

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.11
(156 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
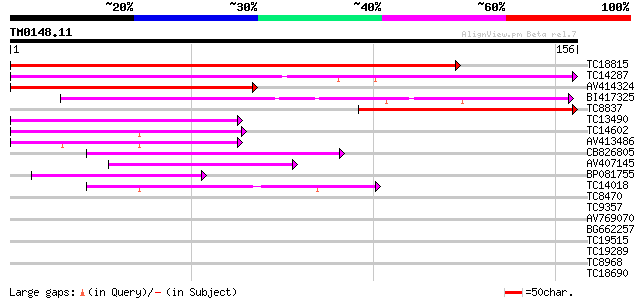
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 246 1e-66
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 132 2e-32
AV414324 86 3e-18
BI417325 85 6e-18
TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein ki... 57 1e-09
TC13490 similar to UP|Q8LPG6 (Q8LPG6) Serine/threonine-protein k... 50 1e-07
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 44 1e-05
AV413486 44 1e-05
CB826805 44 1e-05
AV407145 42 6e-05
BP081755 40 2e-04
TC14018 similar to UP|Q43465 (Q43465) Protein kinase 2, partial ... 39 5e-04
TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase,... 36 0.003
TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein k... 35 0.004
AV769070 30 0.14
BG662257 30 0.19
TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, pa... 29 0.32
TC19289 similar to GB|AAK15470.1|13194578|AF308596 endoribonucle... 29 0.32
TC8968 homologue to UP|Q41619 (Q41619) Protein kinase (Fragment)... 29 0.42
TC18690 similar to UP|P93470 (P93470) Cholinephosphate cytidylyl... 28 0.55
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 246 bits (629), Expect = 1e-66
Identities = 123/124 (99%), Positives = 123/124 (99%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPN RITISSIM
Sbjct: 608 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNRRITISSIM 787
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS
Sbjct: 788 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 967
Query: 121 GFFE 124
GFFE
Sbjct: 968 GFFE 979
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial (75%)
Length = 1999
Score = 132 bits (333), Expect = 2e-32
Identities = 76/166 (45%), Positives = 98/166 (58%), Gaps = 10/166 (6%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL+G LPFQ EN+M +Y K +AE++FP W SP++K LIS +LVADP R +I I+
Sbjct: 1002 LYALLSGYLPFQGENVMRIYRKAFKAEYEFPEWISPQAKNLISNLLVADPEKRYSIPEII 1181
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNS-----SSEQSTKVVA-----AAKFINAFEFI 110
WFQ GF + +ES D E ++VA A F NAFE I
Sbjct: 1182 SDPWFQYGFMRPLAF-SINESAVGEDSTI*SFWWGGEXVPELVAAMGKPARPFYNAFEII 1358
Query: 111 SSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SS+S GFDL FE ++R S+F SK S S + +K+EG AK L FR
Sbjct: 1359 SSLSHGFDLRSLFETRKRSPSMFISKYSASAVVAKLEGMAKKLNFR 1496
>AV414324
Length = 380
Score = 85.9 bits (211), Expect = 3e-18
Identities = 38/68 (55%), Positives = 52/68 (75%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF+ NLM MY K+ +AEF+FP WF+P+ ++L+SKIL +P RI+++ IM
Sbjct: 27 LYVLLAGYLPFRDSNLMEMYRKIGKAEFKFPNWFAPDVRRLLSKILDPNPRTRISLAKIM 206
Query: 61 RVSWFQKG 68
SWF+KG
Sbjct: 207ESSWFRKG 230
>BI417325
Length = 449
Score = 84.7 bits (208), Expect = 6e-18
Identities = 59/149 (39%), Positives = 83/149 (55%), Gaps = 8/149 (5%)
Frame = +2
Query: 15 NLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIMRVSWFQKGFSASIP 74
NLM +Y K+ AEF PPW S ++KLI++IL +P RITI+ I+ WF+K + I
Sbjct: 5 NLMELYKKISAAEFTCPPWLSFSARKLITRILDPNPMTRITIAEILEDEWFKKDYKPPI- 181
Query: 75 IPDPDESNFNSDLNSSSEQSTKVVAAAK------FINAFEFISSMSSGFDLSGFF--EEK 126
+ E+N + D+ + + S + K +NAFE I SMS G +L F E+
Sbjct: 182 FQENGETNLD-DVEAVFKDSEEHHVTEKKEEQPTAMNAFELI-SMSKGLNLENLFDVEQG 355
Query: 127 RRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+ + FTSK +EI SKIE AAK L F
Sbjct: 356 FKRETRFTSKSPANEIISKIEEAAKPLGF 442
>TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(15%)
Length = 606
Score = 57.4 bits (137), Expect = 1e-09
Identities = 30/60 (50%), Positives = 36/60 (60%)
Frame = +1
Query: 97 VVAAAKFINAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
V AA NAFE ISS+S GFDL FE ++R S F SK S S + +K+E AK L R
Sbjct: 37 VEAARPSYNAFEIISSLSHGFDLRNLFETRKRSPSTFISKLSASAVVAKLEAVAKKLNLR 216
>TC13490 similar to UP|Q8LPG6 (Q8LPG6) Serine/threonine-protein kinase-like
protein, partial (31%)
Length = 614
Score = 50.4 bits (119), Expect = 1e-07
Identities = 23/64 (35%), Positives = 36/64 (55%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY ++ G PF + L Y+K++ + P +P+ KKLI +L DP LR+T+S +
Sbjct: 141 LYCMIVGEYPFLGDTLQDTYDKIVNNPIELPNDMNPQLKKLIEGLLFKDPTLRMTLSDVA 320
Query: 61 RVSW 64
SW
Sbjct: 321 VDSW 332
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 44.3 bits (103), Expect = 1e-05
Identities = 22/69 (31%), Positives = 37/69 (52%), Gaps = 4/69 (5%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWF----SPESKKLISKILVADPNLRITI 56
LY +L+G PF ++ ++ V+R +FP SP +K L+ K++ DP+ RI+
Sbjct: 638 LYIMLSGTPPFYGDSAAEIFEAVIRGNLRFPSRIFRNVSPAAKDLLRKMICRDPSNRISA 817
Query: 57 SSIMRVSWF 65
+R WF
Sbjct: 818 EQALRHPWF 844
>AV413486
Length = 402
Score = 43.9 bits (102), Expect = 1e-05
Identities = 25/70 (35%), Positives = 37/70 (52%), Gaps = 6/70 (8%)
Frame = +3
Query: 1 LYALLAGCLPFQH----ENLMTMYNKVLRAEFQFPPWF--SPESKKLISKILVADPNLRI 54
LY +L G PF+ ++ +VL ++ P + SPE + LIS+I V DP RI
Sbjct: 192 LYVMLVGTYPFEDPDDPKDFRKTIQRVLTVQYSIPDFVQVSPECRHLISRIFVFDPAERI 371
Query: 55 TISSIMRVSW 64
TI I++ W
Sbjct: 372 TIPEILKNEW 401
>CB826805
Length = 635
Score = 43.9 bits (102), Expect = 1e-05
Identities = 19/71 (26%), Positives = 38/71 (52%)
Frame = +2
Query: 22 KVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIMRVSWFQKGFSASIPIPDPDES 81
++ + + Q P W S ++ +I +IL +P RIT++ I WF++ ++ + P + D+
Sbjct: 2 QIFKGDVQIPKWLSAGAQNIIKRILDPNPETRITMAGIKEDPWFKQDYTPADPDDEEDDV 181
Query: 82 NFNSDLNSSSE 92
+ D S E
Sbjct: 182YIDDDAFSIHE 214
Score = 30.0 bits (66), Expect = 0.19
Identities = 16/25 (64%), Positives = 19/25 (76%)
Frame = +3
Query: 104 INAFEFISSMSSGFDLSGFFEEKRR 128
INAF+ I MSS DLSGFFE++ R
Sbjct: 396 INAFQLIG-MSSFLDLSGFFEQEVR 467
>AV407145
Length = 232
Score = 41.6 bits (96), Expect = 6e-05
Identities = 19/52 (36%), Positives = 28/52 (53%)
Frame = +1
Query: 28 FQFPPWFSPESKKLISKILVADPNLRITISSIMRVSWFQKGFSASIPIPDPD 79
+ P SP ++ LI ++LV DP R+TI I + WFQ + +P PD
Sbjct: 7 YTLPSHLSPGARDLIPRMLVVDPMKRMTIPEIRQHPWFQARLPRYLAVPPPD 162
>BP081755
Length = 518
Score = 39.7 bits (91), Expect = 2e-04
Identities = 19/48 (39%), Positives = 26/48 (53%)
Frame = -1
Query: 7 GCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRI 54
G PF E+ +Y K+L FP + PE+K LI K+L AD R+
Sbjct: 515 GYPPFFDESPFRIYEKILEGRVDFPRFVDPEAKDLIKKLLTADRTKRL 372
>TC14018 similar to UP|Q43465 (Q43465) Protein kinase 2, partial (27%)
Length = 559
Score = 38.5 bits (88), Expect = 5e-04
Identities = 26/87 (29%), Positives = 44/87 (49%), Gaps = 6/87 (6%)
Frame = +3
Query: 22 KVLRAEFQFPPWF--SPESKKLISKILVADPNLRITISSIMRVSWFQKGFSASIPIPDPD 79
+++ ++ P + S E + L+S+I VADP RITI+ I + WF K S I + +
Sbjct: 18 RIIGVQYSIPDYVRVSAECRNLLSRIFVADPAKRITIAEIKQNPWFLK--SLPKEIIEAE 191
Query: 80 ESNF----NSDLNSSSEQSTKVVAAAK 102
F L+ S E+ ++V A+
Sbjct: 192 TKGFVETKKDQLSQSVEEIMRIVQEAR 272
>TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(29%)
Length = 807
Score = 35.8 bits (81), Expect = 0.003
Identities = 23/67 (34%), Positives = 35/67 (51%), Gaps = 3/67 (4%)
Frame = +1
Query: 92 EQSTKVVAAAKFINAFEFISSMSSGFDLSGFFEEKRRGGSV---FTSKCSVSEIASKIEG 148
++ +V +NAF IS +S GFDLS FEE++R F + S + S++E
Sbjct: 16 QEQEQVQEQPSTMNAFHIIS-LSEGFDLSPLFEERKREEKEEMRFATTRPASSVISRLEE 192
Query: 149 AAKGLRF 155
AK +F
Sbjct: 193 VAKAGKF 213
>TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein kinase,
41K) , partial (71%)
Length = 1181
Score = 35.4 bits (80), Expect = 0.004
Identities = 21/56 (37%), Positives = 29/56 (51%), Gaps = 6/56 (10%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNK----VLRAEFQFPPW--FSPESKKLISKILVADP 50
LY +L G PF+ N + K VL ++ P + SPE + LIS+I V DP
Sbjct: 921 LYVMLVGAYPFEDPNEPKDFRKTIQRVLSVQYSIPDFVQISPECRHLISRIFVFDP 1088
>AV769070
Length = 425
Score = 30.4 bits (67), Expect = 0.14
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = -3
Query: 35 SPESKKLISKILVADPNLRITISSIM 60
SPE+ L+ K+LV DPN RIT+ +
Sbjct: 423 SPEAVDLLEKMLVFDPNKRITVDEAL 346
>BG662257
Length = 336
Score = 30.0 bits (66), Expect = 0.19
Identities = 12/36 (33%), Positives = 21/36 (58%)
Frame = +1
Query: 83 FNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFD 118
F + + +++AAA+ NA +FISS+ G+D
Sbjct: 4 FGTRKGGDAATEAEIIAAAELANAHKFISSLQQGYD 111
>TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, partial (3%)
Length = 504
Score = 29.3 bits (64), Expect = 0.32
Identities = 13/39 (33%), Positives = 21/39 (53%)
Frame = -3
Query: 63 SWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAA 101
SW S S+P P P + +S +SSS S+ ++A+
Sbjct: 193 SWLMSSLSLSLPRPGPKRGSSSSPSSSSSSSSSSSLSAS 77
>TC19289 similar to GB|AAK15470.1|13194578|AF308596 endoribonuclease/protein
kinase IRE1-like protein {Arabidopsis thaliana;},
partial (18%)
Length = 470
Score = 29.3 bits (64), Expect = 0.32
Identities = 22/64 (34%), Positives = 31/64 (48%)
Frame = +1
Query: 2 YALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIMR 61
+ L G PF E+L N V + F F PE++ LIS +L DPNLR ++
Sbjct: 199 FCLTGGRHPFG-EHLERDVNVVKNRKDLFLVEFIPEAEDLISCLLNPDPNLRPKAIEVLH 375
Query: 62 VSWF 65
+F
Sbjct: 376 HPFF 387
>TC8968 homologue to UP|Q41619 (Q41619) Protein kinase (Fragment),
partial (32%)
Length = 612
Score = 28.9 bits (63), Expect = 0.42
Identities = 23/75 (30%), Positives = 32/75 (42%), Gaps = 16/75 (21%)
Frame = +1
Query: 27 EFQFP-----PWFS-------PESKKLISKILVADPNLRITISSIMRVSWFQKGFSASIP 74
EF+FP PW PE+ L+S++L PNLR T + +F + S
Sbjct: 19 EFKFPQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCTALDALTHPFFDELREPSTR 198
Query: 75 IPD----PDESNFNS 85
+P P NF S
Sbjct: 199LPTGRFLPPLFNFKS 243
>TC18690 similar to UP|P93470 (P93470) Cholinephosphate cytidylyltransferase
, partial (80%)
Length = 845
Score = 28.5 bits (62), Expect = 0.55
Identities = 21/54 (38%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Frame = +1
Query: 24 LRAEFQFPPWFSPESKKLISKILVADPNL-RITISSIMRVSWFQKGFSASIPIP 76
LR+ +P WFS S+K+ISK ++A L R + S I R + + G IP P
Sbjct: 109 LRSLPLWPRWFSRASQKIISKYILACWLLQR*SHSQIQRQNCYD*G*KIRIPSP 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,494,553
Number of Sequences: 28460
Number of extensions: 29348
Number of successful extensions: 214
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 212
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 212
length of query: 156
length of database: 4,897,600
effective HSP length: 83
effective length of query: 73
effective length of database: 2,535,420
effective search space: 185085660
effective search space used: 185085660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0148.11


