

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.14
(277 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
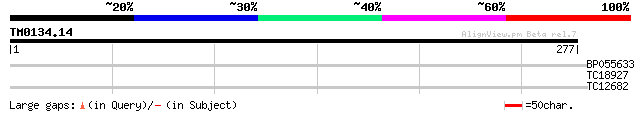
Score E
Sequences producing significant alignments: (bits) Value
BP055633 35 0.011
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 33 0.069
TC12682 26 8.5
>BP055633
Length = 528
Score = 35.4 bits (80), Expect = 0.011
Identities = 32/113 (28%), Positives = 51/113 (44%)
Frame = -2
Query: 132 FRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCK 191
F+ A LE++ TS A +GS V E+ + E A + +DE +
Sbjct: 509 FKEAMLEQFQLTSNSSPFAALLALKQEGS--VEEFVGQFERFAGMLK----GIDEEHYMD 348
Query: 192 RFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSS 244
F+NGL+ +I ++ +V+KA VE KN ++ AG G R +S
Sbjct: 347 IFVNGLKEEIAAEIKLYEPKSLTIMVKKALMVE-QKNLAVSKAGIGSTSRYNS 192
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 32.7 bits (73), Expect = 0.069
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 5/57 (8%)
Frame = -2
Query: 219 KATEVELMKNRRMNGA-----GTGGPMRSSSQDNLGKGRFQMKKPYQRPMGEGYTLG 270
KA +E + N R A G+GGP RS+ + + KKP+QRP G + G
Sbjct: 560 KAKSIEAIDNLRSRPAFRPNQGSGGPNRSAP-GRFDRNKSFQKKPFQRPQNRGTSSG 393
>TC12682
Length = 583
Score = 25.8 bits (55), Expect = 8.5
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 4/72 (5%)
Frame = -2
Query: 108 DAEYW--WKGTRG--IMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTV 163
+A+ W W+G R I++A E FR+AF+ R+ E F + G SMT+
Sbjct: 276 EAQKWGLWEGRRQFKILQAMRREF---MFRSAFVLSMI----RNLLETMFRS-GSASMTI 121
Query: 164 PEYASKLESLAK 175
P Y + L K
Sbjct: 120 PSYTGAIFQLLK 85
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,484,225
Number of Sequences: 28460
Number of extensions: 36401
Number of successful extensions: 207
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 207
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 207
length of query: 277
length of database: 4,897,600
effective HSP length: 89
effective length of query: 188
effective length of database: 2,364,660
effective search space: 444556080
effective search space used: 444556080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0134.14


