sbjct: 448 -----NSFTSSRLTLPAFF*PARNASKSLAGTSNTAPV 546
>TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1 component
alpha subunit, mitochondrial precursor (PDHE1-A) ,
partial (90%)
Length = 1816
Score = 25.8 bits (55), Expect = 7.3
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 7/55 (12%)
Frame = -2
Query: 20 KKVSSHCWRNTQTPFA---LPILKPPI----SNLTSFLSPPELHKSFTTATDNCF 67
+K++++ R Q +A L I + P SN+ L+PPELH + T NC+
Sbjct: 1602 QKLATN*GRQVQGSYAQ*ILQICRTPSCINHSNMLWLLAPPELHGNKY*FTPNCY 1438
>AV767691
Length = 617
Score = 25.8 bits (55), Expect = 7.3
Identities = 19/57 (33%), Positives = 27/57 (47%)
Frame = +2
Query: 181 GGTLDLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKWMK 237
GGT + PIL + +TS L +LPV + GVA + T +T K M+
Sbjct: 53 GGTRSIRPIL*-PISGLYTSG---LEGKLPVNKGDANGVANVVDGTVLTIPDKPCMR 211
>TC11257 UP|Q7ZPS7 (Q7ZPS7) Tat protein (Fragment), partial (18%)
Length = 593
Score = 25.4 bits (54), Expect = 9.5
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = -1
Query: 92 SLCFQIRCSNYRSRRWLSSHNFAAVLP 118
SLC CS Y SR + NFA V+P
Sbjct: 443 SLCL---CSRYTSRNNMEVQNFANVMP 372
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,141,531
Number of Sequences: 28460
Number of extensions: 83168
Number of successful extensions: 515
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 512
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 513
length of query: 247
length of database: 4,897,600
effective HSP length: 88
effective length of query: 159
effective length of database: 2,393,120
effective search space: 380506080
effective search space used: 380506080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0154.8


