

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
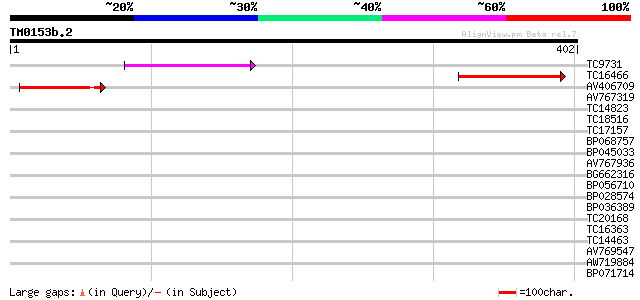
TC9731 81 4e-16
TC16466 weakly similar to UP|TDE2_MOUSE (Q9QZI8) Tumor different... 78 2e-15
AV406709 70 6e-13
AV767319 29 1.6
TC14823 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF... 28 2.1
TC18516 weakly similar to GB|BAB85647.1|19032343|AB076907 inflor... 28 2.1
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 28 2.1
BP068757 28 3.6
BP045033 28 3.6
AV767936 28 3.6
BG662316 28 3.6
BP056710 28 3.6
BP028574 28 3.6
BP036389 27 4.6
TC20168 27 4.6
TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete 27 6.1
TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein (At2g... 27 6.1
AV769547 27 6.1
AW719884 27 6.1
BP071714 27 6.1
>TC9731
Length = 662
Score = 80.9 bits (198), Expect = 4e-16
Identities = 36/93 (38%), Positives = 56/93 (59%)
Frame = +1
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSE 141
T+ VLRVS+G FLFF ++ + RD H G W +KI+ W + IF F LP+E
Sbjct: 358 TDAVLRVSLGNFLFFTILAVLMVGVKNQKDPRDGLHHGGWMMKIICWFLLVIFMFFLPNE 537
Query: 142 LIDLYGEVAHFGAGVFLFIQLISIISFITWLND 174
+I Y ++ FG+G+FL +Q++ ++ F+ ND
Sbjct: 538 IISFYETISKFGSGMFLLVQVVLLLDFVHGWND 636
>TC16466 weakly similar to UP|TDE2_MOUSE (Q9QZI8) Tumor differentially
expressed protein 2 (Tumor differentially expressed 1
protein like) (Membrane protein TMS-2) (Axotomy induced
glyco/Golgi protein 2), partial (6%)
Length = 584
Score = 78.2 bits (191), Expect = 2e-15
Identities = 33/76 (43%), Positives = 46/76 (60%)
Frame = +1
Query: 319 DKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEW 378
+K + V Y Y FFH +F+ +MY AMLL GW++ +DVGW S WVRIV W
Sbjct: 115 EKNEKAKPVTYSYSFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIVTCW 294
Query: 379 LAVCVYLWMLIAPIIW 394
++LW L+API++
Sbjct: 295 ATAILFLWSLVAPIMF 342
>AV406709
Length = 430
Score = 70.1 bits (170), Expect = 6e-13
Identities = 36/61 (59%), Positives = 44/61 (72%)
Frame = +1
Query: 8 SSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKL 67
S+N + K +SW QFRNASNP MARYVYAL+FLV NLLAWA+RD GR LT++
Sbjct: 250 SNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDY--GRGALTEM 423
Query: 68 K 68
+
Sbjct: 424 E 426
>AV767319
Length = 579
Score = 28.9 bits (63), Expect = 1.6
Identities = 19/69 (27%), Positives = 31/69 (44%)
Frame = +2
Query: 155 GVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQ 214
G+ + ++LI LN+ F + C VMLF+T Y +C MY+ +
Sbjct: 218 GMKVLLRLIP*TYLEKKLNEKFRHSTPHQLCNCEVMLFSTG*YCVC------MYLLAFSE 379
Query: 215 PSCLLNISF 223
P C + S+
Sbjct: 380 PGCSSHWSY 406
>TC14823 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF|01169
Uncharacterized (transmembrane domain) protein family.
{Arabidopsis thaliana;}, complete
Length = 1231
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/39 (33%), Positives = 19/39 (48%)
Frame = -1
Query: 274 CIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKC 312
C+R++ S N T W I S G ++ F+ D KC
Sbjct: 673 CLRQNPSKNHTKWVLIRSQTYGSQLALVTPFTKKCDRKC 557
>TC18516 weakly similar to GB|BAB85647.1|19032343|AB076907 inflorescence and
root apices receptor-like kinase {Arabidopsis
thaliana;}, partial (7%)
Length = 654
Score = 28.5 bits (62), Expect = 2.1
Identities = 12/34 (35%), Positives = 21/34 (61%), Gaps = 2/34 (5%)
Frame = -3
Query: 196 SYFICMVGVILMYIWYAPQPSCLLN--ISFITFT 227
SYF C + ++ + W+ PS LL+ +SF +F+
Sbjct: 574 SYFTCFLFLLFILTWFISNPSFLLSHFVSFYSFS 473
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein 1-like
protein 3, partial (81%)
Length = 1298
Score = 28.5 bits (62), Expect = 2.1
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = +3
Query: 321 PAEEDDVPYGYGFFHFVFATGAMYFAMLLV 350
P + D +P YGFFHF F YF+ML++
Sbjct: 1134 PCDVDFIPL-YGFFHFFFIES--YFSMLIL 1214
>BP068757
Length = 519
Score = 27.7 bits (60), Expect = 3.6
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = -2
Query: 114 DTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFGA 154
D W WW ++ L +I P+LLP ++I LY + H G+
Sbjct: 170 DDW---WWGVEFWLSD**SIQPYLLPLDMIVLY--IKHIGS 63
>BP045033
Length = 434
Score = 27.7 bits (60), Expect = 3.6
Identities = 11/30 (36%), Positives = 18/30 (59%)
Frame = -3
Query: 373 RIVNEWLAVCVYLWMLIAPIIWKNIHTEST 402
R + WLA+ V++ +L PI W ++H T
Sbjct: 237 RHLRVWLAITVHIPLLAKPIAWCSLHAVVT 148
>AV767936
Length = 514
Score = 27.7 bits (60), Expect = 3.6
Identities = 14/47 (29%), Positives = 22/47 (46%)
Frame = -3
Query: 250 PGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGI 296
PGL G YV+ + PEG C + D + D + + + VG+
Sbjct: 467 PGLPGCYVLHTLNATVEGLPEGDFCTYEHDEYSCLDDKKVANQAVGV 327
>BG662316
Length = 346
Score = 27.7 bits (60), Expect = 3.6
Identities = 13/44 (29%), Positives = 24/44 (54%)
Frame = -1
Query: 220 NISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWC 263
N S+ FT++LL +T+++ P +++ I GLY+ C
Sbjct: 259 NTSYPKFTILLLHSLTALTTFP*LSSNICKTLPSGLYLTIAACC 128
>BP056710
Length = 486
Score = 27.7 bits (60), Expect = 3.6
Identities = 10/30 (33%), Positives = 14/30 (46%)
Frame = +1
Query: 356 HSMKKWTIDVGWTSAWVRIVNEWLAVCVYL 385
H K W D G T V + W+A +Y+
Sbjct: 316 HQQKLWPTDTGITMLLVMMTKSWMASMIYM 405
>BP028574
Length = 454
Score = 27.7 bits (60), Expect = 3.6
Identities = 15/51 (29%), Positives = 25/51 (48%)
Frame = +1
Query: 78 DCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLW 128
D L +N ++R M F +F + + K ++ R WH GW I+ +W
Sbjct: 292 DPLLSNLLMRSCMVKFKKLRTIFSESFGSQKHHQIRLPWHGGWVLIEFHVW 444
>BP036389
Length = 439
Score = 27.3 bits (59), Expect = 4.6
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = -2
Query: 265 IRSEPEGYECIRKSDSPNKTDW 286
+ S PEGY R+ PN+ DW
Sbjct: 426 LASPPEGYGVGRRKSDPNEIDW 361
>TC20168
Length = 937
Score = 27.3 bits (59), Expect = 4.6
Identities = 19/58 (32%), Positives = 27/58 (45%)
Frame = +2
Query: 247 ILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATF 304
+ S L+ +YV LCW IR R +S T W+ +I I+ L+I TF
Sbjct: 698 VFSTTLLCIYVFALCWVYIR---------RIMESEETTIWKAMIKTPASIV-LIIYTF 841
>TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete
Length = 2164
Score = 26.9 bits (58), Expect = 6.1
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = +1
Query: 127 LWVAVTIFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLN 173
LW VT L ++ ++HF L +Q+ S++SF T L+
Sbjct: 730 LWPLVTFLAMRLEHTVVGTGFLLSHFPLHALLVVQISSLLSFSTLLS 870
>TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein
(At2g16660/T24I21.7), partial (40%)
Length = 1122
Score = 26.9 bits (58), Expect = 6.1
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 6/43 (13%)
Frame = +1
Query: 195 ASYF-IC-----MVGVILMYIWYAPQPSCLLNISFITFTLVLL 231
ASYF +C +V V L+ + P PS L++ F+ LVLL
Sbjct: 739 ASYFGVCNAVAVVVAVYLLAYGFVPSPSSLVSRVFVAVLLVLL 867
>AV769547
Length = 416
Score = 26.9 bits (58), Expect = 6.1
Identities = 10/36 (27%), Positives = 21/36 (57%)
Frame = +2
Query: 38 YVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC 73
Y +++ C+L + SR+ +P +++ K +KTC
Sbjct: 194 YFLSIVISACDLENFGSREHIP*KAIRDTEKKIKTC 301
>AW719884
Length = 373
Score = 26.9 bits (58), Expect = 6.1
Identities = 9/34 (26%), Positives = 19/34 (55%)
Frame = +3
Query: 249 SPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPN 282
SP L+++F CW + + +G++C+ + N
Sbjct: 204 SPSAQVLHILFGCWESAEDKFDGFDCVSAGNCYN 305
>BP071714
Length = 411
Score = 26.9 bits (58), Expect = 6.1
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Frame = +1
Query: 184 RCQIHVMLF---ATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLH 240
R QI M+F +V LM+IW+ + CLL + + + L+ MTS+ +H
Sbjct: 49 RIQIKYMMFNFLTACKIKFSLVRSTLMFIWFLQEFPCLL*V----LSQMHLECMTSIFVH 216
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,519,658
Number of Sequences: 28460
Number of extensions: 159556
Number of successful extensions: 1151
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1138
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1151
length of query: 402
length of database: 4,897,600
effective HSP length: 92
effective length of query: 310
effective length of database: 2,279,280
effective search space: 706576800
effective search space used: 706576800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0153b.2


