

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
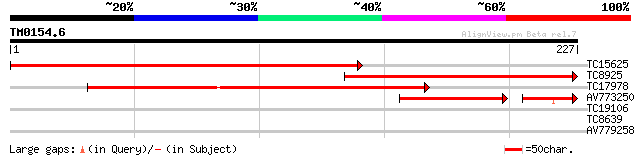
Query= TM0154.6
(227 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC15625 282 4e-77
TC8925 196 4e-51
TC17978 similar to UP|Q9BJD1 (Q9BJD1) SAG1-related sequence 8 (F... 143 3e-35
AV773250 86 1e-20
TC19106 26 6.5
TC8639 similar to UP|FRI4_SOYBN (Q948P5) Ferritin 4, chloroplast... 26 6.5
AV779258 26 6.5
>TC15625
Length = 594
Score = 282 bits (721), Expect = 4e-77
Identities = 141/141 (100%), Positives = 141/141 (100%)
Frame = +2
Query: 1 MAKLYLVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLA 60
MAKLYLVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLA
Sbjct: 170 MAKLYLVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLA 349
Query: 61 CIALQYIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYV 120
CIALQYIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYV
Sbjct: 350 CIALQYIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYV 529
Query: 121 HAPEAFSEVLIQNQRSLEILH 141
HAPEAFSEVLIQNQRSLEILH
Sbjct: 530 HAPEAFSEVLIQNQRSLEILH 592
>TC8925
Length = 535
Score = 196 bits (497), Expect = 4e-51
Identities = 91/93 (97%), Positives = 93/93 (99%)
Frame = +2
Query: 135 RSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKPGCFSGA 194
RSLEILHS+NHTQVGAAVTGTDGGSPYFWCVLFSGG+PNSTFAFEGGVAKLTKPGCFSGA
Sbjct: 2 RSLEILHSQNHTQVGAAVTGTDGGSPYFWCVLFSGGQPNSTFAFEGGVAKLTKPGCFSGA 181
Query: 195 NDECSGAHDWSPLSVMWLFAASVLIALGFAFPL 227
NDECSGAHDWSPLSVMWLFAASVLIALGFAFPL
Sbjct: 182 NDECSGAHDWSPLSVMWLFAASVLIALGFAFPL 280
>TC17978 similar to UP|Q9BJD1 (Q9BJD1) SAG1-related sequence 8 (Fragment),
partial (7%)
Length = 680
Score = 143 bits (360), Expect = 3e-35
Identities = 68/137 (49%), Positives = 93/137 (67%)
Frame = +2
Query: 32 NPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDCGAVGGPDAKKPPESQFA 91
NPA+ +V IN+NRT K+ L D+ GL C+ALQY++ +G+C + K PPE F
Sbjct: 269 NPANDIVDIINKNRTDQKLPHLNDSPGLGCMALQYVELCKGNCTDNNVVNCK-PPEDDFT 445
Query: 92 EVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHSKNHTQVGAA 151
EVFAPNCGV+ + ITG +GCQ KY+ P AFSEVLI++++SL +L +K+HT+VG
Sbjct: 446 EVFAPNCGVELPTFGTITGHIVGCQKKYLEPPLAFSEVLIKDKKSLSLLTNKSHTEVGVX 625
Query: 152 VTGTDGGSPYFWCVLFS 168
+ G P+FWCVLFS
Sbjct: 626 LVGLHNKGPFFWCVLFS 676
>AV773250
Length = 463
Score = 85.5 bits (210), Expect(2) = 1e-20
Identities = 37/43 (86%), Positives = 39/43 (90%)
Frame = -3
Query: 157 GGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKPGCFSGANDECS 199
GGSPYFW VLFSGG+PN AFEGGVAKLTKPGCFSGANDEC+
Sbjct: 461 GGSPYFWFVLFSGGQPNRPLAFEGGVAKLTKPGCFSGANDECT 333
Score = 30.0 bits (66), Expect(2) = 1e-20
Identities = 16/23 (69%), Positives = 17/23 (73%), Gaps = 1/23 (4%)
Frame = -2
Query: 206 PLSVMWLFAAS-VLIALGFAFPL 227
P VMWLFAAS + LGFAFPL
Sbjct: 312 PPPVMWLFAASRSQLHLGFAFPL 244
>TC19106
Length = 725
Score = 25.8 bits (55), Expect = 6.5
Identities = 11/19 (57%), Positives = 12/19 (62%), Gaps = 1/19 (5%)
Frame = -2
Query: 188 PGCFSGANDEC-SGAHDWS 205
PGC S ND C + AH WS
Sbjct: 514 PGCSSPQNDSCRARAHPWS 458
>TC8639 similar to UP|FRI4_SOYBN (Q948P5) Ferritin 4, chloroplast precursor
(SFerH-4), partial (43%)
Length = 515
Score = 25.8 bits (55), Expect = 6.5
Identities = 16/42 (38%), Positives = 16/42 (38%)
Frame = -3
Query: 143 KNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAK 184
K H GA V G GG V K N T EGG K
Sbjct: 240 KQHAGEGAVVAGAFGGGANHHGVRVMAVKRNRTVGVEGGRRK 115
>AV779258
Length = 494
Score = 25.8 bits (55), Expect = 6.5
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = -1
Query: 96 PNCGVKASSLAPITGRFLGC 115
P C +A + AP+T R GC
Sbjct: 395 PGCSCRARTFAPVTPRHKGC 336
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,118,597
Number of Sequences: 28460
Number of extensions: 57248
Number of successful extensions: 251
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 250
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 250
length of query: 227
length of database: 4,897,600
effective HSP length: 87
effective length of query: 140
effective length of database: 2,421,580
effective search space: 339021200
effective search space used: 339021200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0154.6


