

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
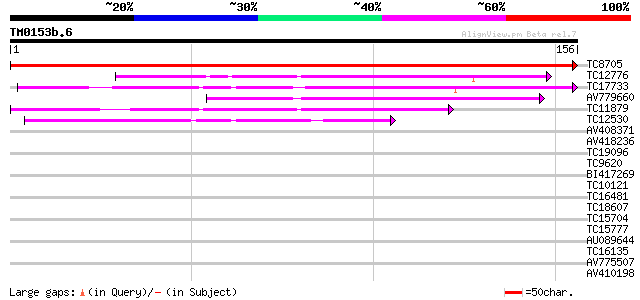
Query= TM0153b.6
(156 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8705 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {Ara... 317 8e-88
TC12776 71 7e-14
TC17733 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (63%) 70 1e-13
AV779660 68 8e-13
TC11879 weakly similar to UP|OREX_HUMAN (O43612) Orexin precurso... 64 2e-11
TC12530 similar to GB|AAB68983.1|500656|YSCH9666 Yhr143wp {Sacch... 39 4e-04
AV408371 35 0.006
AV418236 28 0.93
TC19096 27 1.2
TC9620 similar to UP|Q9ZNX1 (Q9ZNX1) NAD-dependent isocitrate de... 27 1.2
BI417269 27 2.1
TC10121 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (S... 26 2.7
TC16481 similar to UP|O80896 (O80896) At2g32590 protein, partial... 26 2.7
TC18607 similar to UP|O80512 (O80512) At2g44730 protein, partial... 26 2.7
TC15704 similar to UP|O22265 (O22265) Expressed protein (At2g474... 26 3.5
TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing... 25 4.6
AU089644 25 4.6
TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A ... 25 4.6
AV775507 25 4.6
AV410198 25 6.0
>TC8705 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {Arabidopsis
thaliana;}, partial (64%)
Length = 816
Score = 317 bits (811), Expect = 8e-88
Identities = 155/156 (99%), Positives = 155/156 (99%)
Frame = +1
Query: 1 MNPKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN 60
MNPKILILFAL LNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN
Sbjct: 40 MNPKILILFALLLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN 219
Query: 61 QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI 120
QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI
Sbjct: 220 QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI 399
Query: 121 RSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
RSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS
Sbjct: 400 RSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 507
>TC12776
Length = 560
Score = 71.2 bits (173), Expect = 7e-14
Identities = 38/122 (31%), Positives = 67/122 (54%), Gaps = 2/122 (1%)
Frame = +2
Query: 30 STPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSN 89
ST + H L YGFP GL+P+ + + ++ T G F+++L C + D Y+ +
Sbjct: 95 STSTDIHDLLPEYGFPKGLIPNNAI-SYTIS-TDGFFTIQLDSPCYVHFS-DQYLVYFHT 265
Query: 90 TITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDN--IVFEVGMVTAKYPSKNFDDSP 147
+TGK+ G + ++GI+ + F W S+TGI+ D+ + F G ++ K P++ F + P
Sbjct: 266 RLTGKLSYGSVTRVSGIQAQILFLWPSVTGIKVHKDSGMLEFFAGALSQKLPAEEFVNVP 445
Query: 148 AC 149
C
Sbjct: 446 GC 451
>TC17733 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (63%)
Length = 711
Score = 70.5 bits (171), Expect = 1e-13
Identities = 50/157 (31%), Positives = 73/157 (45%), Gaps = 3/157 (1%)
Frame = +1
Query: 3 PKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQT 62
P L + A L S S + +A+ L YG P GLLPDT V G+ ++
Sbjct: 91 PHALFIVATIFTLLSSSLTL------STAEETAYDILPKYGLPSGLLPDT-VTGYTLSD- 246
Query: 63 SGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR- 121
G F V L C I +Y+ Y TITGK+ G I +L GI+V+ W+++ IR
Sbjct: 247 DGRFVVNLAKTCYIKF---DYMVYYEKTITGKLSYGAITDLKGIQVQRLLIWFNVDEIRV 417
Query: 122 --SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
D+I F+VG++ + F +C +SS
Sbjct: 418 DLPPSDSIYFQVGIINKRLDIDQFKSVRSCRKSLASS 528
>AV779660
Length = 559
Score = 67.8 bits (164), Expect = 8e-13
Identities = 31/93 (33%), Positives = 54/93 (57%)
Frame = -1
Query: 55 LGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQW 114
LG+++ + +G+F+V G+C + ++Y +Y TITG I GR+ +L G+ VR W
Sbjct: 544 LGYSLYRQTGEFAVYFDGSCHFNI--ESYRLSYKRTITGVITNGRLYQLKGVSVRVLLLW 371
Query: 115 WSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSP 147
I ++ G ++ F VG+ +A + NF +SP
Sbjct: 370 LDIVEVKIRGGDVEFSVGIASANFGVDNFYESP 272
>TC11879 weakly similar to UP|OREX_HUMAN (O43612) Orexin precursor
(Hypocretin) (Hcrt) [Contains: Orexin-A (Hypocretin-1)
(Hcrt1); Orexin-B (Hypocretin-2) (Hcrt2)], partial (17%)
Length = 683
Score = 63.5 bits (153), Expect = 2e-11
Identities = 40/122 (32%), Positives = 63/122 (50%)
Frame = +1
Query: 1 MNPKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN 60
M +L+L L + L S + P SA+ L + FP GLLP V + ++
Sbjct: 97 MPATLLLLLLLSVPLLSSASSAAAP--------SAYEALAGFNFPAGLLPKG-VTAYELD 249
Query: 61 QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI 120
+++G F L G+C TL +Y +Y++TITG+I R+ +L GI V+ F W +I +
Sbjct: 250 ESTGKFRADLNGSCSFTLE-GSYQLSYNSTITGRISDNRLTDLRGISVKVLFFWLNILEV 426
Query: 121 RS 122
S
Sbjct: 427 GS 432
>TC12530 similar to GB|AAB68983.1|500656|YSCH9666 Yhr143wp {Saccharomyces
cerevisiae;} , partial (5%)
Length = 464
Score = 38.9 bits (89), Expect = 4e-04
Identities = 32/102 (31%), Positives = 44/102 (42%)
Frame = +3
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
I L L L TS S + S ++ + L NYG P+GL P V F V+ G
Sbjct: 171 IQFLLPLLLLSTSSSSSSFASASTSSNATTIYEVLRNYGLPMGLFP-KGVKDFEVDD-DG 344
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGI 106
F V L AC + + Y ++G + KG I L G+
Sbjct: 345 RFWVHLDQACNAKFENELH---YDRNVSGSLSKGMIDALTGL 461
>AV408371
Length = 407
Score = 35.0 bits (79), Expect = 0.006
Identities = 24/69 (34%), Positives = 34/69 (48%)
Frame = +3
Query: 38 ELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVK 97
EL G PVGLLP + + +N TSG+F V + AC + + Y + I G +
Sbjct: 210 ELKAQGLPVGLLP-KGITKYEINGTSGEFVVWMEEACNAKFENEVH---YDSNIKGTLGF 377
Query: 98 GRIAELNGI 106
G I L G+
Sbjct: 378 GWIKGLTGM 404
>AV418236
Length = 411
Score = 27.7 bits (60), Expect = 0.93
Identities = 18/54 (33%), Positives = 27/54 (49%)
Frame = +1
Query: 24 NPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKIT 77
NP+ P +PSS+H ELT G LP+ ++ + TS LG C ++
Sbjct: 82 NPRLPSPSPSSSHSELT--GRNPSTLPEPLLVSSSCPSTS-SLVTGLGRCCLLS 234
>TC19096
Length = 370
Score = 27.3 bits (59), Expect = 1.2
Identities = 17/55 (30%), Positives = 26/55 (46%)
Frame = +2
Query: 23 PNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKIT 77
P P PP + A + +++ FP+GL P AV F+ + GAC +T
Sbjct: 176 PPPLPPLKSKFPALLH-SHHSFPIGLKPKPNFTCQAVFSDDAPFAAAI-GACMLT 334
>TC9620 similar to UP|Q9ZNX1 (Q9ZNX1) NAD-dependent isocitrate
dehydrogenase precursor , partial (41%)
Length = 947
Score = 27.3 bits (59), Expect = 1.2
Identities = 12/27 (44%), Positives = 14/27 (51%)
Frame = -1
Query: 24 NPKPPKSTPSSAHIELTNYGFPVGLLP 50
NPKPP + PSS H T+ P P
Sbjct: 374 NPKPPPAPPSSCHGHRTSRSTPARASP 294
>BI417269
Length = 460
Score = 26.6 bits (57), Expect = 2.1
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 22 NPNPKPPKSTPSSAHIELTNYGFPVGLLPDTT 53
+PNP STPS + + T+ G P LP T
Sbjct: 210 SPNPSTLPSTPSPSTLPTTSSGHPSRALPIPT 305
>TC10121 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (Sucrose-UDP
glucosyltransferase 2) , partial (41%)
Length = 996
Score = 26.2 bits (56), Expect = 2.7
Identities = 14/38 (36%), Positives = 18/38 (46%)
Frame = -3
Query: 30 STPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFS 67
S P HI +T+Y + P VL A QTS F+
Sbjct: 802 SVPGLFHINITSYDDKIDQFPQFVVLRIAFYQTSYVFN 689
>TC16481 similar to UP|O80896 (O80896) At2g32590 protein, partial (19%)
Length = 888
Score = 26.2 bits (56), Expect = 2.7
Identities = 20/57 (35%), Positives = 25/57 (43%), Gaps = 2/57 (3%)
Frame = +3
Query: 17 SKSFQNPNPKPPKST--PSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLG 71
SKS P +PP T P H E + + LLP+ LG + SGD S G
Sbjct: 27 SKSLLLPENRPPCVTKLPEDCHYEPQDL-VKLFLLPNVKCLGRRARKLSGDVSEEQG 194
>TC18607 similar to UP|O80512 (O80512) At2g44730 protein, partial (16%)
Length = 516
Score = 26.2 bits (56), Expect = 2.7
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -2
Query: 17 SKSFQNPNPKPPKSTPSS 34
S SFQ+P+P P S PS+
Sbjct: 113 SASFQSPSPSPSSSAPSA 60
>TC15704 similar to UP|O22265 (O22265) Expressed protein
(At2g47450/T30B22.25), partial (25%)
Length = 664
Score = 25.8 bits (55), Expect = 3.5
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Frame = -3
Query: 9 FALFLNLTSKS-FQNPNPKPPKSTPSSAHIELTNYGFPVGL 48
F L N++S S + N P +TPSS + GFP G+
Sbjct: 134 FPLRSNISSTSAYSNTAPSKTLTTPSSPTLRPNCIGFPFGV 12
>TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (34%)
Length = 566
Score = 25.4 bits (54), Expect = 4.6
Identities = 13/27 (48%), Positives = 16/27 (59%), Gaps = 1/27 (3%)
Frame = +3
Query: 16 TSKSFQNPNPKPPKSTPS-SAHIELTN 41
T SF+ P P +TPS S HI+ TN
Sbjct: 42 TKASFRASTPHPQHTTPSPSYHIQATN 122
>AU089644
Length = 436
Score = 25.4 bits (54), Expect = 4.6
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -3
Query: 6 LILFALFLNLTSKSFQNPNPKPPKSTPSSAH 36
L+ F ++ + TSK +Q+ P PP S SS H
Sbjct: 122 LLSFHIYKHRTSKWYQSNTPVPP-SVTSSCH 33
>TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A apoprotein
A2 (PsaB) (PSI-B), complete
Length = 2647
Score = 25.4 bits (54), Expect = 4.6
Identities = 10/28 (35%), Positives = 16/28 (56%)
Frame = +1
Query: 98 GRIAELNGIRVRAFFQWWSITGIRSSGD 125
G + +N I +QWW G+R++GD
Sbjct: 322 GALGPVN-IAYSGVYQWWYTIGLRTNGD 402
>AV775507
Length = 274
Score = 25.4 bits (54), Expect = 4.6
Identities = 14/44 (31%), Positives = 20/44 (44%)
Frame = +2
Query: 11 LFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTV 54
L L +S P P PPKST SS + + + LP + +
Sbjct: 125 LLLPRSSPPSSQPLPPPPKSTSSSDQSQSQSQNLSLTPLPQSPI 256
>AV410198
Length = 337
Score = 25.0 bits (53), Expect = 6.0
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 3 PKILILFALFLNLTSKSFQNPNPKP 27
P++ L L+ +S F +PNPKP
Sbjct: 44 PQLTSPLLLLLHCSSSLFNSPNPKP 118
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,812,203
Number of Sequences: 28460
Number of extensions: 39311
Number of successful extensions: 441
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 428
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 436
length of query: 156
length of database: 4,897,600
effective HSP length: 83
effective length of query: 73
effective length of database: 2,535,420
effective search space: 185085660
effective search space used: 185085660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0153b.6


