

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
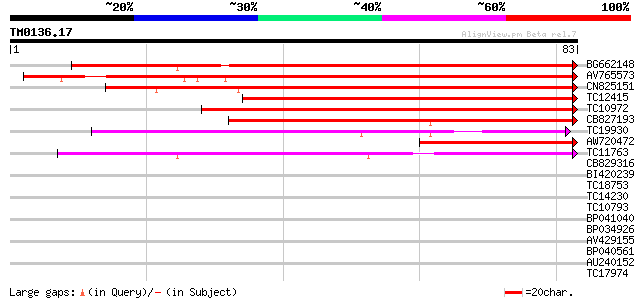
Query= TM0136.17
(83 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662148 108 2e-25
AV765573 106 7e-25
CN825151 69 2e-13
TC12415 similar to UP|Q9LII9 (Q9LII9) Genomic DNA, chromosome 3,... 59 1e-10
TC10972 similar to UP|Q40318 (Q40318) Coil protein, partial (32%) 59 2e-10
CB827193 57 6e-10
TC19930 weakly similar to UP|Q9AX26 (Q9AX26) GDSL-motif lipase/h... 39 2e-04
AW720472 39 2e-04
TC11763 37 6e-04
CB829316 36 0.001
BI420239 33 0.009
TC18753 32 0.016
TC14230 31 0.035
TC10793 30 0.060
BP041040 30 0.078
BP034926 30 0.10
AV429155 28 0.39
BP040561 27 0.51
AU240152 27 0.66
TC17974 UP|AAO45821 (AAO45821) Gamma-glutamylcysteine synthetase... 27 0.66
>BG662148
Length = 415
Score = 108 bits (269), Expect = 2e-25
Identities = 54/78 (69%), Positives = 62/78 (79%), Gaps = 4/78 (5%)
Frame = +2
Query: 10 VSFVSLFVILFTTT----LNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDT 65
++ +SLFV +FT T LNPV+ C+FPAIFN GASNSDTGG AAAF PK P+GDT
Sbjct: 26 ITQISLFVFVFTATSTTLLNPVYGGG-CDFPAIFNFGASNSDTGGYAAAFEGPKPPHGDT 202
Query: 66 YFHRPAGRYSDGRIILDF 83
YFHRPAGRYSDGRII+DF
Sbjct: 203 YFHRPAGRYSDGRIIIDF 256
>AV765573
Length = 440
Score = 106 bits (265), Expect = 7e-25
Identities = 62/87 (71%), Positives = 66/87 (75%), Gaps = 6/87 (6%)
Frame = +2
Query: 3 LMEF-HRHVSFVSLFVILFTTTL-NP--VFAK--KRCNFPAIFNLGASNSDTGGLAAAFL 56
LMEF RH+S LF +LFTT L NP VFA K C+FPAIFN GASNSDTGG AAAFL
Sbjct: 2 LMEFLSRHIS---LFALLFTTVLLNPDVVFATTTKTCDFPAIFNFGASNSDTGGFAAAFL 172
Query: 57 PPKSPYGDTYFHRPAGRYSDGRIILDF 83
P P GDTYF RPAGR+SDGRIILDF
Sbjct: 173QPGPPQGDTYFGRPAGRFSDGRIILDF 253
>CN825151
Length = 740
Score = 68.9 bits (167), Expect = 2e-13
Identities = 36/74 (48%), Positives = 48/74 (64%), Gaps = 5/74 (6%)
Frame = +1
Query: 15 LFVILF---TTTLNPVFAKKR--CNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHR 69
LF+ F T +N V K C FPAI+N G SNSDTGG++AAF P PYG+++ +
Sbjct: 22 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQK 201
Query: 70 PAGRYSDGRIILDF 83
P+ R DGR+I+DF
Sbjct: 202PSARDCDGRLIVDF 243
>TC12415 similar to UP|Q9LII9 (Q9LII9) Genomic DNA, chromosome 3, TAC
clone:K24A2, partial (22%)
Length = 366
Score = 59.3 bits (142), Expect = 1e-10
Identities = 27/49 (55%), Positives = 34/49 (69%)
Frame = +3
Query: 35 FPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+PAIFN G SNSDTG ++AAF P G + +GRYSDGR+I+DF
Sbjct: 78 YPAIFNFGDSNSDTGAISAAFTIVHPPNGQNFLGALSGRYSDGRLIIDF 224
>TC10972 similar to UP|Q40318 (Q40318) Coil protein, partial (32%)
Length = 548
Score = 58.9 bits (141), Expect = 2e-10
Identities = 26/55 (47%), Positives = 40/55 (72%)
Frame = +1
Query: 29 AKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+ +C++PA++N G SNS+TG + AAF +SP G ++F +GR SDGR+I+DF
Sbjct: 196 SSSKCSYPAVYNFGDSNSNTGVVYAAFAGLQSPGGISFFGNLSGRASDGRLIIDF 360
>CB827193
Length = 528
Score = 57.0 bits (136), Expect = 6e-10
Identities = 28/56 (50%), Positives = 36/56 (64%), Gaps = 5/56 (8%)
Frame = +1
Query: 33 CNFPAIFNLGASNSDTGGLAAAFLPPKS-----PYGDTYFHRPAGRYSDGRIILDF 83
C+F +IF+ G S +DTG L + P PYG TYFHRP+ R SDGRII+D+
Sbjct: 67 CSFSSIFSFGDSLADTGNLYLSSALPSHHCFFPPYGRTYFHRPSARCSDGRIIIDY 234
>TC19930 weakly similar to UP|Q9AX26 (Q9AX26) GDSL-motif
lipase/hydrolase-like protein, partial (11%)
Length = 329
Score = 38.9 bits (89), Expect = 2e-04
Identities = 26/79 (32%), Positives = 37/79 (45%), Gaps = 9/79 (11%)
Frame = +2
Query: 13 VSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG------LAAAFLPPKS---PYG 63
++L V+L + +F ++ P +F G S SD+G LA PP P G
Sbjct: 62 IALVVLLVAIAIMQIFVQRNTQVPCLFVFGDSLSDSGNNNNLETLAKVAYPPYGIDFPTG 241
Query: 64 DTYFHRPAGRYSDGRIILD 82
T P GRYS+GR +D
Sbjct: 242 PT----PTGRYSNGRTAVD 286
>AW720472
Length = 518
Score = 38.9 bits (89), Expect = 2e-04
Identities = 15/23 (65%), Positives = 18/23 (78%)
Frame = +2
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG+TYF P GR+ DGR+I DF
Sbjct: 221 PYGETYFKFPTGRFCDGRLISDF 289
>TC11763
Length = 469
Score = 37.0 bits (84), Expect = 6e-04
Identities = 23/85 (27%), Positives = 42/85 (49%), Gaps = 9/85 (10%)
Frame = +3
Query: 8 RHVSFVSLFVILFTTT-------LNPVFAKKRCNFPAIFNLGASNSDTGGL--AAAFLPP 58
+ ++ ++L ++ F T V++ K +F G S DTG + ++ PP
Sbjct: 81 KQITLLTLLLLFFIVTEVEGAKKSYGVYSNKNSIVKKLFVFGDSYVDTGNFVNSGSYKPP 260
Query: 59 KSPYGDTYFHRPAGRYSDGRIILDF 83
G T+ +PAGR+ DGR++ D+
Sbjct: 261 S---GITFPGKPAGRFCDGRVLTDY 326
>CB829316
Length = 555
Score = 35.8 bits (81), Expect = 0.001
Identities = 20/47 (42%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Frame = +1
Query: 38 IFNLGASNSDTGGLAA-AFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+F G S DTG A K PYG T+ P GR+SDGR++ D+
Sbjct: 214 LFVFGDSYVDTGNTRKDAPGSWKEPYGITFPGEPTGRFSDGRVLTDY 354
>BI420239
Length = 621
Score = 33.1 bits (74), Expect = 0.009
Identities = 21/78 (26%), Positives = 35/78 (43%), Gaps = 3/78 (3%)
Frame = +1
Query: 8 RHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG---LAAAFLPPKSPYGD 64
R + ++L ++L + ++ P +F G S SD+G L PYG
Sbjct: 1 RTLLVLALVILLVAIAIRQYCLQRNSQVPCLFIFGDSLSDSGNNNNLETDAKVDYLPYGI 180
Query: 65 TYFHRPAGRYSDGRIILD 82
+ P GRY++GR +D
Sbjct: 181DFPTGPTGRYTNGRNAID 234
>TC18753
Length = 562
Score = 32.3 bits (72), Expect = 0.016
Identities = 24/79 (30%), Positives = 37/79 (46%), Gaps = 3/79 (3%)
Frame = +3
Query: 7 HRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG---LAAAFLPPKSPYG 63
H + L V L++TT V A+ + P F+ G S D G L++ PYG
Sbjct: 45 HGEFIVLVLIVCLWSTTTTGVGAEPQV--PCFFSFGDSLVDNGNNNRLSSLAKANYRPYG 218
Query: 64 DTYFHRPAGRYSDGRIILD 82
+ P GR+S+G+ +D
Sbjct: 219 IDFPGGPTGRFSNGKTSVD 275
>TC14230
Length = 740
Score = 31.2 bits (69), Expect = 0.035
Identities = 25/76 (32%), Positives = 36/76 (46%), Gaps = 4/76 (5%)
Frame = +1
Query: 11 SFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG---LAAAFLPPKSPYG-DTY 66
+ SL ++LF ++++ + A F G S D+G LA PYG D
Sbjct: 76 TITSLVLVLFLSSVSAQHTR------AFFVFGDSLVDSGNNDFLATTARADAPPYGIDFP 237
Query: 67 FHRPAGRYSDGRIILD 82
HRP GR+S+G I D
Sbjct: 238 THRPTGRFSNGLNIPD 285
>TC10793
Length = 556
Score = 30.4 bits (67), Expect = 0.060
Identities = 21/52 (40%), Positives = 26/52 (49%), Gaps = 4/52 (7%)
Frame = +1
Query: 36 PAIFNLGASNSDTGGLAAAFLPPKS---PYG-DTYFHRPAGRYSDGRIILDF 83
PAI G S+ D G +S PYG D +P GR+S+GRI DF
Sbjct: 133 PAIIVFGDSSVDAGNNNFIETVARSNFQPYGRDFQGGKPTGRFSNGRIATDF 288
>BP041040
Length = 550
Score = 30.0 bits (66), Expect = 0.078
Identities = 22/54 (40%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Frame = +3
Query: 36 PAIFNLGASNSDTGGLAAAFLPPKS-----PYG-DTYFHRPAGRYSDGRIILDF 83
PAI G S+ D+G F+P + PYG D P GR+S+GRI DF
Sbjct: 81 PAIIVFGDSSVDSGN--NNFIPTMAKSNFEPYGRDFPDGNPTGRFSNGRIAPDF 236
>BP034926
Length = 607
Score = 29.6 bits (65), Expect = 0.10
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 4/56 (7%)
Frame = +2
Query: 32 RCNFPAIFNLGASNSDTGGLAAAFLPPKS---PYGDTYFHR-PAGRYSDGRIILDF 83
+ N I G S+ D G KS PYG +F+ P GR+ +GR+ DF
Sbjct: 158 KSNVSCILVFGDSSVDPGNNNVLRTSMKSNFPPYGKDFFNSLPTGRFCNGRLATDF 325
>AV429155
Length = 393
Score = 27.7 bits (60), Expect = 0.39
Identities = 21/74 (28%), Positives = 32/74 (42%), Gaps = 3/74 (4%)
Frame = +3
Query: 12 FVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS---PYGDTYFH 68
F+ L +++ + V K P +F G S SD G KS PYG +
Sbjct: 45 FLPLLLLVANCMQHSVHGKPEV--PCLFIFGDSLSDNGNNNNLVTSTKSNYKPYGIDFPT 218
Query: 69 RPAGRYSDGRIILD 82
P GR+++G +D
Sbjct: 219 GPTGRFTNGPTTVD 260
>BP040561
Length = 515
Score = 27.3 bits (59), Expect = 0.51
Identities = 20/50 (40%), Positives = 23/50 (46%), Gaps = 4/50 (8%)
Frame = +1
Query: 37 AIFNLGASNSDTGG---LAAAFLPPKSPYG-DTYFHRPAGRYSDGRIILD 82
A F G S D G LA PYG D H+P GR+S+G I D
Sbjct: 70 AFFVFGDSLVDNGNNDFLATTARADAPPYGIDFPTHKPTGRFSNGLNIPD 219
>AU240152
Length = 300
Score = 26.9 bits (58), Expect = 0.66
Identities = 16/50 (32%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Frame = +1
Query: 36 PAIFNLGASNSDTGG---LAAAFLPPKSPYGDTYFHRPAGRYSDGRIILD 82
P +F G S SD G L + +PYG + P GR+++G +D
Sbjct: 127 PCLFIFGDSLSDDGNNNNLPTSTKSNYNPYGIDFPTGPTGRFTNGPTAID 276
>TC17974 UP|AAO45821 (AAO45821) Gamma-glutamylcysteine synthetase, complete
Length = 2018
Score = 26.9 bits (58), Expect = 0.66
Identities = 15/38 (39%), Positives = 17/38 (44%)
Frame = +1
Query: 7 HRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGAS 44
H H + SLF FT NP F P I LG+S
Sbjct: 82 HSHTTSSSLFSSQFTLHFNPNFNFSFFTMPVISRLGSS 195
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,548,475
Number of Sequences: 28460
Number of extensions: 18727
Number of successful extensions: 176
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 170
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 174
length of query: 83
length of database: 4,897,600
effective HSP length: 59
effective length of query: 24
effective length of database: 3,218,460
effective search space: 77243040
effective search space used: 77243040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0136.17


