

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.7
(841 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
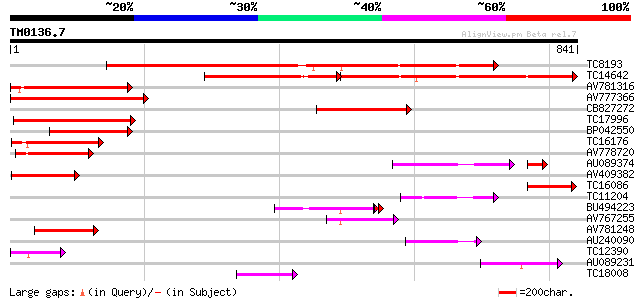
Sequences producing significant alignments: (bits) Value
TC8193 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lactas... 660 0.0
TC14642 homologue to UP|O65761 (O65761) Beta-galactosidase (Lac... 311 e-145
AV781316 327 4e-90
AV777366 288 3e-78
CB827272 284 4e-77
TC17996 similar to UP|O23243 (O23243) Beta-galactosidase like pr... 273 8e-74
BP042550 201 4e-52
TC16176 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lacta... 197 4e-51
AV778720 165 3e-41
AU089374 126 1e-33
AV409382 135 4e-32
TC16086 similar to UP|Q93X57 (Q93X57) Beta-galactosidase (Lacta... 124 8e-29
TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Ar... 107 8e-24
BU494223 100 7e-22
AV767255 97 1e-20
AV781248 95 5e-20
AU240090 82 4e-16
TC12390 82 5e-16
AU089231 81 6e-16
TC18008 weakly similar to UP|O49609 (O49609) Beta-galactosidase ... 74 1e-13
>TC8193 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lactase) ,
partial (80%)
Length = 2257
Score = 660 bits (1704), Expect = 0.0
Identities = 321/593 (54%), Positives = 398/593 (66%), Gaps = 12/593 (2%)
Frame = +2
Query: 144 DNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWA 203
DN PFKA MQ+FT KIV+MMK E L+ +QGGPII+SQIENEYG VE G Y WA
Sbjct: 5 DNGPFKAAMQKFTTKIVNMMKAEKLFQTQGGPIIMSQIENEYGPVEWETGAPGKAYTKWA 184
Query: 204 ASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFG 263
A MA L+TGVPW+MC+QE+APDP+I+TCN +YC++FTPN KPK WTE + GW +G
Sbjct: 185 AQMAVGLNTGVPWIMCKQEDAPDPLIDTCNAYYCEKFTPNKNYKPKMWTENWTGWYTDYG 364
Query: 264 GAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYG 323
VP RP ED+AF+VARF Q GG+ NYYMYHGGTNFGRTS G F+ATSY+YDA IDEYG
Sbjct: 365 SPVPRRPAEDMAFSVARFIQNGGSFVNYYMYHGGTNFGRTSSGLFIATSYEYDAPIDEYG 544
Query: 324 FIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTE-AECVAFLANIDNT 382
+ +PKWGHL+D+HKAIK CE AL++ DPT+T LG LEA V+KT C AFLAN D T
Sbjct: 545 LLNEPKWGHLRDLHKAIKQCEPALVSVDPTLTLLGKGLEAHVFKTSFGACAAFLANYDTT 724
Query: 383 SDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSP 442
S TVKF YNLP WS+SILPDCK V NTA++ +S T +
Sbjct: 725 SSRTVKFGNGQYNLPPWSISILPDCKTDVYNTARVGVKSWEKKMTP-----------VNS 871
Query: 443 GWSWIS---EPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDA-----GAQT 494
++W+S +P S DS + + L EQ+N T D SDYLWY ++++ + G
Sbjct: 872 AFNWLSYNEQPASSSGDDSLTGYALWEQVNVTRDSSDYLWYMTDVNIDSNEGLIKNGQSP 1051
Query: 495 VLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFG 554
VL S GH LH F+NG+L+G+ G N K+ + L G N I LLS++VGL + G
Sbjct: 1052VLTAMSAGHVLHVFVNGQLSGTVYGGLKNPKLTFSDSVKLRVGNNKISLLSVSVGLSNVG 1231
Query: 555 PFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGL--PSGTSG-QWNSQS 611
F+TW G+ GPV LKGL G T D S ++W+Y++GLKGE L L SG+S QW S
Sbjct: 1232SHFETWNVGVLGPVTLKGLNEG-TRDLSRQKWSYKVGLKGEALNLHTTSGSSSVQWTQGS 1408
Query: 612 TLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSS 671
+ K QPLTWYK F+AP+ NDP+A+D T MGKGE W+NGQSIGR+WP YI+ G S
Sbjct: 1409LVAKKQPLTWYKATFNAPASNDPLALDMTSMGKGEVWINGQSIGRHWPGYIA--RGSCGS 1582
Query: 672 CNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISI 724
CNY G Y +KC NCG+PSQ YH+PRSWL P N +V+ EE GGDPT IS+
Sbjct: 1583CNYAGYYTDTKCRTNCGEPSQRWYHIPRSWLNPSGNLVVVLEEFGGDPTGISL 1741
>TC14642 homologue to UP|O65761 (O65761) Beta-galactosidase (Lactase)
(Fragment) , partial (78%)
Length = 2239
Score = 311 bits (798), Expect(2) = e-145
Identities = 158/356 (44%), Positives = 215/356 (60%), Gaps = 5/356 (1%)
Frame = +1
Query: 491 GAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGL 550
G L + S GHALH F+NG+L+G+ G+ + K+ + L AG N I LLS+ VGL
Sbjct: 610 GKNPFLTVLSAGHALHVFVNGQLSGTAYGSLEFPKLTFSEGVKLRAGVNKISLLSVAVGL 789
Query: 551 QHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPS---GTSGQW 607
+ GP F+TW AG+ GP+ L GL G D + ++W+Y+IGLKGE L L S +S +W
Sbjct: 790 PNVGPHFETWNAGVLGPITLNGLNEGRR-DLTWQKWSYKIGLKGESLSLHSLSGSSSVEW 966
Query: 608 NSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDG 667
+ + QPLTWYK F AP+G P+A+D MGKG+ W+NGQS+GRYWP Y + G
Sbjct: 967 IQGYLVSRRQPLTWYKTTFDAPAGVAPLALDMESMGKGQVWINGQSLGRYWPAYKA--SG 1140
Query: 668 CTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIK 727
CNY G Y KC NCG+ SQ YHVP SWL+P N LV+FEE GGDP IS+ +
Sbjct: 1141SCDYCNYAGTYHEKKCGSNCGEASQRWYHVPHSWLKPTGNLLVVFEELGGDPNGISLVRR 1320
Query: 728 LIGSSCSHVSESHPPPVDL-WNSDTESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGN 786
I S+C+ + E P V + + P L C P + I++IKFASFGTP G+
Sbjct: 1321DIDSACADIYEWQPNLVSYQMQASGKVSAPVRPKAHLSCG-PGQKISSIKFASFGTPVGS 1497
Query: 787 CGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
CGN+ G C + ++ +++C+G + C++ V+ F GDPC V K L+VEA CT
Sbjct: 1498CGNYHEGSCHAHKSYDAFERSCVGQNQCAVTVAPEIFGGDPCPNVMKKLSVEAVCT 1665
Score = 223 bits (567), Expect(2) = e-145
Identities = 111/203 (54%), Positives = 140/203 (68%), Gaps = 1/203 (0%)
Frame = +3
Query: 290 NYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIA 349
NYYMYHGGTNFGRT+GGPF+ATSYDYDA +DEYG ++QPKWGHLKD+H+AIKLCE AL++
Sbjct: 9 NYYMYHGGTNFGRTAGGPFIATSYDYDAPLDEYGLLQQPKWGHLKDLHRAIKLCEPALVS 188
Query: 350 TDPTITSLGSNLEAAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCK 408
DPT+T LG+ EA V+K+++ C AFLAN + S ATV F YNLP WS+SILP+CK
Sbjct: 189 GDPTVTRLGNYGEAHVFKSKSGACAAFLANYNPKSYATVVFGNQHYNLPPWSISILPNCK 368
Query: 409 NVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQI 468
+ V NTA++ S+S T + G L W +E + SF+ GLLEQI
Sbjct: 369 HTVYNTARVGSQSTQMKMTRVPIH--GGL-----SWRSFNEETTTTDDSSFTVTGLLEQI 527
Query: 469 NTTADRSDYLWYSLRIDLEDDAG 491
N T D SDYLWYS + + + G
Sbjct: 528 NATRDLSDYLWYSTDVVINPNEG 596
>AV781316
Length = 574
Score = 327 bits (839), Expect = 4e-90
Identities = 157/183 (85%), Positives = 167/183 (90%), Gaps = 3/183 (1%)
Frame = +1
Query: 2 AMRRTQFLLLL---LWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEM 58
AMR TQ +++L LW LC P FC NVTYDHRALVIDGKRRVL+SGSIHYPRSTPEM
Sbjct: 34 AMRATQIIMVLFRFLWCLC---PCLFCTNVTYDHRALVIDGKRRVLVSGSIHYPRSTPEM 204
Query: 59 WPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGP 118
WPDLIQK+KDGGLDVIETYVFWNLHEPV+GQYNFEGRGDLV+FVK VA AGLYVHLRIGP
Sbjct: 205 WPDLIQKSKDGGLDVIETYVFWNLHEPVRGQYNFEGRGDLVQFVKAVAAAGLYVHLRIGP 384
Query: 119 YACAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIIL 178
YACAEWNYGGFPL LHFIPGIQFRT+NEPFKAEM+RFTAKIVDMMKQENLYA +GGPIIL
Sbjct: 385 YACAEWNYGGFPLGLHFIPGIQFRTNNEPFKAEMKRFTAKIVDMMKQENLYAREGGPIIL 564
Query: 179 SQI 181
SQI
Sbjct: 565 SQI 573
>AV777366
Length = 637
Score = 288 bits (736), Expect = 3e-78
Identities = 136/206 (66%), Positives = 162/206 (78%)
Frame = -3
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
M + + +L+L++L A+V+YDH+A+ I+G+RR+L+SGSIHYPRSTPEMWP
Sbjct: 620 MGFKLKTWNVLVLFVLACSLLGHAAASVSYDHKAITINGQRRILLSGSIHYPRSTPEMWP 441
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYA 120
DLIQKAK+GGLDVI+TYVFWN HEP QG+Y FEG DLVKF+K V AGLYVHLRIGPY
Sbjct: 440 DLIQKAKEGGLDVIQTYVFWNGHEPSQGKYYFEGNYDLVKFIKLVQQAGLYVHLRIGPYV 261
Query: 121 CAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQ 180
CAEWN+GGFP+WL ++PGI FRTDN PFK +MQRFT KIV+MMK E LY +QGGPIILSQ
Sbjct: 260 CAEWNFGGFPVWLKYVPGISFRTDNGPFKYQMQRFTNKIVNMMKAERLYETQGGPIILSQ 81
Query: 181 IENEYGNVEGAYGPAAVPYINWAASM 206
IENEYG +E G Y WAASM
Sbjct: 80 IENEYGPMEYEIGSPGKAYSQWAASM 3
>CB827272
Length = 470
Score = 284 bits (727), Expect = 4e-77
Identities = 140/141 (99%), Positives = 140/141 (99%)
Frame = +1
Query: 455 KADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLA 514
KADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLA
Sbjct: 46 KADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLA 225
Query: 515 GSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLK 574
GSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGA ITGPVILKGLK
Sbjct: 226 GSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAXITGPVILKGLK 405
Query: 575 NGSTLDFSSKQWTYQIGLKGE 595
NGSTLDFSSKQWTYQIGLKGE
Sbjct: 406 NGSTLDFSSKQWTYQIGLKGE 468
>TC17996 similar to UP|O23243 (O23243) Beta-galactosidase like protein
(Lactase) , partial (20%)
Length = 677
Score = 273 bits (698), Expect = 8e-74
Identities = 129/181 (71%), Positives = 147/181 (80%)
Frame = +3
Query: 6 TQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQK 65
++FLL L + + VTYD +AL+I+G+RR+LISGSIHYPRSTP+MW DLIQK
Sbjct: 108 SKFLLPFLCFALFSSILVVHSAVTYDSKALLINGQRRILISGSIHYPRSTPDMWEDLIQK 287
Query: 66 AKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWN 125
AKDGGLDVIETYVFWN+HEP QG YNFEGR DLV+FVK + AGLY HLRIGPY CAEWN
Sbjct: 288 AKDGGLDVIETYVFWNVHEPSQGNYNFEGRYDLVRFVKTIQKAGLYAHLRIGPYVCAEWN 467
Query: 126 YGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEY 185
+GGFP+WL ++PGI FRTDNEPFK MQ FT KIV MMK E+LY S GGPIILSQIENEY
Sbjct: 468 FGGFPVWLKYVPGISFRTDNEPFKRAMQGFTEKIVGMMKSEHLYESXGGPIILSQIENEY 647
Query: 186 G 186
G
Sbjct: 648 G 650
>BP042550
Length = 489
Score = 201 bits (511), Expect = 4e-52
Identities = 90/123 (73%), Positives = 102/123 (82%)
Frame = +3
Query: 59 WPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGP 118
W DLIQKAK GGLDVI+TYVFWN+HEP G YNFEGR DLV+F+K V GLYVHLRIGP
Sbjct: 3 WEDLIQKAKHGGLDVIDTYVFWNVHEPSPGNYNFEGRYDLVRFIKTVQKVGLYVHLRIGP 182
Query: 119 YACAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIIL 178
Y CAEWN+GGFP+WL ++PGI FRTDN PFKA M+ FT KIV MMK E L+ SQGGPIIL
Sbjct: 183 YVCAEWNFGGFPVWLKYVPGISFRTDNGPFKAAMRGFTQKIVQMMKNEKLFQSQGGPIIL 362
Query: 179 SQI 181
+Q+
Sbjct: 363 AQV 371
>TC16176 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lactase) ,
partial (18%)
Length = 649
Score = 197 bits (502), Expect = 4e-51
Identities = 93/139 (66%), Positives = 112/139 (79%), Gaps = 3/139 (2%)
Frame = +1
Query: 3 MRRTQFLLLLLWLLCVYAPACFC---ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMW 59
MRR++F +L+ C + FC A+VTYDH+A+++DGKRR+LISGSIHYPRSTP+MW
Sbjct: 244 MRRSKFHVLVFMWFCSW----FCYVTASVTYDHKAILVDGKRRILISGSIHYPRSTPQMW 411
Query: 60 PDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPY 119
PDLIQKAKDGG+DVI+TYVF N HEP G Y FE R DLVKF+K V AGLYV+LRIGPY
Sbjct: 412 PDLIQKAKDGGIDVIQTYVFGNGHEPSPGNYYFEDRLDLVKFIKIVQQAGLYVNLRIGPY 591
Query: 120 ACAEWNYGGFPLWLHFIPG 138
CAEWN GGFP+WL ++PG
Sbjct: 592 VCAEWNLGGFPVWLKYVPG 648
>AV778720
Length = 512
Score = 165 bits (417), Expect = 3e-41
Identities = 79/116 (68%), Positives = 93/116 (80%)
Frame = +1
Query: 9 LLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKD 68
L+LLL +L C NV+YD +AL+I+G+RRVLISGSIHYPRSTPEMW DLIQKAK
Sbjct: 166 LVLLLTVLFAGFELIHC-NVSYDRKALLINGQRRVLISGSIHYPRSTPEMWEDLIQKAKH 342
Query: 69 GGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEW 124
GG+DVI+TYVFW++HEP G YNFEGR DLV+F+K V GLY +LRIGPY CAEW
Sbjct: 343 GGIDVIDTYVFWDVHEPSPGNYNFEGRYDLVRFIKTVQKLGLYANLRIGPYVCAEW 510
>AU089374
Length = 694
Score = 126 bits (317), Expect(2) = 1e-33
Identities = 67/182 (36%), Positives = 98/182 (53%), Gaps = 1/182 (0%)
Frame = +3
Query: 568 VILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGTSGQWNSQSTLPKNQP-LTWYKINF 626
+ + GL +G +D S W +Q+GLKGE + + + + P L+WYK NF
Sbjct: 24 IFIIGLNSGK-IDLSLNGWGHQVGLKGEXNKIFTEKGSKKVEWKDVKGPGPVLSWYKTNF 200
Query: 627 SAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKN 686
+ P G DP+AI GMGKG W+NG+SIGR+W +Y+SP+
Sbjct: 201 ATPEGRDPVAIRMEGMGKGMVWINGKSIGRHWMSYLSPL--------------------- 317
Query: 687 CGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDL 746
GKP+Q+ YH+PRS+L N LV+FEE P +I+I + C ++E+HPP V
Sbjct: 318 -GKPTQSEYHIPRSYLNSKDNLLVVFEEXVXXPXKIAILNVNRDTICXFITENHPPNVKS 494
Query: 747 WN 748
W+
Sbjct: 495 WS 500
Score = 34.3 bits (77), Expect(2) = 1e-33
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 1/31 (3%)
Frame = +2
Query: 768 PN-EVITTIKFASFGTPHGNCGNFSHGDCSS 797
PN + I ++FA FG P G CG F+ G C++
Sbjct: 566 PNXKTIKAVEFAXFGDPEGYCGKFTMGKCNA 658
>AV409382
Length = 430
Score = 135 bits (339), Expect = 4e-32
Identities = 63/103 (61%), Positives = 79/103 (76%), Gaps = 2/103 (1%)
Frame = +2
Query: 3 MRRTQF--LLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
M T F L++ L + + VTYD +A++I+G+RR+L+SGSIHYPRSTP+MW
Sbjct: 122 METTSFSKLVVAFCLALCFGSQLIHSTVTYDRKAILINGQRRILLSGSIHYPRSTPDMWE 301
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVK 103
DLIQKAK+GGLDVIETYVFWN+HEP G YNFEG+ DLVKF+K
Sbjct: 302 DLIQKAKEGGLDVIETYVFWNVHEPSPGNYNFEGKYDLVKFIK 430
>TC16086 similar to UP|Q93X57 (Q93X57) Beta-galactosidase (Lactase) ,
partial (9%)
Length = 551
Score = 124 bits (310), Expect = 8e-29
Identities = 57/72 (79%), Positives = 66/72 (91%)
Frame = +3
Query: 769 NEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCG 828
N+VI++IKFAS+GTP G CGNF HG CSS +ALS+VQKACIGSSSCS+GVSI+TFGDPC
Sbjct: 3 NQVISSIKFASYGTPAGTCGNFYHGRCSSHKALSLVQKACIGSSSCSVGVSIDTFGDPCT 182
Query: 829 GVTKSLAVEASC 840
GVTKSLAVEA+C
Sbjct: 183 GVTKSLAVEATC 218
>TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Arabidopsis
thaliana;}, partial (13%)
Length = 568
Score = 107 bits (267), Expect = 8e-24
Identities = 59/148 (39%), Positives = 79/148 (52%), Gaps = 3/148 (2%)
Frame = +2
Query: 580 DFSSKQWTYQIGLKGEDLGL--PSGTSG-QWNSQSTLPKNQPLTWYKINFSAPSGNDPIA 636
D ++ W YQ+GL GE L + SG+S QW S + PLTWY+ F AP GNDP+
Sbjct: 56 DVTNPSWGYQVGLLGEKLQIYTASGSSKVQWESFQS--ST*PLTWYQTTFDAPEGNDPVV 229
Query: 637 IDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYH 696
++ MGKG WVNGQ IGRYW ++ +P G SQ YH
Sbjct: 230 LNLGSMGKGITWVNGQGIGRYWVSFHTP----------------------DGTSSQNWYH 343
Query: 697 VPRSWLQPDSNTLVLFEESGGDPTQISI 724
+PRS L+ N LV+ EE G+P +I++
Sbjct: 344 IPRSILKSTGNLLVILEEESGNPLEITL 427
>BU494223
Length = 495
Score = 99.8 bits (247), Expect(2) = 7e-22
Identities = 54/159 (33%), Positives = 82/159 (50%), Gaps = 6/159 (3%)
Frame = +2
Query: 394 YNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSWISEPIG- 452
Y+LP WS+SILPDCK V NTAK+ + +LK W E +
Sbjct: 20 YDLPPWSISILPDCKADVFNTAKVRVRTSKVQMIPVNLKLFS--------WETYDEDLSS 175
Query: 453 ISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD-----AGAQTVLHIESLGHALHA 507
++++ + GLLEQ+N T D SDYLWY +D+ G Q ++++S GH +H
Sbjct: 176 LAESSRITAPGLLEQLNVTRDSSDYLWYITSVDISSSESFLGGGHQPSINVQSAGHGIHV 355
Query: 508 FINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSL 546
F+NG+ +GS G ++ P++L AG N + +L
Sbjct: 356 FVNGQYSGSAFGAKEQRSCKFNGPVSLHAGTNKLHFSAL 472
Score = 21.9 bits (45), Expect(2) = 7e-22
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +1
Query: 541 IDLLSLTVGLQHFG 554
I LLS+TVGLQ+ G
Sbjct: 454 IALLSVTVGLQNVG 495
>AV767255
Length = 342
Score = 96.7 bits (239), Expect = 1e-20
Identities = 49/111 (44%), Positives = 64/111 (57%), Gaps = 5/111 (4%)
Frame = +3
Query: 471 TADRSDYLWYSLRIDLEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSK 525
T D SDYLWY +D+ G L ++S GHA+H ING+L+GSG G R++ +
Sbjct: 3 TRDTSDYLWYITSVDIGSSESFLRGGELPTLIVQSTGHAVHIIINGQLSGSGYGTREDRR 182
Query: 526 VNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNG 576
P+ L AG NTI LLS+ VGL + G F+TW GI G + L GL G
Sbjct: 183 FRYTGPVNLRAGTNTIALLSVAVGLPNIGGHFETWNTGILGSIALHGLDKG 335
>AV781248
Length = 488
Score = 94.7 bits (234), Expect = 5e-20
Identities = 44/95 (46%), Positives = 59/95 (61%)
Frame = +2
Query: 38 DGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGD 97
DG+ ++ G +HY R PE W D + KAK GL+ I+ YV WNLHEP QG+ ++G +
Sbjct: 185 DGEPFRIMGGDLHYFRVHPEYWEDRLLKAKALGLNTIQVYVPWNLHEPTQGKLVWDGVAN 364
Query: 98 LVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLW 132
+ F+ GL V +R GPY CAEW+ GGFP W
Sbjct: 365 MESFLNLCHKLGLLVMVRPGPYICAEWDGGGFPAW 469
>AU240090
Length = 300
Score = 82.0 bits (201), Expect = 4e-16
Identities = 43/113 (38%), Positives = 60/113 (53%), Gaps = 1/113 (0%)
Frame = +3
Query: 588 YQIGLKGEDLGLPSGTSGQWNSQSTLP-KNQPLTWYKINFSAPSGNDPIAIDFTGMGKGE 646
+Q+GLKGE G+ + Q S L + ++WYK F+ P G P+AI TGMGKG
Sbjct: 3 HQVGLKGEANGIFTEEGSQKVKWSPLAGPGRAISWYKTRFTTPEGRGPVAIRMTGMGKGM 182
Query: 647 AWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPR 699
WVNG+SIGR+W +++SP+ G P+Q YH+PR
Sbjct: 183 IWVNGRSIGRHWVSFLSPL----------------------GLPTQAEYHIPR 275
>TC12390
Length = 403
Score = 81.6 bits (200), Expect = 5e-16
Identities = 40/101 (39%), Positives = 57/101 (55%), Gaps = 18/101 (17%)
Frame = +2
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCA------------------NVTYDHRALVIDGKRR 42
M R+T L + L + + A + + A NVTYD ++L ++G+R
Sbjct: 101 MTTRKTNILFITLLVTIIIASSIYPASAHGKKKRLAHNRIKNFQNVTYDGKSLFVNGRRE 280
Query: 43 VLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLH 83
+L SGS+HY RSTP MWP ++ A GGL+VI+TY FWN H
Sbjct: 281 LLFSGSVHYTRSTPXMWPVILDXATHGGLNVIQTYXFWNAH 403
>AU089231
Length = 383
Score = 81.3 bits (199), Expect = 6e-16
Identities = 44/125 (35%), Positives = 65/125 (51%), Gaps = 4/125 (3%)
Frame = +1
Query: 699 RSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSG- 757
R++L P N LV+ EE G P +I I + CS + ES PP V+ W S R
Sbjct: 10 RAYLNPKDNLLVILEEDQGTPEKIEIMNVNRDTVCSIIEESDPPNVNSWVSSHGQFRPXV 189
Query: 758 ---GPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSC 814
SL C N ++ ++FASFG P G+CG GDC++ IV++ C+G SC
Sbjct: 190 SNVATQASLSCGSGNXIVA-VEFASFGNPSGSCGKLVLGDCNAAATQXIVEQQCLGKGSC 366
Query: 815 SIGVS 819
++ ++
Sbjct: 367 NVDLN 381
>TC18008 weakly similar to UP|O49609 (O49609) Beta-galactosidase - like
protein (Beta-galactosidase-like protein), partial (8%)
Length = 301
Score = 73.6 bits (179), Expect = 1e-13
Identities = 38/92 (41%), Positives = 52/92 (56%), Gaps = 2/92 (2%)
Frame = +1
Query: 337 HKAIKLCEEALIATDPTITSLGSNLEAAVYKTEAE--CVAFLANIDNTSDATVKFNGNSY 394
HKA+ LC++AL+ P+ T L E VY+ E C AF+ N + ++ F G+ Y
Sbjct: 10 HKAVSLCKKALLTGKPSSTKLSRYHEVIVYEKEGTDLCAAFITNNHTKTAKSINFKGSKY 189
Query: 395 NLPAWSVSILPDCKNVVLNTAKINSESMISSF 426
LP S+SILPDCK VV NT I S+ +F
Sbjct: 190 YLPPRSISILPDCKTVVFNTQNIASQHNSRNF 285
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,309,579
Number of Sequences: 28460
Number of extensions: 276406
Number of successful extensions: 1355
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 1325
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1331
length of query: 841
length of database: 4,897,600
effective HSP length: 98
effective length of query: 743
effective length of database: 2,108,520
effective search space: 1566630360
effective search space used: 1566630360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0136.7


