

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0142.11
(270 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
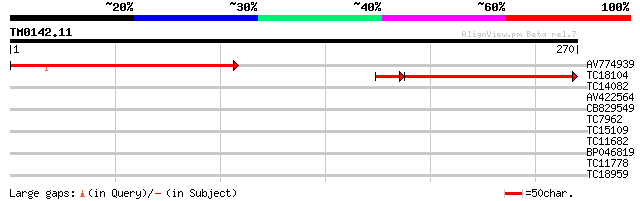
Score E
Sequences producing significant alignments: (bits) Value
AV774939 182 7e-47
TC18104 weakly similar to UP|Q9FPH6 (Q9FPH6) AT3g15840, partial ... 162 5e-44
TC14082 similar to UP|Q7X9A0 (Q7X9A0) RuBisCO activase alpha for... 36 0.008
AV422564 27 4.8
CB829549 27 4.8
TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 ... 26 6.3
TC15109 similar to UP|Q9FVE9 (Q9FVE9) Cytosolic aconitase, parti... 26 6.3
TC11682 homologue to UP|Q7PZ83 (Q7PZ83) AgCP9540 (Fragment), par... 26 6.3
BP046819 26 8.2
TC11778 weakly similar to UP|O80836 (O80836) At2g45830 protein, ... 26 8.2
TC18959 similar to UP|AAR07599 (AAR07599) Fiber protein Fb37, pa... 26 8.2
>AV774939
Length = 486
Score = 182 bits (461), Expect = 7e-47
Identities = 98/144 (68%), Positives = 100/144 (69%), Gaps = 35/144 (24%)
Frame = +2
Query: 1 MYDSSDLLFFSVPPSN-----------------------------------PITSRHTFL 25
MYDSSDLLF SVPPS P T TFL
Sbjct: 53 MYDSSDLLFLSVPPSKSNNKQAPK*CDT*MAATNASIFASSTTQPCLPIRIPNTLASTFL 232
Query: 26 NVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFELGKASVYWKTMNG 85
NVSLPRCYLVKE VKVTRTI AAVAVATTPAQE+EEYKIP+WANFELGKASVYWKTMNG
Sbjct: 233 NVSLPRCYLVKEGKVKVTRTIKAAVAVATTPAQEMEEYKIPAWANFELGKASVYWKTMNG 412
Query: 86 LPPTSGEKLKLFYNPTSTQLAPNE 109
LPPTSGEKLKLFYNPTSTQLAPNE
Sbjct: 413 LPPTSGEKLKLFYNPTSTQLAPNE 484
>TC18104 weakly similar to UP|Q9FPH6 (Q9FPH6) AT3g15840, partial (33%)
Length = 543
Score = 162 bits (411), Expect(2) = 5e-44
Identities = 76/82 (92%), Positives = 80/82 (96%)
Frame = +1
Query: 189 FFNEGLAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHL 248
FFNEGLAE LG+EGACE AIFPDSNKVIT+CAMLG+LTVEGGDRCDLNLVEGCTDPSSHL
Sbjct: 43 FFNEGLAEGLGQEGACEPAIFPDSNKVITQCAMLGHLTVEGGDRCDLNLVEGCTDPSSHL 222
Query: 249 YNPLANVDDGTCPLDLDSDSEA 270
YNPLA+VDDGTCPLDLDSDSEA
Sbjct: 223 YNPLAHVDDGTCPLDLDSDSEA 288
Score = 31.2 bits (69), Expect(2) = 5e-44
Identities = 13/14 (92%), Positives = 14/14 (99%)
Frame = +2
Query: 175 QFQVPKVLQNKPIE 188
QFQVPKVLQNKPI+
Sbjct: 2 QFQVPKVLQNKPID 43
>TC14082 similar to UP|Q7X9A0 (Q7X9A0) RuBisCO activase alpha form precursor,
partial (90%)
Length = 1667
Score = 35.8 bits (81), Expect = 0.008
Identities = 14/38 (36%), Positives = 23/38 (59%)
Frame = +2
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G + + ++ + EGCTDP++ ++P A DDGTC
Sbjct: 1412 GTFYGKAAQQVNVPVPEGCTDPNAKNFDPTARSDDGTC 1525
>AV422564
Length = 441
Score = 26.6 bits (57), Expect = 4.8
Identities = 26/94 (27%), Positives = 42/94 (44%), Gaps = 3/94 (3%)
Frame = +1
Query: 20 SRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFELGKASVY 79
SRH N+S P C+LV R + +T A+ + + +Q + N + S++
Sbjct: 145 SRHPTRNISPPDCFLV--RRIGKFQTFTASKFMVSCTSQNSQNQN-----NGDEAPESLF 303
Query: 80 WKTM--NGLPPTS-GEKLKLFYNPTSTQLAPNEE 110
K + G+ PTS E K N ++ NEE
Sbjct: 304 MKELKRRGMNPTSLLEDYKQSSNVLDEEVYVNEE 405
>CB829549
Length = 534
Score = 26.6 bits (57), Expect = 4.8
Identities = 13/38 (34%), Positives = 21/38 (55%)
Frame = +2
Query: 156 LNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
L L+ T V DGP R++ ++P+ K +E N+G
Sbjct: 383 LALVRQPTTNVGLDGPMRVKMRLPRAQLEKLMEESNDG 496
>TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 , complete
Length = 1489
Score = 26.2 bits (56), Expect = 6.3
Identities = 21/94 (22%), Positives = 37/94 (39%), Gaps = 7/94 (7%)
Frame = +1
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP E NGG ++ + C G +AM+ D ++ + VP
Sbjct: 841 FVNPKDHDKPVQEVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNQDD 1020
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ 183
H +N + T + G Y+ + +P V++
Sbjct: 1021AFQTHPVNFLNERTLKGTFYGNYKPRTDLPNVVE 1122
>TC15109 similar to UP|Q9FVE9 (Q9FVE9) Cytosolic aconitase, partial (46%)
Length = 1237
Score = 26.2 bits (56), Expect = 6.3
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = -1
Query: 226 TVEGGDRCDLNLVEGCTDPSSHLY 249
T GG+ L LV GCT PS L+
Sbjct: 433 TTRGGEAR*LALVNGCTRPSKFLF 362
>TC11682 homologue to UP|Q7PZ83 (Q7PZ83) AgCP9540 (Fragment), partial (11%)
Length = 822
Score = 26.2 bits (56), Expect = 6.3
Identities = 25/95 (26%), Positives = 36/95 (37%), Gaps = 17/95 (17%)
Frame = +1
Query: 30 PRCYLVKERNVKVTRTINAAVAVATT-----PAQEIEEY---KIPSWA-----NFELGKA 76
P+ + K R K T A AT PA E+ P+W + + A
Sbjct: 97 PQFTVKKNRKWKTTPPTTATATTATVVAPSCPATSSTEWPTGSAPTWLLLSSLPWSVALA 276
Query: 77 SVY----WKTMNGLPPTSGEKLKLFYNPTSTQLAP 107
S+ KT PPT ++ + NP S+ AP
Sbjct: 277 SISAPPTTKTTTTTPPTRDDRALMLTNPVSSSAAP 381
>BP046819
Length = 459
Score = 25.8 bits (55), Expect = 8.2
Identities = 12/37 (32%), Positives = 16/37 (42%)
Frame = +2
Query: 136 DRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPY 172
+R + P +S IC N + S TN DW Y
Sbjct: 158 ERDXSSKPPFSFSICSNSQHGNCVASHTNKSDWVSEY 268
>TC11778 weakly similar to UP|O80836 (O80836) At2g45830 protein, partial
(38%)
Length = 816
Score = 25.8 bits (55), Expect = 8.2
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = -2
Query: 8 LFFSVPPSNPITSRHTFLNVSLPRCYL 34
++FSV P NP S FL PR Y+
Sbjct: 665 IWFSVVPFNPFISLLDFLQQMFPRPYI 585
>TC18959 similar to UP|AAR07599 (AAR07599) Fiber protein Fb37, partial (28%)
Length = 364
Score = 25.8 bits (55), Expect = 8.2
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = +2
Query: 4 SSDLLFFSVPPSNPITSRHTFLNVSL 29
S LLFF P SNP++ FL S+
Sbjct: 92 SDSLLFFVNPQSNPLSLSSIFLRFSI 169
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,836,416
Number of Sequences: 28460
Number of extensions: 64263
Number of successful extensions: 322
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 321
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 322
length of query: 270
length of database: 4,897,600
effective HSP length: 89
effective length of query: 181
effective length of database: 2,364,660
effective search space: 428003460
effective search space used: 428003460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0142.11


