

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.7
(571 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
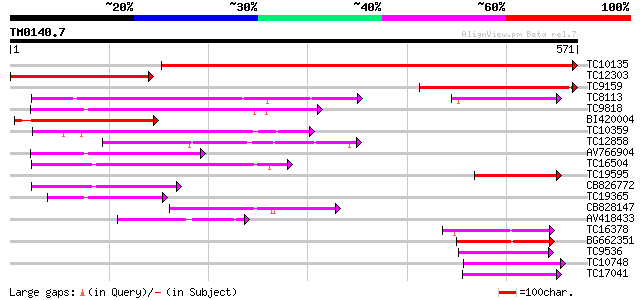
Sequences producing significant alignments: (bits) Value
TC10135 834 0.0
TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transpo... 283 7e-77
TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose tr... 256 5e-69
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 188 2e-48
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 186 7e-48
BI420004 174 3e-44
TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partia... 155 1e-38
TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter,... 137 6e-33
AV766904 127 6e-30
TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporte... 120 5e-28
TC19595 102 1e-22
CB826772 91 4e-19
TC19365 weakly similar to UP|O23213 (O23213) Sugar transporter l... 82 2e-16
CB828147 82 3e-16
AV418433 78 4e-15
TC16378 similar to GB|AAM70554.1|21700861|AY124845 At1g75220/F22... 77 6e-15
BG662351 74 6e-14
TC9536 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, parti... 69 3e-12
TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter,... 68 5e-12
TC17041 homologue to GB|AAL25568.1|16648753|AY058152 AT5g16150/T... 65 4e-11
>TC10135
Length = 1501
Score = 834 bits (2155), Expect = 0.0
Identities = 416/418 (99%), Positives = 417/418 (99%)
Frame = +2
Query: 154 GALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWL 213
GALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWL
Sbjct: 2 GALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWL 181
Query: 214 YRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGL 273
YRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGL
Sbjct: 182 YRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGL 361
Query: 274 YAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFID 333
YAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFID
Sbjct: 362 YAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFID 541
Query: 334 KTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNNTCPDFRAAVNPGRW 393
KTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNNTCPDFRAAVNPGRW
Sbjct: 542 KTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNNTCPDFRAAVNPGRW 721
Query: 394 TCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALV 453
TCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALV
Sbjct: 722 TCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALV 901
Query: 454 GLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIG 513
GLALYIITFSPGMGTVPWVINSEIYP RYRGVCGGIASTTVWISNLIVSQSFLSLTQTIG
Sbjct: 902 GLALYIITFSPGMGTVPWVINSEIYPFRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIG 1081
Query: 514 TAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEKH 571
TAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK+DSGSEKH
Sbjct: 1082TAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKKDSGSEKH 1255
>TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transporter like
protein {Arabidopsis thaliana;}, partial (25%)
Length = 545
Score = 283 bits (723), Expect = 7e-77
Identities = 143/145 (98%), Positives = 144/145 (98%)
Frame = +3
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGVPEADMSAFRECLSLSWKNPYVLR AFSAGIGGLLFGYDTGVISGALLYI+DDFKA
Sbjct: 111 MEGGVPEADMSAFRECLSLSWKNPYVLRRAFSAGIGGLLFGYDTGVISGALLYIKDDFKA 290
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA
Sbjct: 291 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 470
Query: 121 ILLFGRVFVGFGVGMASMASPLYIS 145
ILLFGRVFVGFGVGMASMASPLYIS
Sbjct: 471 ILLFGRVFVGFGVGMASMASPLYIS 545
>TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose transporter
HXT4 (Low-affinity glucose transporter LGT1), partial
(5%)
Length = 728
Score = 256 bits (655), Expect = 5e-69
Identities = 120/160 (75%), Positives = 139/160 (86%), Gaps = 1/160 (0%)
Frame = +3
Query: 413 NKLLPGACLISDDTTKDMCGN-DHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPW 471
+KL PGACL+SDD T+ C HR WYT+GCPSK G+ AL GLALYII FSPGMGTVPW
Sbjct: 3 DKLSPGACLLSDDNTRTQCQKVHHRPWYTQGCPSKLGFLALGGLALYIIFFSPGMGTVPW 182
Query: 472 VINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFF 531
V+NSEIYPLRYRG+CGGIASTTVW+SNL+VSQSFLSLTQTIGTAWTFMMFGI+A++ IFF
Sbjct: 183 VVNSEIYPLRYRGICGGIASTTVWVSNLVVSQSFLSLTQTIGTAWTFMMFGIIAIIGIFF 362
Query: 532 VIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEKH 571
V++FVPETKGVPMEEVE ML +R++ KFW+K SG EK+
Sbjct: 363 VLIFVPETKGVPMEEVESMLNKRAVHLKFWEK-TSGGEKN 479
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (93%)
Length = 1865
Score = 188 bits (477), Expect = 2e-48
Identities = 122/350 (34%), Positives = 181/350 (50%), Gaps = 17/350 (4%)
Frame = +2
Query: 23 NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIG 82
N Y L A + ++ GYDTGV+SGALL+I++D D + QE + A++G
Sbjct: 107 NKYALACVIVASMVSIISGYDTGVMSGALLFIKEDIGISDTQ---QEVLAGILNICALVG 277
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
G +D GRR I +A LFL+G+V M PN AIL+FGR G GVG A +P+
Sbjct: 278 CLAAGKTSDYIGRRYTIFLASILFLVGAVFMGYGPNFAILMFGRCVCGLGVGFALTTAPV 457
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT--WRWMLGVAAAPAIIQI 200
Y +E S RG L SL I G F+ Y+ N T WR MLG+AA P++
Sbjct: 458 YSAELSSASTRGFLTSLPEVCIGLGIFIGYISNYFLGKLALTLGWRLMLGLAAIPSLGLA 637
Query: 201 VLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKI-- 258
+ +L++PESPRWL +GR +K +L ++ + +AE+ V + +E +
Sbjct: 638 LGILTMPESPRWLVMQGRLGCAKKVLLEVSNTTE-EAELRFRDIIVSAGFDEKCNDEFVK 814
Query: 259 ------------SMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFAS 306
+ L T VRR L A +G+ FF+ GI VM Y P I + AG +
Sbjct: 815 QPQKSHHGEGVWKELFLRPTPPVRRMLIAAVGIHFFEHATGIEAVMLYGPRIFKKAG-VT 991
Query: 307 NRTALLLSLITSGLNAFGSI-LSIYFIDKTGRKKLALISLCGVVLSLALL 355
+ LLL+ I +GL + +S + +D+ GR++L IS+ G++ L +L
Sbjct: 992 TKDRLLLATIGTGLTKITFLTISTFLLDRVGRRRLLQISVAGMIFGLTIL 1141
Score = 68.2 bits (165), Expect = 4e-12
Identities = 39/113 (34%), Positives = 59/113 (51%), Gaps = 3/113 (2%)
Frame = +2
Query: 446 KFGWA---ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVS 502
K WA ++V Y+ F+ G+ V WV +SEI+PLR R I N +S
Sbjct: 1178 KLVWALSLSIVATYTYVAFFNVGLAPVTWVYSSEIFPLRLRAQGNSIGVAVNRGMNAAIS 1357
Query: 503 QSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
SF+S+ + I F +F ++VVA F +PETKG +EE+E + ++S
Sbjct: 1358 MSFISIYKAITIGGAFFLFAGMSVVAWVFFYFCLPETKGKALEEMEMVFSKKS 1516
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 186 bits (473), Expect = 7e-48
Identities = 111/308 (36%), Positives = 167/308 (54%), Gaps = 14/308 (4%)
Frame = +2
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
+N Y A A + +L GYD GV+SGA +YI+ D K D K + I++ ++I
Sbjct: 224 RNKYAFACAMLASMTSILLGYDIGVMSGAAIYIKRDLKVSDVKIEILLGIINLY---SLI 394
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ + G +D GRR I+ A +F +G+++M +PN L+FGR G G+G A M +P
Sbjct: 395 GSGLAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYWFLMFGRFIAGIGIGYALMIAP 574
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT--NAPGTWRWMLGVAAAPAIIQ 199
+Y +E SP RG L S I GG L Y+ N AF+ + WR MLGV A P++I
Sbjct: 575 VYTAEVSPASSRGFLTSFPEVFINGGILLGYISNFAFSKLSLKVGWRMMLGVGALPSVIL 754
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKIY-APEDVDAEIEALKES-------VESEIE 251
V +L++PESPRWL +GR ++ +L K +PE+ + +K + + +E
Sbjct: 755 GVGVLAMPESPRWLVMRGRLGDAIKVLNKTSDSPEEAQLRLADIKRAAGIPESCTDDVVE 934
Query: 252 ESKTSK----ISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASN 307
SK S + L T +R + A +G+ FFQQ GI+ V+ YSPTI + AG S+
Sbjct: 935 VSKRSTGEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGIDAVVLYSPTIFEKAGIKSD 1114
Query: 308 RTALLLSL 315
LL ++
Sbjct: 1115TDKLLATV 1138
Score = 26.9 bits (58), Expect = 9.0
Identities = 14/42 (33%), Positives = 24/42 (56%), Gaps = 4/42 (9%)
Frame = -1
Query: 45 GVISGALLYIRD----DFKAVDRKTWLQEAIVSTAIAGAIIG 82
GV +G LL D D+ D + W++E ++ ++AGA +G
Sbjct: 1063 GVDAGGLLEEVDAEGGDYDVADCRGWVEEELLPDSLAGASLG 938
>BI420004
Length = 547
Score = 174 bits (441), Expect = 3e-44
Identities = 86/145 (59%), Positives = 111/145 (76%)
Frame = +1
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE MS F KNPY+L + +AGIGGLLFGYDTGVISGALLYI+DDF+ V R +
Sbjct: 133 PERKMSYF--------KNPYILGVTAAAGIGGLLFGYDTGVISGALLYIKDDFEEVRRSS 288
Query: 66 WLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFG 125
LQE IVS A+ GAIIGAA GGWIND FGR+ A + AD +F +GS++MAAAP+ ++FG
Sbjct: 289 LLQETIVSMALVGAIIGAATGGWINDAFGRKKATLSADVVFTLGSLVMAAAPDAYFVIFG 468
Query: 126 RVFVGFGVGMASMASPLYISEASPT 150
R+ VG G+G+AS+ +P+YI+E+SP+
Sbjct: 469 RLLVGLGIGVASVTAPVYIAESSPS 543
>TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partial (90%)
Length = 1099
Score = 155 bits (393), Expect = 1e-38
Identities = 101/304 (33%), Positives = 150/304 (49%), Gaps = 20/304 (6%)
Frame = +2
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKTWLQEAI--------- 71
P V+ A GGL+FGYD GV G ++ F V +KT L++ +
Sbjct: 203 PIVIISCILAATGGLMFGYDVGVSGGVTSMPHFLHKFFPDVYKKTVLEKELDSNYCKYDN 382
Query: 72 ------VSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFG 125
S+ ++ + GRR+ ++IA F+ G V A N A+L+ G
Sbjct: 383 QGLQLFTSSLYLAGLVSTFFASYTTRTLGRRMTMLIAGVFFIAGVVFNVTAQNLAMLIVG 562
Query: 126 RVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTW 185
R+ +G GVG A+ A PL++SE +P+R+RGAL L IT G + LIN G W
Sbjct: 563 RILLGCGVGFANQAVPLFLSEIAPSRIRGALNILFQLNITIGILFANLINYGTNKIKGGW 742
Query: 186 RW--MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALK 243
W LG+A PA++ V L + ++P L +GR EE K++LKKI ++V+ E L
Sbjct: 743 GWRLSLGLAGIPAVLLTVGALLVVDTPNSLIERGRTEEGKAVLKKIRGTDNVEPEFLELV 922
Query: 244 ESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAG 303
E+ +K K LLK R + + LQ FQQF GIN +M+Y+P + G
Sbjct: 923 EA----SRVAKEVKDPFRNLLKRRN-RPQIIISIALQIFQQFTGINAIMFYAPVLFNTLG 1087
Query: 304 FASN 307
F ++
Sbjct: 1088FKND 1099
>TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter, partial
(51%)
Length = 882
Score = 137 bits (344), Expect = 6e-33
Identities = 85/269 (31%), Positives = 144/269 (52%), Gaps = 8/269 (2%)
Frame = +3
Query: 94 GRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVR 153
G R +I+ F+ G++ A +L+ GR+ +GFG+G A+ + P+Y+SE +P + R
Sbjct: 36 GMRATMIVGGLFFVSGALFNGLADGIWMLIVGRLLLGFGIGCANQSVPIYLSEMAPYKYR 215
Query: 154 GALVSLNSFLITGGQFLSYLINLAFT---NAPGTWRWMLGVAAAPAIIQIVLMLSLPESP 210
G L IT G F++ L N F N G WR LG+ A PA+I +V L LP+SP
Sbjct: 216 GGLNMCFQLSITIGIFVANLFNYYFAKILNGQG-WRLSLGLGAVPAVIFVVGSLCLPDSP 392
Query: 211 RWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVR 270
L +GR E ++ L KI D++AE+ + + + E + K TLL+ R
Sbjct: 393 SSLVARGRHEAARQELVKIRGTTDIEAEL----KDIITASEALENVKHPWKTLLE-RKYR 557
Query: 271 RGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIY 330
L + + FFQQF G+N + +Y+P + + GF +L+ ++I +++SI+
Sbjct: 558 PQLVFAVCIPFFQQFTGLNVITFYAPILFRTIGFGPT-ASLMSAVIIGSFKPVSTLISIF 734
Query: 331 FIDKTGRKKL-----ALISLCGVVLSLAL 354
+DK GR+ L A + +C +++++A+
Sbjct: 735 VVDKFGRRTLFLEGGAQMLICQIIMTIAI 821
>AV766904
Length = 611
Score = 127 bits (318), Expect = 6e-30
Identities = 67/178 (37%), Positives = 102/178 (56%), Gaps = 2/178 (1%)
Frame = +3
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
+N + L A A + +L GYD GV+SGA++YI+ D K D + + I++ + I
Sbjct: 78 RNKFALACAILASMTSILLGYDIGVMSGAVIYIKRDLKISDVQIEILSGIMNIY---SPI 248
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ + G +D GRR I++A +F +G+++M +PN A L+FGR G G+G A + +P
Sbjct: 249 GSYIAGRTSDWIGRRYTIVLAGLIFFVGAILMGFSPNYAFLMFGRFVAGVGIGFAFLIAP 428
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT--WRWMLGVAAAPAI 197
+Y SE SPT RG L SL + G + Y+ N F+ P WR MLG+ A P+I
Sbjct: 429 VYTSEISPTLSRGFLTSLPEVFLNAGILIGYISNYTFSKLPLRLGWRLMLGIGAIPSI 602
>TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporter, partial
(37%)
Length = 903
Score = 120 bits (302), Expect = 5e-28
Identities = 88/279 (31%), Positives = 134/279 (47%), Gaps = 17/279 (6%)
Frame = +2
Query: 23 NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIG 82
N Y +A + L+GY V++G LL+I++D + D + ++ + A +
Sbjct: 77 NMYACASVVAASVIFALYGYVMAVMTGVLLFIKEDLQLSDLQVQFSVGLLHMSALPACMT 256
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
A G I D GRR II+ +FL+GSV+M P+ IL+ G G G+G + + +P+
Sbjct: 257 A---GRIADYIGRRYTIILTAIIFLVGSVLMGYGPSYPILMIGLSISGIGLGFSLIVAPV 427
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAF--TNAPGTWRWMLGVAAAPAIIQI 200
Y +E SP RG L S G L Y+ N F + WR ML V A P++ +
Sbjct: 428 YCAEISPPSHRGFLTSFLEVSTNAGIALGYVSNYFFGKLSIRLGWRLMLVVHAVPSLALV 607
Query: 201 VLMLSLPESPRWLYRKGREEESKSILKKI-YAPEDVDAEIEALKESVESEIEESKTSKIS 259
LML+ ESPRWL +GR E++ +L + E+ + ++ +K +V I+E T I
Sbjct: 608 ALMLNWVESPRWLVMQGRVGEARKVLLLVSNTKEEAEQRLKEIKVAV--GIDEKCTQDIV 781
Query: 260 M--------------MTLLKTTTVRRGLYAGMGLQFFQQ 284
M T VRR L A +G+ FQQ
Sbjct: 782 QVPNRTRNGGGAFREMFCKPTPPVRRILIAAVGVHAFQQ 898
>TC19595
Length = 500
Score = 102 bits (255), Expect = 1e-22
Identities = 43/87 (49%), Positives = 67/87 (76%)
Frame = +1
Query: 469 VPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVA 528
VPW +NSEIYP +RGVCGG+++T W+ ++I+SQSFLS++ +G +F + G++AVVA
Sbjct: 4 VPWTVNSEIYPEEFRGVCGGMSATVNWVCSVIMSQSFLSISDAVGLGGSFAILGVIAVVA 183
Query: 529 IFFVIVFVPETKGVPMEEVEKMLEQRS 555
FV++FVPETKG+ EE+ + ++R+
Sbjct: 184 FLFVLLFVPETKGLTFEEMTLIWKRRA 264
>CB826772
Length = 490
Score = 91.3 bits (225), Expect = 4e-19
Identities = 54/151 (35%), Positives = 78/151 (50%)
Frame = +1
Query: 23 NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIG 82
N Y +A I ++GY+ GV++GALL+I++D + D + I A +
Sbjct: 46 NLYACVSVMAASILSAIYGYEVGVMTGALLFIKEDLQISDLQLQFLVGIFHMCGLPASMA 225
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
A G I+D GRR II+A FL GS++ +P+ IL+ G G G G + +PL
Sbjct: 226 A---GRISDYIGRRYTIILASIAFLSGSILTGYSPSYLILMIGNFISGIGAGFTLIVAPL 396
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYL 173
Y +E SP RG+L SL F I G L Y+
Sbjct: 397 YCAEISPPSHRGSLTSLTEFSINIGILLGYM 489
>TC19365 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (24%)
Length = 441
Score = 82.0 bits (201), Expect = 2e-16
Identities = 46/121 (38%), Positives = 71/121 (58%)
Frame = +2
Query: 39 LFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVA 98
+FGY TGV++GALL+I+++ D + L I++ A+ V G +D GRR
Sbjct: 83 IFGYVTGVMAGALLFIKEELGINDMQVQLLGGILNVC---ALPACMVAGRTSDYVGRRYT 253
Query: 99 IIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVS 158
II++ + L+GS++M P+ IL+ GR G GVG+A + +P+Y +E S RG L S
Sbjct: 254 IILSAVILLLGSILMGYGPSFPILMIGRCTAGLGVGLALIIAPVYSTEISLPSNRGFLTS 433
Query: 159 L 159
L
Sbjct: 434 L 436
>CB828147
Length = 570
Score = 81.6 bits (200), Expect = 3e-16
Identities = 61/189 (32%), Positives = 89/189 (46%), Gaps = 17/189 (8%)
Frame = +1
Query: 162 FLITGGQFLSYLINLAFTNAPGT--WRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGRE 219
F I G L Y+ F WR ML V AAP+I I LML L ESPRWL +GR
Sbjct: 4 FSINIGILLGYMSTFFFEKLRLNLGWRMMLAVPAAPSIALIFLMLKLGESPRWLIMQGRV 183
Query: 220 EESKSILKKIY-APEDVDAEIEALKESVESEIEESKTSKISMMT--------LLK----- 265
++K +L ++ ++ + + +K +V IE + T +I + LK
Sbjct: 184 GDAKKVLLLVFNTKQEAEQRLAEIKTAV--GIEPNNTQEIVQVPKKTRHGGGALKELFCK 357
Query: 266 -TTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFG 324
T VRR + A +GL F GI ++ YSP I + G S LL ++ +
Sbjct: 358 PTPLVRRIMVAAVGLHVFMHLGGIGVILLYSPRIFEKTGITSKSKLLLCTIGMGVMKLVF 537
Query: 325 SILSIYFID 333
S +SI+F+D
Sbjct: 538 SFISIFFMD 564
>AV418433
Length = 401
Score = 77.8 bits (190), Expect = 4e-15
Identities = 41/133 (30%), Positives = 74/133 (54%)
Frame = +3
Query: 109 GSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQ 168
G +++ + P +L GRV G+G+G+ S P++I+E +P +RGAL +LN F++
Sbjct: 9 GWLVIYFSQGPVLLDIGRVATGYGMGVFSYVVPVFIAEIAPKELRGALTTLNQFMLVSAM 188
Query: 169 FLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKK 228
+S++I +WR + P + ++ + +PESPRWL ++G E+ ++ L +
Sbjct: 189 SVSFVIGNVL-----SWRALALTGLIPTAVLLLGLFFIPESPRWLAKRGCREDFEAAL-Q 350
Query: 229 IYAPEDVDAEIEA 241
I +D D EA
Sbjct: 351 ILRGKDADISQEA 389
>TC16378 similar to GB|AAM70554.1|21700861|AY124845 At1g75220/F22H5_6
{Arabidopsis thaliana;}, partial (31%)
Length = 738
Score = 77.4 bits (189), Expect = 6e-15
Identities = 40/119 (33%), Positives = 69/119 (57%), Gaps = 7/119 (5%)
Frame = +2
Query: 437 AWYTRGCPSK-------FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGI 489
A+Y G S+ G ++VGL + +I FS G+G +PWVI SEI P+ +G+ G
Sbjct: 92 AFYLEGTVSEDSHLYKILGIVSVVGLVVMVIGFSLGLGPIPWVIMSEILPVSIKGLAGST 271
Query: 490 ASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
A+ W+++ ++ + +L T + TF M+ ++A + F ++VPETKG +EE++
Sbjct: 272 ATMANWLTSWAITMT-ANLLLTWSSGGTFTMYTVVAAFTVVFTALWVPETKGRTLEEIQ 445
Score = 28.9 bits (63), Expect = 2.4
Identities = 12/32 (37%), Positives = 23/32 (71%)
Frame = +2
Query: 328 SIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
S + +DK+GR+ L L+S + +SL ++++AF
Sbjct: 2 STWLVDKSGRRLLLLLSSSIMTVSLVVVSIAF 97
>BG662351
Length = 407
Score = 73.9 bits (180), Expect = 6e-14
Identities = 33/98 (33%), Positives = 60/98 (60%)
Frame = +1
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQ 510
A+ G+ +Y+ +F+ GMG VPWV+ SEI+P+ +G G +A+ W + S +F +
Sbjct: 88 AVTGILVYVGSFAIGMGAVPWVVMSEIFPVNIKGQAGSLATLVNWFGAWLCSYTF-NFLM 264
Query: 511 TIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
+ T TF+++ + + I ++V VPETKG +E+++
Sbjct: 265 SWSTYGTFILYAAINALGILSIVVVVPETKGKSLEQLQ 378
>TC9536 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, partial (17%)
Length = 537
Score = 68.6 bits (166), Expect = 3e-12
Identities = 32/95 (33%), Positives = 56/95 (58%)
Frame = +2
Query: 453 VGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTI 512
V + +Y F G +P ++ SEI+P R RG+C I + WI ++IV+ + + ++
Sbjct: 32 VCVVVYFCFFVMAYGPIPNILCSEIFPTRVRGLCIAICALVFWIGDIIVTYTLPVMLSSM 211
Query: 513 GTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEV 547
G A F ++ I+ ++ FV + VPETKG+P+E +
Sbjct: 212 GLAGVFGLYAIVCCISWVFVYLKVPETKGMPLEVI 316
>TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(21%)
Length = 550
Score = 67.8 bits (164), Expect = 5e-12
Identities = 34/102 (33%), Positives = 56/102 (54%)
Frame = +2
Query: 458 YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWT 517
Y+ TFS G G + WV +SEI+PLR R + +++ ++S +FLSL++ I
Sbjct: 14 YVATFSIGAGPITWVYSSEIFPLRLRAQGCAMGVVVNRVTSGVISMTFLSLSKGITIGGA 193
Query: 518 FMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFK 559
F +FG +A+ F +PET+G +E++E Q + K
Sbjct: 194 FFLFGGIAICGWIFFYTMLPETRGKTLEDMEGSFGQSKSKIK 319
>TC17041 homologue to GB|AAL25568.1|16648753|AY058152 AT5g16150/T21H19_70
{Arabidopsis thaliana;} , partial (18%)
Length = 632
Score = 64.7 bits (156), Expect = 4e-11
Identities = 33/99 (33%), Positives = 54/99 (54%)
Frame = +2
Query: 457 LYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAW 516
LY+++FS G G VP ++ EI+ R R ++ WISN ++ FLS+ G +
Sbjct: 8 LYVLSFSLGAGPVPALLLPEIFASRIRAKAVSLSLGMHWISNFVIGLYFLSVVNKFGISS 187
Query: 517 TFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
++ F + V+A+ ++ V ETKG +EE+E L S
Sbjct: 188 VYLGFSAVCVLAVLYIAGNVVETKGRSLEEIEHALSSSS 304
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,139,627
Number of Sequences: 28460
Number of extensions: 147190
Number of successful extensions: 858
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 812
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 824
length of query: 571
length of database: 4,897,600
effective HSP length: 95
effective length of query: 476
effective length of database: 2,193,900
effective search space: 1044296400
effective search space used: 1044296400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0140.7


