

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
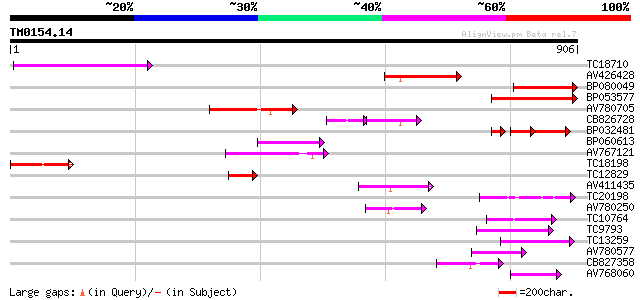
Query= TM0154.14
(906 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18710 176 1e-44
AV426428 174 7e-44
BP080049 150 7e-37
BP053577 138 3e-33
AV780705 111 6e-25
CB826728 70 1e-19
BP032481 55 2e-19
BP060613 87 2e-17
AV767121 84 8e-17
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 76 3e-14
TC12829 75 6e-14
AV411435 70 2e-12
TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial... 69 4e-12
AV780250 68 6e-12
TC10764 64 1e-10
TC9793 62 5e-10
TC13259 61 9e-10
AV780577 60 2e-09
CB827358 59 4e-09
AV768060 59 4e-09
>TC18710
Length = 843
Score = 176 bits (446), Expect = 1e-44
Identities = 97/221 (43%), Positives = 127/221 (56%)
Frame = +2
Query: 7 FQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVLIPKVKK 66
F + ++ G DG+ LFYQK ++ D V+ L L + IN+TL+ LIPKV
Sbjct: 161 FLLETSQPRGMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLVTLIPKVPH 340
Query: 67 PMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNALVAFECF 126
P NQFR IS C ++K+I+K RLK +PDLI Q+ F+ GR I DN L+ E F
Sbjct: 341 PESINQFRPISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDNLLIVQEAF 520
Query: 127 HYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVDFSI 186
H + + + +KLDM+KAYDRVEW FL S L GF +WV +IM VS V ++
Sbjct: 521 HAINRPGALGRNHSIIKLDMNKAYDRVEWKFLESSLLAFGFSTNWVKMIMILVSGVSYNY 700
Query: 187 MLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKA 227
+NG P RGLRQGDP SPYLF+ EV S LI +
Sbjct: 701 KINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLSLLIQNS 823
>AV426428
Length = 413
Score = 174 bits (440), Expect = 7e-44
Identities = 88/137 (64%), Positives = 99/137 (72%), Gaps = 14/137 (10%)
Frame = +3
Query: 599 VASSSQVPSLPRADWRKLWKADAL--------------LPVRAYLHARGLVTDPTCPRCL 644
VASSSQVPSLP D RKLWKA+AL LPV+A LHARGL DP CPRCL
Sbjct: 3 VASSSQVPSLPSVD*RKLWKAEALPRCCETGWRACLGVLPVKASLHARGLAEDPICPRCL 182
Query: 645 VALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLLYA 704
A +T HA+LFCP V PIWFASSLGFRL EC++HEFV+DFMQVAD + G FLT+LYA
Sbjct: 183 AAPKTADHAVLFCP*VHPIWFASSLGFRLNQECKMHEFVADFMQVAD*DTCGEFLTILYA 362
Query: 705 IWMARNELCFKEKASTL 721
IW A+NELCF+ TL
Sbjct: 363 IWTAQNELCFRYTTVTL 413
>BP080049
Length = 356
Score = 150 bits (380), Expect = 7e-37
Identities = 72/101 (71%), Positives = 80/101 (78%)
Frame = -1
Query: 806 LSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFS 865
LS RWALGLA +LGFR +VFE DCL LYEAWKRRGGCS+LD+LILDCR L S FD F S
Sbjct: 356 LSLRWALGLASELGFR*IVFETDCLHLYEAWKRRGGCSVLDTLILDCRNLLSYFDVFQLS 177
Query: 866 FIRRTGNSVADALAKRSFTLGVWVWIEEIPQDFYGLVQSDV 906
F+RRT N V DALAK F LG VW+EE+P + YGLVQSDV
Sbjct: 176 FVRRTSNCVVDALAKLCFQLGNVVWVEELPHELYGLVQSDV 54
>BP053577
Length = 463
Score = 138 bits (348), Expect = 3e-33
Identities = 72/137 (52%), Positives = 89/137 (64%), Gaps = 1/137 (0%)
Frame = +3
Query: 771 GDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCL 830
G AG G+VARN GEV+A+ACSS +S LLAE L RW + LA DLGFRR+ E DCL
Sbjct: 6 GIAGLGMVARNCDGEVMASACSSPVSISSPLLAEALGLRWTMQLATDLGFRRITLETDCL 185
Query: 831 LLYEAWK-RRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALAKRSFTLGVWV 889
L+ W+ + G S L S+I DCR L S FD SF+RRTGN+VAD LA+ S T V
Sbjct: 186 QLFNTWRSQEEGRSYLSSIISDCRILISAFDYVSVSFVRRTGNTVADFLARNSDTYANMV 365
Query: 890 WIEEIPQDFYGLVQSDV 906
W+EE+P L+ +DV
Sbjct: 366 WVEEVPLTVVSLIDADV 416
>AV780705
Length = 524
Score = 111 bits (277), Expect = 6e-25
Identities = 55/147 (37%), Positives = 90/147 (60%), Gaps = 5/147 (3%)
Frame = -2
Query: 319 YLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREVLVKAVAHAIPSYVMSCFL 378
YLGLP I G++K+ +++E+V +KL+GWKE L +AG+EVL+KA+ AIPSY M+
Sbjct: 484 YLGLPAIWGRNKSHSLVWIEEKVKEKLEGWKETLLNQAGKEVLIKAIIQAIPSYAMTIVH 305
Query: 379 LPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLC-----KAKDNGRLGFRDFKMFNSA 433
P CN + +++ F+W +G + + +K+ G +GFRD + N A
Sbjct: 304 FPKTFCNSLYALVADFWW----KSQGEGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQNLA 137
Query: 434 LVAKNWWRIKSNPESLMGKVYRAVYFP 460
+A+ WR+ +NPE+L +V +++YFP
Sbjct: 136 FLARQAWRVLTNPEALWVRVMKSLYFP 56
>CB826728
Length = 509
Score = 69.7 bits (169), Expect(2) = 1e-19
Identities = 38/101 (37%), Positives = 55/101 (53%), Gaps = 14/101 (13%)
Frame = +1
Query: 571 VLYWPFSSDGHYSTKSGYQFLRHGAEAVVASSSQVPSLPRADWRKLWKADAL-------- 622
V +WP SSDG YS+KSGY+F+R ++++ S+S ++P W+ +W A +L
Sbjct: 205 VFFWPGSSDGWYSSKSGYEFIRLENKSLLPSTSPAAAVPSLIWKTVWSASSLPRCKEVMW 384
Query: 623 ------LPVRAYLHARGLVTDPTCPRCLVALETVQHALLFC 657
LPVR+ L RGL DP+ P + E H LL C
Sbjct: 385 RACAGYLPVRSALKRRGLDVDPSSPWSGLEDEAEVHVLLTC 507
Score = 44.3 bits (103), Expect(2) = 1e-19
Identities = 22/68 (32%), Positives = 35/68 (51%)
Frame = +3
Query: 506 DKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLAWNRELINFICCPPTARAIMAIPLPL 565
D W+P + LVY L + V+DL+ G WN L++ + P T +A+ LPL
Sbjct: 15 DAWLPNSAPLVYCEDLFVDLQLKKVADLIENG--RWNENLVSMVFSPSTTAVFLAVSLPL 188
Query: 566 VPHDDVLY 573
++D L+
Sbjct: 189 QSYEDCLF 212
>BP032481
Length = 447
Score = 55.5 bits (132), Expect(3) = 2e-19
Identities = 28/65 (43%), Positives = 40/65 (61%), Gaps = 1/65 (1%)
Frame = -1
Query: 832 LYEAWKRRG-GCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALAKRSFTLGVWVW 890
L+++WK + S L S++ DCR L + FD SF+RRTGN+VA A+ + T VW
Sbjct: 258 LFKSWKAKEVE*SYLSSILQDCRLLFTAFDVVSLSFVRRTGNTVAGFFARNAETYANLVW 79
Query: 891 IEEIP 895
EE+P
Sbjct: 78 PEEVP 64
Score = 45.8 bits (107), Expect(3) = 2e-19
Identities = 21/39 (53%), Positives = 25/39 (63%)
Frame = -2
Query: 801 LLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRR 839
LLAE +S RW + LA+DLGFRR+ E DCL K R
Sbjct: 350 LLAEAMSMRWIMQLAIDLGFRRICLETDCLTFSSLGKXR 234
Score = 31.6 bits (70), Expect(3) = 2e-19
Identities = 13/22 (59%), Positives = 16/22 (72%)
Frame = -3
Query: 771 GDAGFGLVARNSRGEVLAAACS 792
G G G+VARNS G+V+A CS
Sbjct: 439 GLGGMGMVARNSEGDVMATTCS 374
>BP060613
Length = 378
Score = 86.7 bits (213), Expect = 2e-17
Identities = 39/107 (36%), Positives = 61/107 (56%)
Frame = +1
Query: 396 WGGDVTRRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYR 455
W RG+H W L + KD G LGF+DF++ N AL+AK WRI NP++L ++ +
Sbjct: 46 WWSSKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILK 225
Query: 456 AVYFPRGTIRGAKRGYRPSYAWSSIMRSGEIFATGGMWNIGNGKSVK 502
A+YFP + S+ WSS++ E+ G W +G+G++V+
Sbjct: 226 ALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSGETVQ 366
>AV767121
Length = 525
Score = 84.3 bits (207), Expect = 8e-17
Identities = 51/172 (29%), Positives = 83/172 (47%), Gaps = 7/172 (4%)
Frame = +2
Query: 345 LKGWKERSLYRAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRG 404
L WK++++ GR L+++V A+P + +S F LP G+ ++ F WGG
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 405 MHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAV--YFPRG 462
+ + W D+CK K+ G LG +D FN AL+ K WR + P+SL +V A ++ G
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQPDHYSCG 361
Query: 463 TIRGAKRGYRPSYAWSSIMR-----SGEIFATGGMWNIGNGKSVKAWSDKWV 509
S W+ I+ + F++G +G G K WS+ W+
Sbjct: 362 -----------SSWWNDILSLCPEDADGWFSSGLKKLVGEGDQTKFWSEDWL 484
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 75.9 bits (185), Expect = 3e-14
Identities = 42/103 (40%), Positives = 65/103 (62%), Gaps = 1/103 (0%)
Frame = -3
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVL-HGDIS*SIINKTLLV 59
+++EA+F M KAPG DG+ LFYQ+ I+ +DV++ L HG++ + N+TL+
Sbjct: 317 EIKEAVFSMGDLKAPGMDGINGLFYQQNREIVKNDVNNAILDFFDHGELPVEL-NETLVT 141
Query: 60 LIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLI 102
LIPK+ QFR IS C+ I+K+I+K + RLK PD++
Sbjct: 140 LIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARLK---PDMV 21
>TC12829
Length = 448
Score = 74.7 bits (182), Expect = 6e-14
Identities = 31/46 (67%), Positives = 40/46 (86%)
Frame = +3
Query: 350 ERSLYRAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFY 395
E+ LYRAGREVL+K+V AIP+Y+MSCF LPD +C+QIEGM+ +FY
Sbjct: 252 EKFLYRAGREVLIKSVTKAIPTYIMSCFALPDSICSQIEGMVGKFY 389
Score = 37.0 bits (84), Expect = 0.014
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +1
Query: 386 QIEGMISRFYWGGDVTRRGMHCLSWKDLC 414
+++ + F W GDV+RR +H L WK LC
Sbjct: 361 RLKAWLVSFIWSGDVSRRSIHWLGWKKLC 447
>AV411435
Length = 426
Score = 69.7 bits (169), Expect = 2e-12
Identities = 45/133 (33%), Positives = 64/133 (47%), Gaps = 14/133 (10%)
Frame = +3
Query: 558 IMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAV--VASSSQVPS-------- 607
IM P + +D L+WPF G YS KSGY+ ++ + +ASSS PS
Sbjct: 3 IMQTPFSITGSEDRLFWPFIPSGEYSVKSGYRAVKATQFNLSNLASSSSHPSSFLWHTIW 182
Query: 608 ---LPRADWRKLWKA-DALLPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPI 663
+P+ LW+A + +PV+ L R + D CP C E + HALL C + +
Sbjct: 183 GAQVPKKVRSFLWRAANNAIPVKRNLKRRNMGRDDFCPICNKGPEDINHALLACEWTRAV 362
Query: 664 WFASSLGFRLAHE 676
WF L F + HE
Sbjct: 363 WFGCPLNF-VVHE 398
>TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial (13%)
Length = 630
Score = 68.6 bits (166), Expect = 4e-12
Identities = 53/158 (33%), Positives = 79/158 (49%), Gaps = 4/158 (2%)
Frame = +3
Query: 751 WSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRG-EVLAAACSSHGPAASSLLAEGLSFR 809
W P G FK+NFD ++ + AG LV R+ G EV + G AS + +G S+
Sbjct: 93 WPPPPDGVFKLNFDTSVLASS*AGLSLVVRDCAG*EVFSIQN*VFG--ASLISRDG*SY- 263
Query: 810 WALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRR---LSSGFDAFDFSF 866
+ +DL V+ + DC L+ WK++ + + CR L + F D SF
Sbjct: 264 ---SVLVDLFLTSVILKTDCYNLFVQWKKKHVSASYFGVY--CRTFFLLCACFSFVDLSF 428
Query: 867 IRRTGNSVADALAKRSFTLGVWVWIEEIPQDFYGLVQS 904
+RR GN VA+ LAK +F+ +VWIE+ P L+ S
Sbjct: 429 VRR-GNKVANDLAKLAFSSHGYVWIEDSPIGLVALLSS 539
>AV780250
Length = 494
Score = 68.2 bits (165), Expect = 6e-12
Identities = 43/110 (39%), Positives = 58/110 (52%), Gaps = 12/110 (10%)
Frame = +3
Query: 569 DDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVASSS----------QVPSLPRADWRKLWK 618
+DVL WP +S+G Y++KSGY +R+ + SSS + P+LPR R+L
Sbjct: 171 NDVLAWPHTSNGEYTSKSGYAVVRNKVVHALPSSSTALQVSPVVWKAPTLPRC--RELIG 344
Query: 619 ADAL--LPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFA 666
L LP R+ L ARG+ D CP C E+ LLF P+V WFA
Sbjct: 345 RAFLKILPTRSALRARGMDVDELCPSCETHRESPAQVLLFVPVVSKWWFA 494
>TC10764
Length = 555
Score = 63.9 bits (154), Expect = 1e-10
Identities = 44/112 (39%), Positives = 59/112 (52%), Gaps = 1/112 (0%)
Frame = +1
Query: 763 FDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRR 822
FD ++ D GFGL+A + GEVLAA + L+AE +F WAL M F
Sbjct: 76 FDVSVVEHLDVGFGLLAGDHEGEVLAATSAFPLSVLPPLIAE--AFCWALSSDMTSCFGS 249
Query: 823 VVFEIDCLLLYEAW-KRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNS 873
FEI L LYE W K G S + +L+ DCR L+ FD + SF+ + +S
Sbjct: 250 FCFEIGYLQLYETWMKT*PGSSYVCTLVNDCRLLACYFDFIEPSFVCTSSHS 405
>TC9793
Length = 568
Score = 61.6 bits (148), Expect = 5e-10
Identities = 34/123 (27%), Positives = 57/123 (45%)
Frame = +2
Query: 747 RPNTWSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGL 806
R W P G K+N DA + + GF LVAR+ +L AA LAE L
Sbjct: 50 RRMVWEPPPEGIIKVNIDAGWSGSCSTGFSLVARSHNAVMLVAATHLEETQIEPTLAEAL 229
Query: 807 SFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSF 866
+ RW L + ++ R++ E L++ ++ ++ ++ D R L++ F+ +F
Sbjct: 230 ALRWCLSMIKEMEMERIIVESYSLVVLNTLHQKTKRPDIELVVYDSRELAAEFNFINFKH 409
Query: 867 IRR 869
I R
Sbjct: 410 IGR 418
>TC13259
Length = 506
Score = 60.8 bits (146), Expect = 9e-10
Identities = 39/119 (32%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Frame = +1
Query: 785 EVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWK-RRGGCS 843
+VLA A S S +E ++ RW L L+++LGF V EID ++ +WK + S
Sbjct: 55 QVLATASSKWPRTVSVATSEAMALRWGLQLSLELGFFEVEVEIDNQVVVNSWKGTKKHVS 234
Query: 844 ILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALAKRSFTLGVWVWIEEIPQDFYGLV 902
L ++I D R+S F F +++ R + AD LA+ + + +V IEE P G++
Sbjct: 235 YLANVIADVVRISQCFRFFSLAYVPRLSHLAADYLAQFALSDYCFVGIEEHPAGLAGVL 411
>AV780577
Length = 486
Score = 60.1 bits (144), Expect = 2e-09
Identities = 33/88 (37%), Positives = 48/88 (54%), Gaps = 1/88 (1%)
Frame = +2
Query: 739 GPMVPQVHRPNTWSRPHPGTFKINFDAAMAPTGDA-GFGLVARNSRGEVLAAACSSHGPA 797
GP+ + P+TW P G KIN + P D GFG V R++ G VL + S H +
Sbjct: 200 GPVKGLILGPSTWWPPPHGKLKINIVDVVVPQVDMMGFGFVMRDANGLVLVSGASKHRRS 379
Query: 798 ASSLLAEGLSFRWALGLAMDLGFRRVVF 825
S + E L+ RWA+ ++ DLG R ++F
Sbjct: 380 LSVAMGEALA*RWAMQVSFDLGHRVLMF 463
>CB827358
Length = 453
Score = 58.5 bits (140), Expect = 4e-09
Identities = 36/114 (31%), Positives = 54/114 (46%), Gaps = 6/114 (5%)
Frame = +2
Query: 682 FVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPL------P 735
F+ F+ AD + + A+W A N+L F+E E + A S++ + P
Sbjct: 125 FLLQFLVEADSDVVASCQMASCALWEAHNKLIFQEVPFRCEAAIQRATSMSQVDVSTVDP 304
Query: 736 PLEGPMVPQVHRPNTWSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAA 789
GP+ + W RP G FK+NFD + +G G+VARN G +LAA
Sbjct: 305 SANGPVHDAI-----WRRPPRGVFKVNFDGSWKSDSVSGIGMVARNHDGLILAA 451
>AV768060
Length = 416
Score = 58.5 bits (140), Expect = 4e-09
Identities = 34/82 (41%), Positives = 44/82 (53%)
Frame = -3
Query: 801 LLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFD 860
L+AE + FRWAL LA + G +VF DC L +A+ L ++LDC +L + F
Sbjct: 297 LIAEAMCFRWALNLAANWGLDSLVFSSDCQQLVKAFHCPSEFPSLAHIMLDCTQLVNSFR 118
Query: 861 AFDFSFIRRTGNSVADALAKRS 882
F F F R N VA ALA S
Sbjct: 117 NFSFCFEYRQTNRVAHALAAES 52
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,848,422
Number of Sequences: 28460
Number of extensions: 300723
Number of successful extensions: 1855
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 1814
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1830
length of query: 906
length of database: 4,897,600
effective HSP length: 99
effective length of query: 807
effective length of database: 2,080,060
effective search space: 1678608420
effective search space used: 1678608420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0154.14


