

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
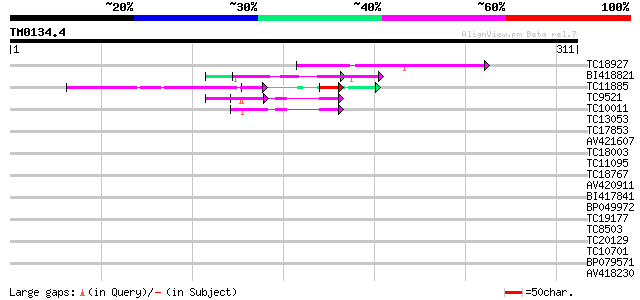
Query= TM0134.4
(311 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 70 5e-13
BI418821 53 8e-08
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 47 1e-06
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 42 1e-04
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 41 2e-04
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 36 0.007
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 35 0.012
AV421607 34 0.036
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 33 0.081
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 32 0.14
TC18767 32 0.14
AV420911 31 0.31
BI417841 30 0.52
BP049972 29 0.89
TC19177 29 0.89
TC8503 similar to GB|AAD30253.1|4835787|F25C20 ESTs gb|R65381 an... 29 0.89
TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-li... 29 0.89
TC10701 similar to UP|Q94A82 (Q94A82) AT5g20070/F28I16_220, part... 29 1.2
BP079571 23 3.1
AV418230 27 4.4
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 70.1 bits (170), Expect = 5e-13
Identities = 39/116 (33%), Positives = 55/116 (46%), Gaps = 10/116 (8%)
Frame = -2
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTA-- 215
P + ++ C+RC + GHFA+RC C+NC + GH C PKVE + N
Sbjct: 365 PRPTQSDTSEIVCHRCSKKGHFANRCPDLV--CWNCQKTGHSGKDCTNPKVEAATNAIAA 192
Query: 216 --------RGKHPAARGKVYTMDGEEAEGADELVKGERKNDGNLLTILSHSSIIRS 263
+GK P A +VYT+ G E+ AD L++ + LTIL S S
Sbjct: 191 RRPAPAANKGKRPVASARVYTVSGAESHRADGLIRSVGSVNCKPLTILFDSGATHS 24
>BI418821
Length = 614
Score = 52.8 bits (125), Expect = 8e-08
Identities = 31/90 (34%), Positives = 40/90 (44%), Gaps = 7/90 (7%)
Frame = +2
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
G G C+NC GHLA +C S+N G A +CY CG+ GH A
Sbjct: 377 GGGGGCYNCGDTGHLARDCHR-----SNNNGGGGGGA---------ACYNCGDAGHLARD 514
Query: 183 CSAT-------GPRCFNCHRVGHLAVVCKK 205
C+ + G C+NC GHLA C +
Sbjct: 515 CNRSNNNSGGGGAGCYNCGDTGHLARDCNR 604
Score = 50.8 bits (120), Expect = 3e-07
Identities = 28/85 (32%), Positives = 34/85 (39%), Gaps = 8/85 (9%)
Frame = +2
Query: 108 CFRCGKNGHYANACL--------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
C+ CG GH A C G G C+NC GHLA +C S+N G
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNR-----SNNNSGG---- 544
Query: 160 IARGTEKQVSCYRCGEIGHFASRCS 184
CY CG+ GH A C+
Sbjct: 545 ------GGAGCYNCGDTGHLARDCN 601
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 47.0 bits (110), Expect(2) = 1e-06
Identities = 30/110 (27%), Positives = 47/110 (42%)
Frame = +2
Query: 32 EAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFKSST 91
E +E + + S+ + R+ +RS YR PY+ RGF+ R
Sbjct: 179 EV*KEEKRKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRD-APYRRDSRRGFS-RDNLCKN 352
Query: 92 TATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC 141
PG ++ C CG GH A+ C + C+NC + GH+A +C
Sbjct: 353 CKRPGHFARECPNVAICHNCGLPGHIASECTTKS-LCWNCKEPGHMASSC 499
Score = 41.2 bits (95), Expect = 2e-04
Identities = 25/76 (32%), Positives = 31/76 (39%)
Frame = +2
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
C NC + GH A C P +A C+ CG GH AS C+ T
Sbjct: 344 CKNCKRPGHFAREC----------------PNVA-------ICHNCGLPGHIASECT-TK 451
Query: 188 PRCFNCHRVGHLAVVC 203
C+NC GH+A C
Sbjct: 452 SLCWNCKEPGHMASSC 499
Score = 21.2 bits (43), Expect(2) = 1e-06
Identities = 6/13 (46%), Positives = 9/13 (69%)
Frame = +1
Query: 171 YRCGEIGHFASRC 183
+ CG++GH A C
Sbjct: 517 HTCGKVGHRAREC 555
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 42.0 bits (97), Expect = 1e-04
Identities = 26/67 (38%), Positives = 29/67 (42%), Gaps = 5/67 (7%)
Frame = +1
Query: 122 LGQGP-----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEI 176
+G+GP RCFNC GH A +CK AG NK CYRCGE
Sbjct: 358 MGRGPPPGSGRCFNCGIDGHWARDCK---AGDWKNK-----------------CYRCGER 477
Query: 177 GHFASRC 183
GH C
Sbjct: 478 GHIEKNC 498
Score = 39.3 bits (90), Expect = 9e-04
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = +1
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCK 142
CF CG +GH+A C +C+ C + GH+ NCK
Sbjct: 391 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 501
Score = 33.5 bits (75), Expect = 0.047
Identities = 18/57 (31%), Positives = 25/57 (43%), Gaps = 2/57 (3%)
Frame = +1
Query: 150 DNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG--PRCFNCHRVGHLAVVCK 204
D + G+ P G C+ CG GH+A C A +C+ C GH+ CK
Sbjct: 346 DREYMGRGPPPGSGR-----CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 501
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 41.2 bits (95), Expect = 2e-04
Identities = 26/67 (38%), Positives = 29/67 (42%), Gaps = 5/67 (7%)
Frame = +2
Query: 122 LGQGP-----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEI 176
LG+GP RCFNC GH A +CK AG NK CYRCG+
Sbjct: 338 LGRGPPPGSGRCFNCGLDGHWARDCK---AGDWKNK-----------------CYRCGDR 457
Query: 177 GHFASRC 183
GH C
Sbjct: 458 GHVERNC 478
Score = 38.1 bits (87), Expect = 0.002
Identities = 15/37 (40%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCK 142
CF CG +GH+A C +C+ C GH+ NCK
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCK 481
Score = 32.0 bits (71), Expect = 0.14
Identities = 20/63 (31%), Positives = 28/63 (43%), Gaps = 3/63 (4%)
Frame = +2
Query: 145 QAGPSDNK-MKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG--PRCFNCHRVGHLAV 201
+ GP N+ G+ P G C+ CG GH+A C A +C+ C GH+
Sbjct: 308 KGGPRGNREYLGRGPPPGSGR-----CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVER 472
Query: 202 VCK 204
CK
Sbjct: 473 NCK 481
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 36.2 bits (82), Expect = 0.007
Identities = 15/36 (41%), Positives = 18/36 (49%)
Frame = +3
Query: 106 VTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC 141
V C C + GH + C+G C NC GHLA C
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
Score = 33.1 bits (74), Expect = 0.062
Identities = 14/36 (38%), Positives = 17/36 (46%)
Frame = +3
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
V C C ++GH + C C NC GHLA C
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 35.4 bits (80), Expect = 0.012
Identities = 19/62 (30%), Positives = 27/62 (42%)
Frame = +2
Query: 64 GSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLG 123
GS R G +GR R + + G+R + CF+CGK GH+A C
Sbjct: 296 GSSRDRDDGDRYRERGRDRDDRGDRDRSRGYGGSRGSNGGE---CFKCGKPGHFARECPS 466
Query: 124 QG 125
+G
Sbjct: 467 EG 472
Score = 29.3 bits (64), Expect = 0.89
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +2
Query: 170 CYRCGEIGHFASRCSATG 187
C++CG+ GHFA C + G
Sbjct: 419 CFKCGKPGHFARECPSEG 472
>AV421607
Length = 245
Score = 33.9 bits (76), Expect = 0.036
Identities = 12/36 (33%), Positives = 19/36 (52%)
Frame = +3
Query: 153 MKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGP 188
M+G P G+ CY+CG GH++ C ++ P
Sbjct: 3 MEGSGPGSGSGSGTATGCYKCGRPGHWSRDCPSSAP 110
Score = 27.3 bits (59), Expect = 3.4
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = +3
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGP 126
PG+ S S C++CG+ GH++ C P
Sbjct: 18 PGSGSGSG-TATGCYKCGRPGHWSRDCPSSAP 110
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 32.7 bits (73), Expect = 0.081
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = +1
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATG 187
RG E + CY CGE GHFA C + G
Sbjct: 145 RGGE-DLKCYECGEPGHFARECRSRG 219
Score = 28.5 bits (62), Expect = 1.5
Identities = 8/21 (38%), Positives = 14/21 (66%)
Frame = +1
Query: 105 EVTCFRCGKNGHYANACLGQG 125
++ C+ CG+ GH+A C +G
Sbjct: 157 DLKCYECGEPGHFARECRSRG 219
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 32.0 bits (71), Expect = 0.14
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +1
Query: 163 GTEKQVSCYRCGEIGHFASRCSATG 187
G + CY CGE GHFA C G
Sbjct: 358 GGGSDMKCYECGEPGHFARECRMRG 432
Score = 27.3 bits (59), Expect = 3.4
Identities = 8/21 (38%), Positives = 14/21 (66%)
Frame = +1
Query: 105 EVTCFRCGKNGHYANACLGQG 125
++ C+ CG+ GH+A C +G
Sbjct: 370 DMKCYECGEPGHFARECRMRG 432
>TC18767
Length = 1004
Score = 32.0 bits (71), Expect = 0.14
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +2
Query: 189 RCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGK 225
RCFNC H C +P+ +VN+AR + + R +
Sbjct: 164 RCFNCGSYNHSLRECSRPRDNVAVNSARKQRKSRRNQ 274
>AV420911
Length = 418
Score = 30.8 bits (68), Expect = 0.31
Identities = 13/22 (59%), Positives = 14/22 (63%)
Frame = +2
Query: 162 RGTEKQVSCYRCGEIGHFASRC 183
RG E + CY CGE GHFA C
Sbjct: 353 RGGE-DLKCYECGEPGHFAREC 415
Score = 26.6 bits (57), Expect = 5.8
Identities = 7/17 (41%), Positives = 12/17 (70%)
Frame = +2
Query: 105 EVTCFRCGKNGHYANAC 121
++ C+ CG+ GH+A C
Sbjct: 365 DLKCYECGEPGHFAREC 415
>BI417841
Length = 617
Score = 30.0 bits (66), Expect = 0.52
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Frame = -3
Query: 240 LVKGERKNDGNLLTILSHSSIIRSITLVAHVT--HLFYLYCLSIDFIVITPNNTL 292
L K E K +L SH+S R +T+ +VT ++ CL+I F VI P N +
Sbjct: 615 LPKAESKKKLPMLDTASHAS--RKLTIFFNVTSWEIYTQICLNISFRVILPRNMI 457
>BP049972
Length = 496
Score = 29.3 bits (64), Expect = 0.89
Identities = 15/39 (38%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = -1
Query: 148 PSDNKMKGKFPAIARGTEKQV-SCYRCGEIGHFASRCSA 185
P K + A RG K+V C RC + GHF + C+A
Sbjct: 304 PPGRPRKKRVRAEDRGRVKRVVHCSRCNQTGHFRTTCAA 188
>TC19177
Length = 538
Score = 29.3 bits (64), Expect = 0.89
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 6/65 (9%)
Frame = +2
Query: 246 KNDGNLLTILSHSSIIRSITLVAHVTHLFYLYCLSIDFIVITPNNTLFDRCSC------L 299
K NL+ I ++S S H+ HL+Y YC+ FI IT + F C
Sbjct: 188 KQGSNLMYIHLYASFFLSNF---HLFHLYYYYCMLF*FICITIFSVSFFFCRLYLGTKPC 358
Query: 300 NYRSY 304
N+RS+
Sbjct: 359 NFRSF 373
>TC8503 similar to GB|AAD30253.1|4835787|F25C20 ESTs gb|R65381 and
gb|T44635 come from this gene. {Arabidopsis thaliana;},
partial (26%)
Length = 1151
Score = 29.3 bits (64), Expect = 0.89
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = -3
Query: 81 GFTPRSFKSSTTATPGARSQSAHPEVTC 108
G T S S +++TPG+ S +PEV+C
Sbjct: 591 GTTSNSLSSRSSSTPGSDSTELNPEVSC 508
>TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (12%)
Length = 588
Score = 29.3 bits (64), Expect = 0.89
Identities = 15/39 (38%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = +3
Query: 148 PSDNKMKGKFPAIARGTEKQV-SCYRCGEIGHFASRCSA 185
P K + A RG K+V C RC + GHF + C+A
Sbjct: 156 PPGRPRKKRVRAEDRGRVKRVVHCSRCNQTGHFRTTCAA 272
>TC10701 similar to UP|Q94A82 (Q94A82) AT5g20070/F28I16_220, partial (32%)
Length = 668
Score = 28.9 bits (63), Expect = 1.2
Identities = 14/47 (29%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Frame = -2
Query: 242 KGERKNDGNLLTILSHSSIIRSITLVAHVTHLFYLYCLS-IDFIVIT 287
KG R G+ + ++++ +THLFYL C S + F+V++
Sbjct: 385 KGSRNKHGSKFATFNVEICSQAVSFFYSLTHLFYLSCCSFLGFLVLS 245
>BP079571
Length = 414
Score = 23.5 bits (49), Expect(2) = 3.1
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = -1
Query: 125 GPRCFNCNQIGHLAVNC 141
G CF ++GHLA +C
Sbjct: 366 GGGCFRFGEVGHLARDC 316
Score = 22.3 bits (46), Expect(2) = 3.1
Identities = 6/13 (46%), Positives = 9/13 (69%)
Frame = -3
Query: 167 QVSCYRCGEIGHF 179
+ C+ CG+ GHF
Sbjct: 271 ETHCFHCGKPGHF 233
>AV418230
Length = 405
Score = 26.9 bits (58), Expect = 4.4
Identities = 19/52 (36%), Positives = 24/52 (45%), Gaps = 1/52 (1%)
Frame = -1
Query: 51 RLDRAGTGGPVRSGSQEYRQLGPYQH-PKGRGFTPRSFKSSTTATPGARSQS 101
R +RAG+ P RSG+ P H K G +P + TTA RS S
Sbjct: 180 RGERAGSSSPRRSGAARISARPPSPHRQKSAGGSPPGEATRTTAVG*LRSPS 25
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,790,056
Number of Sequences: 28460
Number of extensions: 89361
Number of successful extensions: 589
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 495
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 559
length of query: 311
length of database: 4,897,600
effective HSP length: 90
effective length of query: 221
effective length of database: 2,336,200
effective search space: 516300200
effective search space used: 516300200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0134.4


