

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.3
(700 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
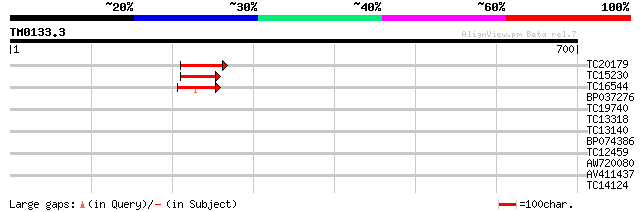
Score E
Sequences producing significant alignments: (bits) Value
TC20179 54 8e-08
TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Ar... 45 4e-05
TC16544 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g07170... 42 3e-04
BP037276 34 0.090
TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, part... 32 0.34
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 29 2.9
TC13140 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger pro... 29 2.9
BP074386 28 5.0
TC12459 28 5.0
AW720080 28 5.0
AV411437 28 6.5
TC14124 UP|Q8H945 (Q8H945) Phosphoenolpyruvate carboxylase , co... 28 6.5
>TC20179
Length = 511
Score = 53.9 bits (128), Expect = 8e-08
Identities = 27/58 (46%), Positives = 37/58 (63%)
Frame = +3
Query: 211 WLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGIA 268
WL LGLG+Y VF EVD + L LT EDL MG++A+G R+K+ A+ + KG +
Sbjct: 24 WLNGLGLGRYAPVFEVHEVDDEVLPMLTLEDLKDMGISAVGSRRKMYCAIQKLGKGFS 197
>TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Arabidopsis
thaliana;}, partial (11%)
Length = 611
Score = 45.1 bits (105), Expect = 4e-05
Identities = 25/49 (51%), Positives = 32/49 (65%)
Frame = +2
Query: 212 LRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
L LGLGKY +F EEVD L+ + E DL +G+ +GPRKKI+ AL
Sbjct: 116 LHALGLGKYVILFKAEEVDMTALKQMGENDLKELGI-PMGPRKKILLAL 259
>TC16544 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g07170/T1B9_17),
partial (33%)
Length = 546
Score = 42.0 bits (97), Expect = 3e-04
Identities = 27/57 (47%), Positives = 36/57 (62%), Gaps = 4/57 (7%)
Frame = +2
Query: 208 VVDWLRDLGLGKYEDVFVREE----VDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V ++L+ LGL KY F EE VD L +T+EDL +MG+ +GPRKKI+ AL
Sbjct: 146 VDEFLQSLGLEKYLISFQAEELLFQVDMTALNHMTDEDLKAMGI-PMGPRKKILLAL 313
>BP037276
Length = 516
Score = 33.9 bits (76), Expect = 0.090
Identities = 24/81 (29%), Positives = 37/81 (45%)
Frame = +2
Query: 40 LKKHKLETTAFGGSGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSS 99
LK H E + SGK+ P + S S + + S + PS+ V + T S+
Sbjct: 260 LKSHTYEKSELINSGKQISPMSKSKSKSVESSFSFPQGSNERTPSSSPLVKSLATAESNG 439
Query: 100 SSSASVSAFVISSPVKVKMKG 120
S SAS + F + S + V + G
Sbjct: 440 SPSASSNNFAMKSSLCVCIGG 502
>TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, partial (4%)
Length = 622
Score = 32.0 bits (71), Expect = 0.34
Identities = 21/58 (36%), Positives = 35/58 (60%)
Frame = +1
Query: 58 VPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVISSPVK 115
VPP++S+S + + C++ I + S+ + T S+SSSS+ + S F ISSPV+
Sbjct: 55 VPPHSSTSL------QSQ*CTMTLITPSTSSISISST-STSSSSNNNRSIFTISSPVQ 207
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 28.9 bits (63), Expect = 2.9
Identities = 17/52 (32%), Positives = 29/52 (55%)
Frame = -3
Query: 60 PNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVIS 111
P+ S+S++ + S PS+ S ++ + SSSSSSS+S S+ +S
Sbjct: 400 PSTSTSSLPSSSSSSSSSSSSSSPSSSSSSSSSSSSSSSSSSSSSSSSSSLS 245
>TC13140 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger protein 12
(LIM domain interacting RING finger protein) (RING
finger LIM domain-binding protein) (R-LIM) (NY-REN-43
antigen), partial (4%)
Length = 526
Score = 28.9 bits (63), Expect = 2.9
Identities = 16/34 (47%), Positives = 22/34 (64%)
Frame = +3
Query: 82 IPSTIDSVGTALTDSSSSSSSASVSAFVISSPVK 115
I S+ S ++L DSSS SSS S S V+ SP++
Sbjct: 174 ISSSAVSSASSLLDSSSPSSSPSSSPRVLDSPIR 275
>BP074386
Length = 496
Score = 28.1 bits (61), Expect = 5.0
Identities = 39/166 (23%), Positives = 64/166 (38%), Gaps = 3/166 (1%)
Frame = +3
Query: 262 EFRKGIAASNEKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFAT 321
E RKG +++EKP DA R +N+ + K KK Y +
Sbjct: 72 EKRKGAGSTSEKPLDAKTLRRLAQNR-------------EAAKKSRLRKK--AYVQQLES 206
Query: 322 NGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKYLR 381
+ K+S ++ Q ++S G G G N S+ + F ++ ++L
Sbjct: 207 SRLKLSHLEQDLQRLRSQGLFLGCGVAGGNLSSGAAM-------------FDMEYARWLE 347
Query: 382 GDCSHWFLTHFHLDRTRL-LDLNIFDTDLRQM--GYLIKFECFARL 424
D H H+ R L + D +LR + GYL ++ RL
Sbjct: 348 DD-------HQHMAELRTGLQAPLSDNELRVIVDGYLYHYDELFRL 464
>TC12459
Length = 797
Score = 28.1 bits (61), Expect = 5.0
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = +3
Query: 313 TEYFPGFATNGKKVSTAPEEQQEMKSSGS 341
T YFP G+ +PE QQE K GS
Sbjct: 27 TRYFPASTCGGEDDRPSPEIQQEQKRRGS 113
>AW720080
Length = 394
Score = 28.1 bits (61), Expect = 5.0
Identities = 19/59 (32%), Positives = 25/59 (42%)
Frame = +3
Query: 288 NVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGKKVSTAPEEQQEMKSSGSVSGRN 346
+V M P VDGT P+ + K TNG V +A E S+ SG+N
Sbjct: 90 SVSMAPSEEAXAVDGTLNPSEHGKSE------ITNGATVDSAAIGGAESASNAETSGKN 248
>AV411437
Length = 417
Score = 27.7 bits (60), Expect = 6.5
Identities = 20/53 (37%), Positives = 28/53 (52%)
Frame = -1
Query: 59 PPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVIS 111
PP+ S+S+ + NCS PST SSSSSSS+S S+ ++S
Sbjct: 171 PPSVSNSSSSESVSSSSNCSNS--PST--------PSSSSSSSSSSSSSAIVS 43
Score = 27.3 bits (59), Expect = 8.5
Identities = 21/63 (33%), Positives = 32/63 (50%)
Frame = -1
Query: 50 FGGSGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFV 109
F +G E PP +SS+ + S + + S+ + + T SSSSSSS+S S+
Sbjct: 228 FSTNGDEVEPPTWASSSSPPPSVS-NSSSSESVSSSSNCSNSPSTPSSSSSSSSSSSSSA 52
Query: 110 ISS 112
I S
Sbjct: 51 IVS 43
>TC14124 UP|Q8H945 (Q8H945) Phosphoenolpyruvate carboxylase , complete
Length = 3367
Score = 27.7 bits (60), Expect = 6.5
Identities = 16/40 (40%), Positives = 22/40 (55%), Gaps = 4/40 (10%)
Frame = +2
Query: 283 RNRNQNVKMQPHRPERKVDGTSKPAAN----KKITEYFPG 318
R+ N NVK++PH + +D SKPA +EY PG
Sbjct: 2852 RDPNYNVKLRPHISKEAID-VSKPADELVTLNPTSEYAPG 2968
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,001,755
Number of Sequences: 28460
Number of extensions: 173976
Number of successful extensions: 1215
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1020
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1132
length of query: 700
length of database: 4,897,600
effective HSP length: 97
effective length of query: 603
effective length of database: 2,136,980
effective search space: 1288598940
effective search space used: 1288598940
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0133.3


