

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
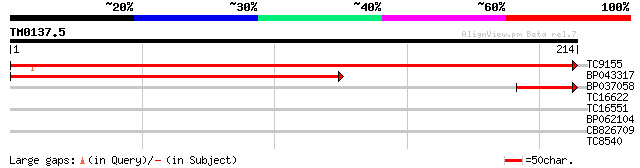
Query= TM0137.5
(214 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9155 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, complete 436 e-123
BP043317 239 2e-64
BP037058 47 3e-06
TC16622 similar to UP|Q8L759 (Q8L759) Expressed protein (At5g576... 28 1.2
TC16551 similar to UP|Q9SEZ7 (Q9SEZ7) CBL-interacting protein ki... 26 4.6
BP062104 26 6.0
CB826709 26 6.0
TC8540 similar to GB|AAM16209.1|20334906|AY094053 AT5g02020/T7H2... 25 7.8
>TC9155 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, complete
Length = 1087
Score = 436 bits (1120), Expect = e-123
Identities = 213/215 (99%), Positives = 213/215 (99%), Gaps = 1/215 (0%)
Frame = +1
Query: 1 MSLFGLG-RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 59
MSLFGLG RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV
Sbjct: 214 MSLFGLGSRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 393
Query: 60 NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 119
NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM
Sbjct: 394 NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 573
Query: 120 DWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 179
DWIESQLDDETIFPQKLGAPFPP FRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL
Sbjct: 574 DWIESQLDDETIFPQKLGAPFPPXFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 753
Query: 180 NTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL 214
NTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL
Sbjct: 754 NTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL 858
>BP043317
Length = 448
Score = 239 bits (611), Expect = 2e-64
Identities = 111/126 (88%), Positives = 121/126 (95%)
Frame = -3
Query: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVN 60
MSLFGLGRNQ+TFRPKKS PSGSKGAQL+KHIDATLGSGNLREAVKLPPGED+NEWLAVN
Sbjct: 380 MSLFGLGRNQRTFRPKKSTPSGSKGAQLRKHIDATLGSGNLREAVKLPPGEDLNEWLAVN 201
Query: 61 TVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMD 120
TVDFFNQVN+L+GTLTEFCT NC +M+AGPKYEYRWADGV IK+PIEV+APKYVEYLMD
Sbjct: 200 TVDFFNQVNLLYGTLTEFCTAENCRTMSAGPKYEYRWADGVQIKRPIEVTAPKYVEYLMD 21
Query: 121 WIESQL 126
WIE+QL
Sbjct: 20 WIETQL 3
>BP037058
Length = 544
Score = 46.6 bits (109), Expect = 3e-06
Identities = 23/23 (100%), Positives = 23/23 (100%)
Frame = -2
Query: 192 EFRLIDKAELAPLEDLVDSIIQL 214
EFRLIDKAELAPLEDLVDSIIQL
Sbjct: 258 EFRLIDKAELAPLEDLVDSIIQL 190
>TC16622 similar to UP|Q8L759 (Q8L759) Expressed protein (At5g57655),
partial (37%)
Length = 649
Score = 28.1 bits (61), Expect = 1.2
Identities = 11/18 (61%), Positives = 16/18 (88%)
Frame = +1
Query: 34 ATLGSGNLREAVKLPPGE 51
A++G GNLRE++K+ PGE
Sbjct: 73 ASMG*GNLRESMKMKPGE 126
>TC16551 similar to UP|Q9SEZ7 (Q9SEZ7) CBL-interacting protein kinase 16,
partial (5%)
Length = 543
Score = 26.2 bits (56), Expect = 4.6
Identities = 22/70 (31%), Positives = 34/70 (48%)
Frame = -1
Query: 40 NLREAVKLPPGEDINEWLAVNTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWAD 99
N RE + +P E N L+ +T +FN +I + F PS S GP++ Y W
Sbjct: 261 NQRENLLVP*VEAPNSTLSRST*LYFNITHI-YHCHFNFTKPSI--ST*LGPRHHYTWHA 91
Query: 100 GVTIKKPIEV 109
+T+ +P V
Sbjct: 90 LLTLPRPNNV 61
>BP062104
Length = 321
Score = 25.8 bits (55), Expect = 6.0
Identities = 16/59 (27%), Positives = 25/59 (42%)
Frame = -2
Query: 136 LGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFR 194
L PF + +V IF +L R +YH+ F + F HF++ T+ R
Sbjct: 170 LETPFSLDSSSLVGAIFNKLSRSSVPMYHTAFSE------------SFTHFIIVTFSAR 30
>CB826709
Length = 432
Score = 25.8 bits (55), Expect = 6.0
Identities = 9/34 (26%), Positives = 21/34 (61%)
Frame = +3
Query: 98 ADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETI 131
++G I + +++ Y+E + D +E Q+DD+ +
Sbjct: 246 SEGAEIDERVDLDEENYMEEMDDDVEEQVDDDGV 347
>TC8540 similar to GB|AAM16209.1|20334906|AY094053 AT5g02020/T7H20_70
{Arabidopsis thaliana;}, partial (17%)
Length = 607
Score = 25.4 bits (54), Expect = 7.8
Identities = 9/26 (34%), Positives = 18/26 (68%)
Frame = +2
Query: 185 HFVLFTWEFRLIDKAELAPLEDLVDS 210
HFV + + F+L+D+ ++ ++LV S
Sbjct: 200 HFVSYIYRFKLVDRRKMEGRKNLVSS 277
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.139 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,840,768
Number of Sequences: 28460
Number of extensions: 52155
Number of successful extensions: 284
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 284
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 284
length of query: 214
length of database: 4,897,600
effective HSP length: 86
effective length of query: 128
effective length of database: 2,450,040
effective search space: 313605120
effective search space used: 313605120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0137.5


