

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
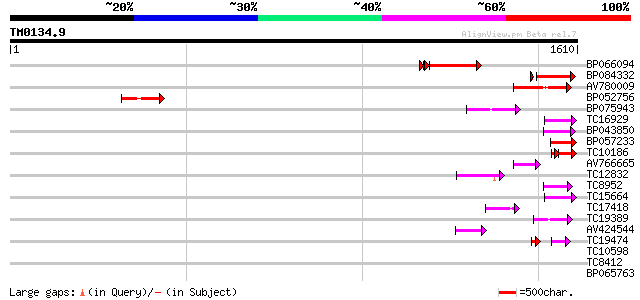
Query= TM0134.9
(1610 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP066094 279 2e-80
BP084332 229 4e-60
AV780009 132 3e-31
BP052756 89 5e-18
BP075943 75 1e-13
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 72 7e-13
BP043850 72 9e-13
BP057233 71 2e-12
TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprot... 50 8e-11
AV766665 61 2e-09
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 59 5e-09
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 57 2e-08
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 57 3e-08
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 53 3e-07
TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 52 7e-07
AV424544 52 7e-07
TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retro... 37 2e-05
TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526 At2g01... 38 0.011
TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromos... 36 0.042
BP065763 35 0.093
>BP066094
Length = 532
Score = 279 bits (714), Expect(3) = 2e-80
Identities = 137/149 (91%), Positives = 142/149 (94%)
Frame = +2
Query: 1191 ARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKK 1250
++ EAIRLLISFSVNHNI+LHQM+VKSAFLNGYISEEVYVHQPPG EDEK DH+FKLKK
Sbjct: 86 SKTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKK 265
Query: 1251 SLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQ 1310
SLYGLKQAPRAWYERLSSFLLENE VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSAN
Sbjct: 266 SLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANP 445
Query: 1311 SLCKEFSEMMQAEFEMSMMGELKYFLGIQ 1339
SLCKEFSEMMQAEFEM MMGELKYFLGIQ
Sbjct: 446 SLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Score = 37.4 bits (85), Expect(3) = 2e-80
Identities = 17/18 (94%), Positives = 17/18 (94%)
Frame = +3
Query: 1175 YSQQEGIDYTETFAPVAR 1192
YSQQ GIDYTETFAPVAR
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
Score = 23.1 bits (48), Expect(3) = 2e-80
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = +1
Query: 1165 RNKARLVAQG 1174
RNKARLVAQG
Sbjct: 10 RNKARLVAQG 39
>BP084332
Length = 368
Score = 229 bits (583), Expect(2) = 4e-60
Identities = 112/112 (100%), Positives = 112/112 (100%)
Frame = +2
Query: 1495 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 1554
QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 211
Query: 1555 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 1606
RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL
Sbjct: 212 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 367
Score = 21.6 bits (44), Expect(2) = 4e-60
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = +1
Query: 1480 CQFLGSNL 1487
CQFLGSNL
Sbjct: 4 CQFLGSNL 27
>AV780009
Length = 529
Score = 132 bits (333), Expect = 3e-31
Identities = 68/168 (40%), Positives = 105/168 (62%), Gaps = 6/168 (3%)
Frame = -1
Query: 1432 AVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWA 1491
A R+LRY+KG GL + KL Y D+D+AG R+S +G FLG++L+SW
Sbjct: 526 AATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWR 347
Query: 1492 SKRQSTIALSTAEAEY--ISAAICSTQMLWMKHQLEDYQILESN----IPIYCDNTAAIS 1545
+K+Q+T++ S++EAEY ++A +C Q W+ + +Q L+ N +P++CDN +A+
Sbjct: 346 TKKQTTVSRSSSEAEYRALAATVCEVQ--WLSYL---FQFLKLNVPLPVPLFCDNQSALH 182
Query: 1546 LSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKP 1593
++ NP H R KHIE+ H +R +Q G++ L + T HQ ADIFTKP
Sbjct: 181 IAHNPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKP 38
>BP052756
Length = 488
Score = 89.0 bits (219), Expect = 5e-18
Identities = 54/124 (43%), Positives = 84/124 (67%), Gaps = 2/124 (1%)
Frame = +1
Query: 318 KKPKKHFKTKKSLMVTFDESESEDVDS--DGEVQGLMAIVKDKGAESKEAVDSDSESEGD 375
K+P+K F TKK+L +TFD ++S++ +S D V+ LMA +GAE+ SE D
Sbjct: 151 KRPRKVF-TKKAL-ITFDATDSDEAESEEDEAVEALMATTS-RGAEA---------SEDD 294
Query: 376 PDSDDENEVFASFSTSELKHALSDIMDKYNSLLSKHKKLKKNLSAVSKTPSEHEKIISDL 435
DS++++EVF+ FS SEL+ ALS+IM K++ LL+KHK+LKK + + + HEK +S+L
Sbjct: 295 SDSENDDEVFSDFSLSELRTALSEIMKKHDILLNKHKELKKKYAVKTSSSEVHEKSVSEL 474
Query: 436 KNDN 439
++N
Sbjct: 475 IDEN 486
>BP075943
Length = 547
Score = 74.7 bits (182), Expect = 1e-13
Identities = 44/155 (28%), Positives = 76/155 (48%), Gaps = 2/155 (1%)
Frame = -3
Query: 1298 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1357
+YVDD++ + + + +F + +GE KYFLG+++ ++ G ++Q KY +
Sbjct: 515 LYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYALQ 336
Query: 1358 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKL--YRGMIGSLLYLTASRPDILFSV 1415
L+ + H T +G + YR ++G LLYL +RPDI F+V
Sbjct: 335 LISDSGHF-GFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDITFAV 159
Query: 1416 HLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMY 1450
+ ++F S P + H + L+YL G+ GL Y
Sbjct: 158 NQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFY 54
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 72.0 bits (175), Expect = 7e-13
Identities = 32/91 (35%), Positives = 54/91 (59%), Gaps = 1/91 (1%)
Frame = +1
Query: 1519 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLL 1577
W+ + L+D ++ E +YCDN +A ++ NP+ H R KHIE+ H +R+ +QKG++ L
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 1578 KFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
+ + ADI+TK L+ F+ I L +
Sbjct: 184 LPISSSEPLADIYTKALSPQNFHQICAKLGL 276
>BP043850
Length = 515
Score = 71.6 bits (174), Expect = 9e-13
Identities = 34/93 (36%), Positives = 55/93 (58%), Gaps = 1/93 (1%)
Frame = -1
Query: 1515 TQMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKG 1573
++++W++ L + L+S P++ DNT+AI ++ NP+ H +HIEV H +R+ +
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 1574 VLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 1606
V+ L V T Q ADI TK L R NF++ L
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKL 237
>BP057233
Length = 473
Score = 70.9 bits (172), Expect = 2e-12
Identities = 31/73 (42%), Positives = 50/73 (68%)
Frame = -2
Query: 1536 IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLA 1595
++CDN +A +L+ NP+LH+R+KHIE+ H+IRD V + +++ +V T Q AD TKPL+
Sbjct: 445 LWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIADCLTKPLS 266
Query: 1596 EDRFNFILKNLNM 1608
RF+ + L +
Sbjct: 265 HTRFSQLRDKLGV 227
>TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein, partial
(4%)
Length = 528
Score = 49.7 bits (117), Expect(2) = 8e-11
Identities = 26/51 (50%), Positives = 32/51 (61%)
Frame = +3
Query: 1558 HIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
+IE KYHF+RD V KG + LK TD Q ADI TK L +RF + LN+
Sbjct: 69 NIETKYHFLRDQVTKGKISLKHCGTDLQVADIMTKGLKTERFRNMRAMLNV 221
Score = 35.4 bits (80), Expect(2) = 8e-11
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = +2
Query: 1539 DNTAAISLSKNPILHSRAKHIE 1560
DN +AI L+KNP+ H R+KHIE
Sbjct: 5 DNKSAIDLAKNPVSHGRSKHIE 70
>AV766665
Length = 601
Score = 60.8 bits (146), Expect = 2e-09
Identities = 32/76 (42%), Positives = 44/76 (57%)
Frame = +3
Query: 1431 TAVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSW 1490
T RI YLK + G +++K + + G+ ADY G +R ST G FL NLV+W
Sbjct: 360 TFTTRIF*YLKANSRRGPLFQKEGKSSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTW 539
Query: 1491 ASKRQSTIALSTAEAE 1506
SK+Q+ IA S+ EAE
Sbjct: 540 RSKQQNIIARSSGEAE 587
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 59.3 bits (142), Expect = 5e-09
Identities = 37/145 (25%), Positives = 71/145 (48%), Gaps = 9/145 (6%)
Frame = +2
Query: 1268 SFLLENEFVRGKVDTTLFCKTYKD-DILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEM 1326
SF++ + R D + K + D D +I+ +YVDD++ N+ +E + EF+M
Sbjct: 8 SFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDM 187
Query: 1327 SMMGELKYFLGIQV--DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTP------MHPTC 1378
+G LG+Q+ D+ ++ Q Y +++L++FNM + TP + +
Sbjct: 188 KDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSM 367
Query: 1379 ILEKEDKSGKVCQKLYRGMIGSLLY 1403
I E + ++ + Y +GSL+Y
Sbjct: 368 IPSSEAERMEMSRVPYASAVGSLMY 442
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 57.4 bits (137), Expect = 2e-08
Identities = 29/82 (35%), Positives = 48/82 (58%), Gaps = 1/82 (1%)
Frame = +3
Query: 1516 QMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGV 1574
+ LW+ + L D +I + IYCDN +A+ L+ N + H R ++IE+ H + V G+
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 1575 LLLKFVDTDHQWADIFTKPLAE 1596
L L V + Q AD+FTK +++
Sbjct: 201 LHLLHVPSSDQVADVFTKTISQ 266
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 56.6 bits (135), Expect = 3e-08
Identities = 27/91 (29%), Positives = 51/91 (55%), Gaps = 1/91 (1%)
Frame = +1
Query: 1519 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLL 1577
W+ + L+D ++ S +YCD+ +A ++ N + H R KH+++ H +R+ +Q + L
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 1578 KFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
+ + Q ADI TKPL F+ ++ L +
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKLGV 285
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 53.1 bits (126), Expect = 3e-07
Identities = 36/99 (36%), Positives = 49/99 (49%), Gaps = 1/99 (1%)
Frame = -1
Query: 1351 QSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGK-VCQKLYRGMIGSLLYLTASRP 1409
Q Y ++LKKF M S T + I GK V Y+ +IGS+ YL R
Sbjct: 739 QKIYVDDILKKFKMTNSKYISTTIGGKEIEAGRRNGGKRVDSTYYKSLIGSVRYLNTVRS 560
Query: 1410 DILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGL 1448
DI+ V L +RF +P + H +R LRY+KGT G+
Sbjct: 559 DIVCGVGLRSRFM-EP*DCH*QGAQRSLRYIKGTLKDGI 446
>TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (6%)
Length = 498
Score = 52.0 bits (123), Expect = 7e-07
Identities = 35/114 (30%), Positives = 59/114 (51%), Gaps = 3/114 (2%)
Frame = +3
Query: 1488 VSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYC---DNTAAI 1544
V+W S+ Q +ALSTAEAE+I+A ++LWMK+ L++ + PI C A
Sbjct: 27 VAWPSRLQKCVALSTAEAEFIAATEACHELLWMKNFLQNAWF--HSHPILCCIVITKALF 200
Query: 1545 SLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDR 1598
+L++ + + +RD + +L L+ + TD AD+ TK L ++
Sbjct: 201 TLARILLFIQDPSTLMFVIIGLRDVLNSKLLELEKIHTDDDGADMMTKSLPREK 362
>AV424544
Length = 276
Score = 52.0 bits (123), Expect = 7e-07
Identities = 26/90 (28%), Positives = 49/90 (53%)
Frame = +3
Query: 1265 RLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEF 1324
+LSS+L +++ D +LF K ++ +YVDD+I + + + + +F
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 1325 EMSMMGELKYFLGIQVDQTPEGTYIHQSKY 1354
+ +G LKYFLG++V ++ G ++ Q KY
Sbjct: 186 RIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 517
Score = 37.0 bits (84), Expect(2) = 2e-05
Identities = 21/54 (38%), Positives = 32/54 (58%)
Frame = -2
Query: 1539 DNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTK 1592
DN AA+ + K RAKHIE+ +HF+ D + +FV+++ Q D+FTK
Sbjct: 354 DNKAALHMPKI*F-SMRAKHIEIDFHFL*DRRLYLDISTRFVNSNDQLTDVFTK 196
Score = 29.6 bits (65), Expect(2) = 2e-05
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -1
Query: 1483 LGSNLVSWASKRQSTIALSTAEAEY 1507
L NL+SW SK +A S AEAE+
Sbjct: 517 LSDNLISWKSKETIIVARSRAEAEF 443
>TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526
At2g01100/F23H14.7 {Arabidopsis thaliana;}, partial
(33%)
Length = 902
Score = 38.1 bits (87), Expect = 0.011
Identities = 49/174 (28%), Positives = 73/174 (41%), Gaps = 7/174 (4%)
Frame = +3
Query: 356 KDKGAESKEAVDSDSESEGDPDSDDENEVFASFSTSELKHALSDIMDKYNSLLSKHKKLK 415
K K A+ K+ DSDSES D DSDDE++ TS H Y+ S H+K K
Sbjct: 369 KLKAAKHKKRADSDSES--DYDSDDESK-----RTSRRSHRKHKKRSHYD---SDHEKRK 518
Query: 416 KNLS--AVSKTPSEHEKIISDLKNDNHALVNSNSVLKNQIAKLEEVIACDASDSKHE--- 470
+ LS K SE D ++++ + K + ++V D+SDS
Sbjct: 519 EKLSKRKTKKWSSESSDFSGD-ESESRSEEERRRKRKQKKKLRDQVSRSDSSDSDSSADE 695
Query: 471 -SKYEKSFQRF-LAKSVDRSLMASMIYGVSRNGMHGIGYSKPIRNESSVSKAKS 522
SK ++ +R + KS + R HG + + +ESS S S
Sbjct: 696 VSKRKRQHKRHRVTKSSGSDFSSDEENSPVRKRSHGKHHKRHRHSESSESDLSS 857
Score = 36.6 bits (83), Expect = 0.032
Identities = 30/118 (25%), Positives = 49/118 (41%), Gaps = 1/118 (0%)
Frame = +3
Query: 209 SEMQDLRKKSIALKSKSEKAKVEKSKA-LQAEEEESEEASEDSGEDELTLISKRLNRIWK 267
SE D +S+ E+ + K K L+ + S+ + DS DE +SKR + +
Sbjct: 555 SESSDFSGDESESRSEEERRRKRKQKKKLRDQVSRSDSSDSDSSADE---VSKRKRQHKR 725
Query: 268 HRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPKLKKDKKPKKHFK 325
HR +K GS + + S +K+S + ES + K +KH K
Sbjct: 726 HRVTKSSGSDFSSDEENSPVRKRSHGKHHKRHRHSESSESDLSSDDIHHKKSHRKHHK 899
>TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromosome 3, P1
clone: MSJ11, partial (39%)
Length = 1403
Score = 36.2 bits (82), Expect = 0.042
Identities = 40/184 (21%), Positives = 70/184 (37%), Gaps = 31/184 (16%)
Frame = +3
Query: 228 AKVEKSKALQAEEEESEEASEDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSG 287
+K + + L+A + D G E S R++ K + K +GK S+
Sbjct: 234 SKKQLEQYLKAHPGGPAASEFDWGTGETPRRSARISEKVKVTPTPESDPPKKRGKRSSAS 413
Query: 288 QKKSS----IREVTCFECKESGHYKSDCPKLKKDKKPK-----KHFKTKKSLMVTFDESE 338
+K +S E E +E+G K + + + K + + K ++ DE
Sbjct: 414 KKDASGEEEKEETKDVEMQEAGETKDAVKENQDENKAEDTDVNQDEKKAEATDANQDEKR 593
Query: 339 SEDVD----------------------SDGEVQGLMAIVKDKGAESKEAVDSDSESEGDP 376
+ED D +D E+Q V DKGAE +AV + + G+P
Sbjct: 594 AEDADVKEPTHPGENTDGPNDEGKLNTADEELQASKEKVDDKGAEGSDAVQNKDDGSGEP 773
Query: 377 DSDD 380
+ +
Sbjct: 774 EKSE 785
>BP065763
Length = 537
Score = 35.0 bits (79), Expect = 0.093
Identities = 18/46 (39%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Frame = -1
Query: 271 SKFKGSGKAKGKYESSGQKKSSI-REVTCFECKESGHYKSDCPKLK 315
SK K K + K ++ K+ I +E C+ CK++GH+K CPK K
Sbjct: 465 SKAKPGRKEQKKDQALKVKEGRIHKEHVCYFCKKAGHFKKYCPKRK 328
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.335 0.145 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,542,721
Number of Sequences: 28460
Number of extensions: 363342
Number of successful extensions: 3041
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 2929
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3024
length of query: 1610
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1507
effective length of database: 1,966,220
effective search space: 2963093540
effective search space used: 2963093540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0134.9


