

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
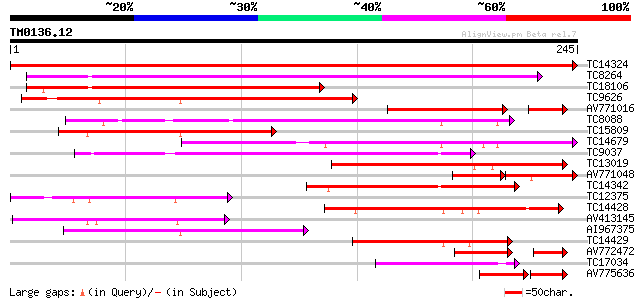
Query= TM0136.12
(245 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14324 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan pro... 479 e-136
TC8264 similar to GB|AAK20861.1|13377784|AF333974 fasciclin-like... 184 2e-47
TC18106 similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-li... 147 2e-36
TC9626 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan prot... 142 4e-35
AV771016 103 6e-27
TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like arabinogalac... 101 1e-22
TC15809 weakly similar to PIR|JN0747|JN0747 histone H1-I - Volvo... 99 7e-22
TC14679 similar to UP|FLA7_ARATH (Q9SJ81) Fasciclin-like arabino... 98 1e-21
TC9037 similar to GB|AAG24276.1|10880493|AF195889 fasciclin-like... 94 2e-20
TC13019 85 1e-17
AV771048 57 2e-17
TC14342 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan pro... 82 6e-17
TC12375 weakly similar to GB|AAA59989.1|189479|HUMP53T p53 cellu... 79 7e-16
TC14428 weakly similar to UP|Q9SP59 (Q9SP59) Arabinogalactan pro... 79 9e-16
AV413145 76 6e-15
AI967375 76 6e-15
TC14429 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan pro... 65 8e-12
AV772472 45 5e-09
TC17034 56 6e-09
AV775636 40 4e-08
>TC14324 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan protein-like,
partial (63%)
Length = 1172
Score = 479 bits (1234), Expect = e-136
Identities = 244/245 (99%), Positives = 244/245 (99%)
Frame = +3
Query: 1 MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQ 60
MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQ
Sbjct: 102 MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQ 281
Query: 61 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 120
VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF
Sbjct: 282 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 461
Query: 121 DTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLL 180
DTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLL
Sbjct: 462 DTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLL 641
Query: 181 PRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVVA 240
PRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPL VAMVVVA
Sbjct: 642 PRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLPVAMVVVA 821
Query: 241 AFSSW 245
AFSSW
Sbjct: 822 AFSSW 836
Score = 26.9 bits (58), Expect = 3.2
Identities = 14/38 (36%), Positives = 21/38 (54%), Gaps = 2/38 (5%)
Frame = -3
Query: 190 PAPAPAPEKAKASKKKSADDTEGPA--GDDSAAVSVKQ 225
P+P+PAPE A A+ ++ + + E P G A KQ
Sbjct: 702 PSPSPAPEPAPAA*QRRSHEGEAPCLLGKPQAGYQSKQ 589
>TC8264 similar to GB|AAK20861.1|13377784|AF333974 fasciclin-like
arabinogalactan-protein 9 {Arabidopsis thaliana;},
partial (38%)
Length = 942
Score = 184 bits (466), Expect = 2e-47
Identities = 99/223 (44%), Positives = 134/223 (59%)
Frame = +3
Query: 8 LPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINA 67
L L+ L F + + A +PAPAPS P ++ IL+KAG YTTLIRLL+ +Q TQI +
Sbjct: 36 LALILLTFLSLFASKIQAQAPAPAPSG-PVNLTAILEKAGQYTTLIRLLKESQQLTQIES 212
Query: 68 QLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPV 127
QL ++ G T FAP DNAF NLK G +N L D QK +LI +H+ P + S+S+ T+SNPV
Sbjct: 213 QLNSTTQGFTLFAPTDNAFQNLKSGAINDLTDDQKVKLILYHVTPKYYSLSDLQTVSNPV 392
Query: 128 RTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA 187
RTQA E LN GN VN+TTG+V ++ + LA+YQVD+VLLP + F A
Sbjct: 393 RTQASEKEGSWGLNFKGQGNQVNVTTGVVTTSINNDLRQQFPLAIYQVDRVLLPLELFGA 572
Query: 188 KPPAPAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWG 230
K P+ AP+P+ ++ + + G S A S K V G
Sbjct: 573 KSPSSAPSPKSSETTPSDETPSSGKKGGAPSPAASQKDNGVAG 701
>TC18106 similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-like, partial
(42%)
Length = 436
Score = 147 bits (370), Expect = 2e-36
Identities = 78/130 (60%), Positives = 98/130 (75%), Gaps = 1/130 (0%)
Frame = +3
Query: 8 LPLLAL-MFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQIN 66
LPLL L MF F T A ++ APAP+ PT+I +IL+KAG +TT I++L+ TQVA +IN
Sbjct: 36 LPLLLLNMFLFLFQTISAQTAAAPAPAG-PTNITQILEKAGQFTTFIKILKATQVADRIN 212
Query: 67 AQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNP 126
+QL NSN GLT FAP DNAFS+LKPG LNS+N Q + +LIQFH+LPT SMS F+T SNP
Sbjct: 213 SQLNNSNQGLTIFAPTDNAFSSLKPGTLNSINTQDQLQLIQFHILPTLYSMSQFETASNP 392
Query: 127 VRTQAGENPD 136
+ TQAG + D
Sbjct: 393 LHTQAGNSGD 422
>TC9626 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan protein-like,
partial (24%)
Length = 605
Score = 142 bits (359), Expect = 4e-35
Identities = 82/176 (46%), Positives = 107/176 (60%), Gaps = 31/176 (17%)
Frame = +1
Query: 6 FTLPLLALMFFFHSTTTLAASSPAPAPSSSPT---------------------------- 37
F+L L A +FF TT++A SPA +P SP
Sbjct: 88 FSLLLFAPIFF----TTISAQSPAASPKKSPAKPSPASLAPAPAKPLVPSLPQSPSSDSS 255
Query: 38 --DIIRILKKAGGYTTLIRLLQTTQVATQINAQLINS-NAGLTFFAPNDNAFSNLKPGFL 94
DII+IL+KA + TLIRLL+TTQ+ Q+NAQL+ + N GLT AP+D AFS LK G+
Sbjct: 256 GQDIIKILRKAKSFNTLIRLLKTTQIINQVNAQLVTTKNGGLTILAPDDGAFSQLKAGYF 435
Query: 95 NSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVN 150
NSL+ +Q+ ELIQFH+LP +VS SNFD LSNPV T A ++P +NVT+ GN+VN
Sbjct: 436 NSLDGRQQKELIQFHVLPQYVSSSNFDALSNPVLTLASDSPKGYQINVTAYGNSVN 603
>AV771016
Length = 506
Score = 103 bits (256), Expect(2) = 6e-27
Identities = 50/52 (96%), Positives = 50/52 (96%)
Frame = -3
Query: 164 VYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAG 215
VYSDNQL VY VDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAG
Sbjct: 504 VYSDNQLPVYLVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAG 349
Score = 33.5 bits (75), Expect(2) = 6e-27
Identities = 15/17 (88%), Positives = 16/17 (93%)
Frame = -1
Query: 225 QRLVWGPLAVAMVVVAA 241
+RLVWGPL VAMVVVAA
Sbjct: 317 ERLVWGPLPVAMVVVAA 267
>TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (69%)
Length = 1527
Score = 101 bits (252), Expect = 1e-22
Identities = 67/209 (32%), Positives = 109/209 (52%), Gaps = 15/209 (7%)
Frame = +1
Query: 25 ASSPAPAPSSSPTDI--IRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPN 82
+S+ A AP+++P+DI I I+ K G LL+T++ N N GLT F P
Sbjct: 586 SSADAEAPTAAPSDIDVISIMSKQG-CKAFADLLRTSKALPTFKE---NVNGGLTVFCPT 753
Query: 83 DNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNV 142
D+A + P + N L D QK L+ +H +P + S+ + + + T A E ++ V
Sbjct: 754 DSAVNGFLPKYKN-LTDSQKVSLLLYHGVPVYQSLQMLKSNNGIMNTLATEGANKYDFTV 930
Query: 143 TSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF-------VAKPPAPAPA 195
+ G VN+ T + + G++ + VY++ KVL+PR+ F V+ +P PA
Sbjct: 931 QNDGEDVNLETKVNTAGIIGTLIDQDPFVVYKISKVLMPRELFKGVTEKTVSPAESPKPA 1110
Query: 196 PEKAKASKKKSADD------TEGPAGDDS 218
+K+K++KK +ADD +GPA DDS
Sbjct: 1111KKKSKSTKKGNADDDAADSPADGPADDDS 1197
>TC15809 weakly similar to PIR|JN0747|JN0747 histone H1-I - Volvox carteri
{Volvox carteri;}, partial (10%)
Length = 526
Score = 99.0 bits (245), Expect = 7e-22
Identities = 53/107 (49%), Positives = 73/107 (67%), Gaps = 13/107 (12%)
Frame = +2
Query: 22 TLAASSPAPAP------------SSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQL 69
T A++P+P P S+P DI +IL+KA ++ LIRLL+TT++ IN+QL
Sbjct: 206 TPKATAPSPKPLVPTLPQSPDSSDSTPDDITKILRKAKIFSVLIRLLKTTEIMNNINSQL 385
Query: 70 INS-NAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFV 115
I + N G+T AP+D+AFS+LK GFLNSLN+ QK EL QFH+LP +V
Sbjct: 386 ITAKNGGITILAPDDSAFSHLKAGFLNSLNENQKIELCQFHILPQYV 526
>TC14679 similar to UP|FLA7_ARATH (Q9SJ81) Fasciclin-like arabinogalactan
protein 7 precursor, partial (50%)
Length = 724
Score = 98.2 bits (243), Expect = 1e-21
Identities = 65/183 (35%), Positives = 95/183 (51%), Gaps = 12/183 (6%)
Frame = +2
Query: 75 GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGEN 134
G+T F P D++FS LK L+ L Q ++I FH LP + S+++F LS Q G
Sbjct: 8 GITIFVPKDSSFSALKKPSLSKLTSDQLKQVILFHALPKYYSLADFKNLS-----QTGST 172
Query: 135 P----DRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF---VA 187
P +LN T TV++ +G V +V+S + +A+Y+V KVLLP F +
Sbjct: 173 PTFAGGSYSLNFTDDSGTVHINSGWSKTKVTSAVHSTDPVAIYEVGKVLLPEAVFGTDIP 352
Query: 188 KPPAPAPAPEKAKASK---KKSADD--TEGPAGDDSAAVSVKQRLVWGPLAVAMVVVAAF 242
PAPAPAPE A A+ +KSAD + + D S++ + +WG L +A V A
Sbjct: 353 PTPAPAPAPEIAPAADSPIEKSADSKASSPSSPDGSSSYKILSYGIWGNLVLATFGVLAM 532
Query: 243 SSW 245
W
Sbjct: 533 L*W 541
>TC9037 similar to GB|AAG24276.1|10880493|AF195889 fasciclin-like
arabinogalactan protein FLA8 {Arabidopsis thaliana;} ,
partial (61%)
Length = 989
Score = 94.4 bits (233), Expect = 2e-20
Identities = 58/173 (33%), Positives = 94/173 (53%)
Frame = +2
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
AP PSSS ++ +++KAG T +L + T +A ++ GLT FAP+D AF
Sbjct: 38 APPPSSS-VNLTALIEKAGCKTFASLVLSNGLIKTFQSA----ADKGLTIFAPSDEAFKA 202
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L+ L + + L+Q+H + ++ + + T +P+ T A + V+ +G++
Sbjct: 203 RGVPDLSKLTNAEVVSLLQYHAVAKYLPVGSLKTTKDPISTLATNGAGKFEYTVSVAGDS 382
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKA 201
V + TG+ + V +V L++Y VD VLLP + F AK P+PAPAPE A A
Sbjct: 383 VTLHTGVDSSRVADTVLDSTPLSIYSVDSVLLPPELF-AKSPSPAPAPEPASA 538
>TC13019
Length = 565
Score = 85.1 bits (209), Expect = 1e-17
Identities = 50/106 (47%), Positives = 69/106 (64%), Gaps = 4/106 (3%)
Frame = +1
Query: 140 LNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKA 199
LNVT+ GN+VN++TG+V+VT+ G VY D LA+Y+VDKVL+P DF KP APAPA KA
Sbjct: 7 LNVTAYGNSVNISTGVVDVTITGIVYFDKTLAIYRVDKVLIPLDFKKPKPIAPAPALAKA 186
Query: 200 -KASKKKSA---DDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVVAA 241
KA K+ S+ DD +G +S+ + + G V +V+AA
Sbjct: 187 PKADKENSSAEDDDDQGQGTKNSSGANRLISSIRGTTLVLSLVLAA 324
>AV771048
Length = 430
Score = 56.6 bits (135), Expect(2) = 2e-17
Identities = 29/32 (90%), Positives = 30/32 (93%), Gaps = 1/32 (3%)
Frame = -3
Query: 215 GDDSAAVSVK-QRLVWGPLAVAMVVVAAFSSW 245
GDDSAAVSVK +RLVWGPL VAMVVVAAFSSW
Sbjct: 359 GDDSAAVSVKHERLVWGPLPVAMVVVAAFSSW 264
Score = 47.8 bits (112), Expect(2) = 2e-17
Identities = 22/23 (95%), Positives = 22/23 (95%)
Frame = -2
Query: 192 PAPAPEKAKASKKKSADDTEGPA 214
PAPAPEKAKASKKKSADD EGPA
Sbjct: 429 PAPAPEKAKASKKKSADDPEGPA 361
>TC14342 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-like,
partial (27%)
Length = 724
Score = 82.4 bits (202), Expect = 6e-17
Identities = 45/93 (48%), Positives = 59/93 (63%), Gaps = 1/93 (1%)
Frame = +2
Query: 129 TQAGENPD-RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA 187
TQAG + D LNVT+SGN V++TTG+V TV ++YSD LAVYQV+KVLLP F
Sbjct: 2 TQAGNSGDGEYPLNVTTSGNQVHITTGVVETTVSNTIYSDKTLAVYQVEKVLLPLALF-G 178
Query: 188 KPPAPAPAPEKAKASKKKSADDTEGPAGDDSAA 220
+ P PAPA A +K+ ++ P+G S A
Sbjct: 179 QAPTPAPAEAPAPTQPEKNVRASDSPSGSSSDA 277
>TC12375 weakly similar to GB|AAA59989.1|189479|HUMP53T p53 cellular tumor
antigen {Homo sapiens;} , partial (6%)
Length = 468
Score = 79.0 bits (193), Expect = 7e-16
Identities = 55/133 (41%), Positives = 68/133 (50%), Gaps = 37/133 (27%)
Frame = +1
Query: 1 MKHQLFTLPLLALMFFFHSTTTLAAS-------------------SPAPAPS-------- 33
MK LF+ LA +F S TTLA S +PAPAPS
Sbjct: 79 MKQILFSFLFLAPIF---SITTLAQSPAAAPKPPAKPAPTKPSPATPAPAPSKPLVPALP 249
Query: 34 ---------SSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLI-NSNAGLTFFAPND 83
S D+++IL+KA + TLIRLL+TTQ+ Q+NAQL+ N GLT AP+D
Sbjct: 250 QSPMVNPDSSGNQDVVKILRKAKSFNTLIRLLKTTQIINQVNAQLVATKNGGLTILAPDD 429
Query: 84 NAFSNLKPGFLNS 96
AFS LK GF NS
Sbjct: 430 GAFSQLKAGFFNS 468
>TC14428 weakly similar to UP|Q9SP59 (Q9SP59) Arabinogalactan protein
Pop14A9, partial (15%)
Length = 573
Score = 78.6 bits (192), Expect = 9e-16
Identities = 53/111 (47%), Positives = 70/111 (62%), Gaps = 8/111 (7%)
Frame = +1
Query: 137 RLALNVTSSGNT-VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFV-AKPPAPAP 194
RL LNV + GN+ VN++TG+VN ++ G VYSD LAVY VDKVLLP DFF+ AK PA AP
Sbjct: 1 RLPLNVNALGNSIVNISTGVVNASIIGVVYSDRNLAVYHVDKVLLPLDFFLTAKAPALAP 180
Query: 195 -----APEKAKA-SKKKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVV 239
AP+ AK S + D+T + S AVS+ + + L VA+V +
Sbjct: 181 SLSAKAPKAAKENSSAEDEDETNQDQDNKSGAVSL-VKTTFMSLGVALVAI 330
>AV413145
Length = 397
Score = 75.9 bits (185), Expect = 6e-15
Identities = 50/127 (39%), Positives = 66/127 (51%), Gaps = 33/127 (25%)
Frame = +1
Query: 2 KHQLFTLPLLALMFFFHSTTTLAASSPAPAP----------------------------S 33
K L +L L L+ +STTTLA SPA AP S
Sbjct: 16 KQSLLSLSLALLVSLLYSTTTLAQLSPASAPLQPAPPVPTTPASAPKPLVPSLPESPSDS 195
Query: 34 SSP----TDIIRILKKAGGYTTLIRLLQTTQVATQINAQLIN-SNAGLTFFAPNDNAFSN 88
++P DI+ IL+KA + LIRL++TTQ+ Q+N+QL+ GLT AP+D+AFS
Sbjct: 196 TAPDTAAVDIVGILRKAKSFNVLIRLMKTTQLINQLNSQLLTIKTGGLTILAPDDSAFSE 375
Query: 89 LKPGFLN 95
LK GFLN
Sbjct: 376 LKAGFLN 396
>AI967375
Length = 397
Score = 75.9 bits (185), Expect = 6e-15
Identities = 45/107 (42%), Positives = 63/107 (58%), Gaps = 1/107 (0%)
Frame = +1
Query: 24 AASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINS-NAGLTFFAPN 82
A+++ +P+ SS DII+IL+KA + TLIRLL+TTQ+ Q+ AQL+ + N GLT AP+
Sbjct: 64 ASATQSPSSDSSGQDIIKILRKAKSFNTLIRLLKTTQIINQVXAQLVTTKNGGLTILAPD 243
Query: 83 DNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRT 129
D AF + K G S K L + NFD L++PV T
Sbjct: 244 DGAFHSSKLGTQLSGRSPTKGTDTVPMFLHNMCQVPNFDGLNHPVLT 384
>TC14429 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-like,
partial (19%)
Length = 530
Score = 65.5 bits (158), Expect = 8e-12
Identities = 37/72 (51%), Positives = 48/72 (66%), Gaps = 3/72 (4%)
Frame = +2
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFV-AKPPAPAPAPE--KAKASKKK 205
VN++TG+VN ++ G VYSD LAVY VDKVLLP DFF+ AK PA AP+ KA+K+
Sbjct: 5 VNISTGVVNASIIGVVYSDRNLAVYHVDKVLLPLDFFLTAKAPALAPSLSA*APKAAKEN 184
Query: 206 SADDTEGPAGDD 217
S+ + E D
Sbjct: 185 SSAEDEDETNQD 220
>AV772472
Length = 423
Score = 45.1 bits (105), Expect(2) = 5e-09
Identities = 20/25 (80%), Positives = 23/25 (92%)
Frame = -3
Query: 193 APAPEKAKASKKKSADDTEGPAGDD 217
APAPEKAKAS+KKS DD EGPAG++
Sbjct: 421 APAPEKAKASEKKSHDDPEGPAGEE 347
Score = 30.8 bits (68), Expect(2) = 5e-09
Identities = 14/15 (93%), Positives = 14/15 (93%)
Frame = -2
Query: 227 LVWGPLAVAMVVVAA 241
LVWGPL VAMVVVAA
Sbjct: 317 LVWGPLPVAMVVVAA 273
>TC17034
Length = 519
Score = 55.8 bits (133), Expect = 6e-09
Identities = 30/62 (48%), Positives = 37/62 (59%)
Frame = +2
Query: 159 TVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDS 218
T+ G VYSD LA+Y VD+VL+P DF K PAPAP A S K ++ D + GDD
Sbjct: 5 TLTGIVYSDKTLAIYHVDQVLVPLDFSKPKSPAPAPTLANAPESDKDNSSDED---GDDQ 175
Query: 219 AA 220
A
Sbjct: 176 GA 181
>AV775636
Length = 412
Score = 40.0 bits (92), Expect(2) = 4e-08
Identities = 18/21 (85%), Positives = 20/21 (94%)
Frame = -3
Query: 204 KKSADDTEGPAGDDSAAVSVK 224
+ SADD+EGPAGDDSAAVSVK
Sbjct: 386 RNSADDSEGPAGDDSAAVSVK 324
Score = 32.7 bits (73), Expect(2) = 4e-08
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = -1
Query: 226 RLVWGPLAVAMVVVAA 241
RLVWGPL VAMVVVAA
Sbjct: 319 RLVWGPLPVAMVVVAA 272
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.130 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,726,056
Number of Sequences: 28460
Number of extensions: 52347
Number of successful extensions: 874
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 730
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 838
length of query: 245
length of database: 4,897,600
effective HSP length: 88
effective length of query: 157
effective length of database: 2,393,120
effective search space: 375719840
effective search space used: 375719840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0136.12


